Top 10 Significant Motifs - Global Matching (Highest to Lowest)
| Dataset #: 1 |
Motif ID: 5 |
Motif name: ZEB1 |
| Original motif Consensus sequence: CACCTD |
Reverse complement motif Consensus sequence: HAGGTG |
|
|
 |
 |
Best Matches for Top Significant Motif ID 5 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
6 |
| Similarity score: |
0.0349782 |
Alignment:
AMTGTGACACCACAGT
-------CACCTD---
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
6 |
| Similarity score: |
0.0362768 |
Alignment:
TGTGGTGTCACAGTGCTCC
-HAGGTG------------
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
12 |
| Number of overlap: |
6 |
| Similarity score: |
0.0409613 |
Alignment:
AGTGCTCCACTGTGGTGTCACAGT
-----------HAGGTG-------
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
33 |
| Motif name: |
srACyCCGAyr |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
6 |
| Similarity score: |
0.0426002 |
Alignment:
KKTCGGKGTKS
--HAGGTG---
| Original motif Consensus sequence: SRACYCCGAYR |
Reverse complement motif Consensus sequence: KKTCGGKGTKS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
6 |
| Similarity score: |
0.0496179 |
Alignment:
TGGAGCACTGTGACAVCACAGTGG
-------------CACCTD-----
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
6 |
| Similarity score: |
0.0538743 |
Alignment:
AGTGCTCCACTGTGDTG
-----------HAGGTG
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
19 |
| Motif name: |
atactttggc |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
6 |
| Similarity score: |
0.0553034 |
Alignment:
ATACTTTGGC
-CACCTD---
| Original motif Consensus sequence: ATACTTTGGC |
Reverse complement motif Consensus sequence: GCCAAAGTAT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
19 |
| Number of overlap: |
6 |
| Similarity score: |
0.0564649 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
------------------CACCTD-
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
6 |
| Similarity score: |
0.0571854 |
Alignment:
SBCCCCGCCSBB
-----CACCTD-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
8 |
| Number of overlap: |
6 |
| Similarity score: |
0.0594701 |
Alignment:
ABCACAGTGGAGCACTG
----HAGGTG-------
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 6 |
Motif name: Mycn |
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
Best Matches for Top Significant Motif ID 6 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0654819 |
Alignment:
BSCCCCGCGBS
-GCCACGTGSD
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0704988 |
Alignment:
BBSGGCGGGGBS
-HSCACGTGGC-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
10 |
| Similarity score: |
0.0883224 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
------GCCACGTGSD---------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0943014 |
Alignment:
ABCACAGTGGAGCACTG
-GCCACGTGSD------
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.0949359 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
---------GCCACGTGSD-----
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0955115 |
Alignment:
TGGAGCACTGTGACAVCACAGTGG
--------------HSCACGTGGC
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.0959628 |
Alignment:
CAHCACAGTGGAGCACT
--GCCACGTGSD-----
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0989932 |
Alignment:
GCGCGCSCGCG
-HSCACGTGGC
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
10 |
| Similarity score: |
0.0991692 |
Alignment:
GGAGCACTGTGACACCACA
---GCCACGTGSD------
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
28 |
| Motif name: |
ssGCkTGCssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.101333 |
Alignment:
YSSGCAYGCSS
HSCACGTGGC-
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 8 |
Motif name: Myc |
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
Best Matches for Top Significant Motif ID 8 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.064853 |
Alignment:
BSCCCCGCGBS
-DCCACGTGCV
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0728588 |
Alignment:
SBCCCCGCCSBB
-DCCACGTGCV-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.0888909 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
----------DCCACGTGCV-----
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0955234 |
Alignment:
ABCACAGTGGAGCACTG
-DCCACGTGCV------
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
11 |
| Number of overlap: |
10 |
| Similarity score: |
0.0971125 |
Alignment:
TGGAGCACTGTGACAVCACAGTGG
----DCCACGTGCV----------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0974038 |
Alignment:
VDGACTRTGGDD
-VGCACGTGGH-
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.0979703 |
Alignment:
CAHCACAGTGGAGCACT
--DCCACGTGCV-----
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.0980018 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
---------DCCACGTGCV-----
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0989125 |
Alignment:
CGCGSGCGCGC
VGCACGTGGH-
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0993196 |
Alignment:
GGAGCACTGTGACACCACA
---DCCACGTGCV------
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 11 |
Motif name: NR4A2 |
| Original motif Consensus sequence: AAGGTCAC |
Reverse complement motif Consensus sequence: GTGACCTT |
|
|
 |
 |
Best Matches for Top Significant Motif ID 11 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
8 |
| Similarity score: |
0.0557243 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
------AAGGTCAC-----------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
8 |
| Similarity score: |
0.060729 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
-------AAGGTCAC---------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
0.0645146 |
Alignment:
ACTGTGGTGTCACART
-----AAGGTCAC---
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
0.0695512 |
Alignment:
TGTGGTGTCACAGTGCTCC
---AAGGTCAC--------
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
0.0709831 |
Alignment:
AGTGCTCCACTGTGGTGTCACAGT
-------------AAGGTCAC---
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
8 |
| Similarity score: |
0.0772292 |
Alignment:
CCGCGGRCACG
--AAGGTCAC-
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
8 |
| Similarity score: |
0.0788401 |
Alignment:
DDCCAMAGTCHB
----AAGGTCAC
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
10 |
| Number of overlap: |
8 |
| Similarity score: |
0.0864039 |
Alignment:
ABCACAGTGGAGCACTG
---------GTGACCTT
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
28 |
| Motif name: |
ssGCkTGCssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
7 |
| Similarity score: |
0.568556 |
Alignment:
SSGCKTGCSSK-
----GTGACCTT
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
11 |
| Number of overlap: |
7 |
| Similarity score: |
0.581814 |
Alignment:
CAHCACAGTGGAGCACT-
----------GTGACCTT
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 25 |
Motif name: wwCCAmAGTCmt |
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
Best Matches for Top Significant Motif ID 25 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.032442 |
Alignment:
RGGBCAAAGKYCA
-DDCCAMAGTCHB
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
12 |
| Similarity score: |
0.0391228 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
---DDCCAMAGTCHB------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
12 |
| Similarity score: |
0.0407304 |
Alignment:
VDBHMAGGTCACCCTGACCY
DDCCAMAGTCHB--------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.0463384 |
Alignment:
AAAGGTCAAAGGTCAAC
----DDCCAMAGTCHB-
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
12 |
| Similarity score: |
0.0480406 |
Alignment:
TGRCCTKTGHCCKAB
--DDCCAMAGTCHB-
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
12 |
| Similarity score: |
0.0506507 |
Alignment:
TGAMCTTTGMMCYT
--VDGACTRTGGDD
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
8 |
| Number of overlap: |
12 |
| Similarity score: |
0.0523578 |
Alignment:
BMSMGCCYMCTKSTGGMHM
DDCCAMAGTCHB-------
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
12 |
| Similarity score: |
0.0576723 |
Alignment:
VHRGGTCABDBTGMCCTB
---DDCCAMAGTCHB---
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.525631 |
Alignment:
VBBYCAAGGTCA-
-DDCCAMAGTCHB
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
1.0578 |
Alignment:
--VGCACGTGGH
VDGACTRTGGDD
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 28 |
Motif name: ssGCkTGCssk |
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
Best Matches for Top Significant Motif ID 28 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
11 |
| Similarity score: |
0.0349439 |
Alignment:
YDRCCASYAGRKGGCRSYV
-------SSGCKTGCSSK-
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0358701 |
Alignment:
VBBYCAAGGTCA
-YSSGCAYGCSS
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.0379074 |
Alignment:
VHRGGTCABDBTGMCCTB
-----YSSGCAYGCSS--
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
11 |
| Similarity score: |
0.0409322 |
Alignment:
TTCAGCACCATGGACAGCKCC
-YSSGCAYGCSS---------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.041419 |
Alignment:
BTRGGDCARAGGKCA
--YSSGCAYGCSS--
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.0437981 |
Alignment:
RGGBCAAAGKYCA
--YSSGCAYGCSS
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.53887 |
Alignment:
GCCACGTGSD-
SSGCKTGCSSK
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.542915 |
Alignment:
DCCACGTGCV-
SSGCKTGCSSK
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
8 |
| Similarity score: |
1.53583 |
Alignment:
---AAAGGTCAAAGGTCAAC
YSSGCAYGCSS---------
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
15 |
| Motif name: |
Tcfcp2l1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
8 |
| Similarity score: |
1.53704 |
Alignment:
---CCAGYYHVADCCRG
YSSGCAYGCSS------
| Original motif Consensus sequence: CCAGYYHVADCCRG |
Reverse complement motif Consensus sequence: CKGGDTBDMMCTGG |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 24 |
Motif name: ssCCCCGCSssk |
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
Best Matches for Top Significant Motif ID 24 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.063076 |
Alignment:
MGGTCAGGGTGACCTRDBHV
-------BBSGGCGGGGBS-
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.0642087 |
Alignment:
YDRCCASYAGRKGGCRSYV
-BBSGGCGGGGBS------
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
12 |
| Similarity score: |
0.0671548 |
Alignment:
TGRCCTKTGHCCKAB
-SBCCCCGCCSBB--
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.0734481 |
Alignment:
BAGGYCABHBTGACCKHV
-SBCCCCGCCSBB-----
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
12 |
| Similarity score: |
0.0751084 |
Alignment:
TTCAGCACCATGGACAGCKCC
BBSGGCGGGGBS---------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.571364 |
Alignment:
VBBYCAAGGTCA-
-BBSGGCGGGGBS
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.573202 |
Alignment:
-TGMYCTTTGBCCK
SBCCCCGCCSBB--
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
15 |
| Motif name: |
Tcfcp2l1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
9 |
| Similarity score: |
1.54913 |
Alignment:
CKGGDTBDMMCTGG---
-----BBSGGCGGGGBS
| Original motif Consensus sequence: CCAGYYHVADCCRG |
Reverse complement motif Consensus sequence: CKGGDTBDMMCTGG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
1.57507 |
Alignment:
---HSCACGTGGC
SBCCCCGCCSBB-
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
1.57737 |
Alignment:
VGCACGTGGH---
-BBSGGCGGGGBS
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 29 |
Motif name: cCGCGGrCACG |
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
Best Matches for Top Significant Motif ID 29 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
11 |
| Similarity score: |
0.0484441 |
Alignment:
TTCAGCACCATGGACAGCKCC
-------CCGCGGRCACG---
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.0486581 |
Alignment:
MGGTCAGGGTGACCTRDBHV
-------CGTGKCCGCGG--
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
11 |
| Similarity score: |
0.0588663 |
Alignment:
YDRCCASYAGRKGGCRSYV
-------CCGCGGRCACG-
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0629476 |
Alignment:
TGRCCTKTGHCCKAB
---CGTGKCCGCGG-
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
9 |
| Number of overlap: |
10 |
| Similarity score: |
0.551597 |
Alignment:
BAGGYCABHBTGACCKHV-
--------CGTGKCCGCGG
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
1.0488 |
Alignment:
VGCACGTGGH--
-CGTGKCCGCGG
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
1.04892 |
Alignment:
HSCACGTGGC--
-CGTGKCCGCGG
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
13 |
| Motif name: |
MIZF |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
1.04904 |
Alignment:
--BAACGTCCGC
CGTGKCCGCGG-
| Original motif Consensus sequence: BAACGTCCGC |
Reverse complement motif Consensus sequence: GCGGACGTTV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
9 |
| Similarity score: |
1.05291 |
Alignment:
VBBYCAAGGTCA--
---CCGCGGRCACG
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
8 |
| Similarity score: |
1.55795 |
Alignment:
TGMYCTTTGBCCK---
-----CGTGKCCGCGG
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 4 |
Motif name: Esrrb |
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
Best Matches for Top Significant Motif ID 4 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
0.0772556 |
Alignment:
TGTGGTGTCACAGTGCTCC
-------TGACCTTGMBBB
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
12 |
| Number of overlap: |
12 |
| Similarity score: |
0.0785385 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
-----------TGACCTTGMBBB-
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
11 |
| Number of overlap: |
12 |
| Similarity score: |
0.0832722 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
---VBBYCAAGGTCA----------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.558919 |
Alignment:
-DDCCAMAGTCHB
VBBYCAAGGTCA-
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
28 |
| Motif name: |
ssGCkTGCssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.581406 |
Alignment:
-SSGCKTGCSSK
TGACCTTGMBBB
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.583683 |
Alignment:
-SBCCCCGCCSBB
TGACCTTGMBBB-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
15 |
| Number of overlap: |
10 |
| Similarity score: |
1.08683 |
Alignment:
AGTGCTCCACTGTGGTGTCACAGT--
--------------VBBYCAAGGTCA
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
19 |
| Motif name: |
atactttggc |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
1.08726 |
Alignment:
--GCCAAAGTAT
VBBYCAAGGTCA
| Original motif Consensus sequence: ATACTTTGGC |
Reverse complement motif Consensus sequence: GCCAAAGTAT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
9 |
| Similarity score: |
1.57579 |
Alignment:
---CCGCGGRCACG
VBBYCAAGGTCA--
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
7 |
| Similarity score: |
2.57595 |
Alignment:
-----AMTGTGACACCACAGT
VBBYCAAGGTCA---------
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 13 |
Motif name: MIZF |
| Original motif Consensus sequence: BAACGTCCGC |
Reverse complement motif Consensus sequence: GCGGACGTTV |
|
|
 |
 |
Best Matches for Top Significant Motif ID 13 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0651598 |
Alignment:
GCGCGCSCGCG
BAACGTCCGC-
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
19 |
| Motif name: |
atactttggc |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0803571 |
Alignment:
ATACTTTGGC
BAACGTCCGC
| Original motif Consensus sequence: ATACTTTGGC |
Reverse complement motif Consensus sequence: GCCAAAGTAT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
9 |
| Similarity score: |
0.560913 |
Alignment:
CCGCGGRCACG-
--GCGGACGTTV
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
31 |
| Motif name: |
ctatacggacg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
9 |
| Similarity score: |
0.562302 |
Alignment:
-CGTCCGTATAG
GCGGACGTTV--
| Original motif Consensus sequence: CTATACGGACG |
Reverse complement motif Consensus sequence: CGTCCGTATAG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
9 |
| Similarity score: |
0.562428 |
Alignment:
DDCCAMAGTCHB-
---BAACGTCCGC
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
1.06446 |
Alignment:
SBCGCGGGGSB--
---GCGGACGTTV
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
8 |
| Similarity score: |
1.07197 |
Alignment:
BBSGGCGGGGBS--
----GCGGACGTTV
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
33 |
| Motif name: |
srACyCCGAyr |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
7 |
| Similarity score: |
1.55 |
Alignment:
SRACYCCGAYR---
----GCGGACGTTV
| Original motif Consensus sequence: SRACYCCGAYR |
Reverse complement motif Consensus sequence: KKTCGGKGTKS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
18 |
| Number of overlap: |
7 |
| Similarity score: |
1.5706 |
Alignment:
---AGTGCTCCACTGTGGTGTCACAGT
BAACGTCCGC-----------------
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
11 |
| Number of overlap: |
7 |
| Similarity score: |
1.57176 |
Alignment:
CAHCACAGTGGAGCACT---
----------GCGGACGTTV
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
Significant Motifs - Global and Local Matching (Highest to Lowest)
| Dataset #: 1 |
Motif ID: 5 |
Motif name: ZEB1 |
| Original motif Consensus sequence: CACCTD |
Reverse complement motif Consensus sequence: HAGGTG |
|
|
 |
 |
Best Matches for Significant Motif ID 5 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
6 |
| Similarity score: |
0 |
Alignment:
YDRCCASYAGRKGGCRSYV
-------HAGGTG------
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
12 |
| Motif name: |
RORA_1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
6 |
| Similarity score: |
0.00688881 |
Alignment:
TGACCTDVWT
-CACCTD---
| Original motif Consensus sequence: AWVDAGGTCA |
Reverse complement motif Consensus sequence: TGACCTDVWT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
6 |
| Similarity score: |
0.00707323 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
----------HAGGTG-----
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
6 |
| Similarity score: |
0.013046 |
Alignment:
VBBYCAAGGTCA
-HAGGTG-----
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
6 |
| Similarity score: |
0.0160352 |
Alignment:
GTTGACCTTTGACCTTT
----------CACCTD-
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
6 |
| Similarity score: |
0.0192435 |
Alignment:
TGRCCTKTGHCCKAB
-CACCTD--------
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
11 |
| Motif name: |
NR4A2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
6 |
| Similarity score: |
0.0216496 |
Alignment:
GTGACCTT
--CACCTD
| Original motif Consensus sequence: AAGGTCAC |
Reverse complement motif Consensus sequence: GTGACCTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
6 |
| Similarity score: |
0.022947 |
Alignment:
MGGTCAGGGTGACCTRDBHV
----CACCTD----------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
6 |
| Similarity score: |
0.0246463 |
Alignment:
HSCACGTGGC
--CACCTD--
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
6 |
| Similarity score: |
0.025129 |
Alignment:
DCCACGTGCV
--HAGGTG--
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 28 |
Motif name: ssGCkTGCssk |
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
Best Matches for Significant Motif ID 28 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.00456539 |
Alignment:
GCGCGCSCGCG
SSGCKTGCSSK
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0176347 |
Alignment:
BBSGGCGGGGBS
-YSSGCAYGCSS
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0190954 |
Alignment:
SBCGCGGGGSB
YSSGCAYGCSS
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0276515 |
Alignment:
CCGCGGRCACG
SSGCKTGCSSK
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
11 |
| Similarity score: |
0.0349439 |
Alignment:
YDRCCASYAGRKGGCRSYV
-------SSGCKTGCSSK-
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0358701 |
Alignment:
VBBYCAAGGTCA
-YSSGCAYGCSS
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.0379074 |
Alignment:
VHRGGTCABDBTGMCCTB
-----YSSGCAYGCSS--
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
11 |
| Similarity score: |
0.0409322 |
Alignment:
TTCAGCACCATGGACAGCKCC
-YSSGCAYGCSS---------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.041419 |
Alignment:
BTRGGDCARAGGKCA
--YSSGCAYGCSS--
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.0437981 |
Alignment:
RGGBCAAAGKYCA
--YSSGCAYGCSS
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 25 |
Motif name: wwCCAmAGTCmt |
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
Best Matches for Significant Motif ID 25 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0263648 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
--------DDCCAMAGTCHB-----
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0282828 |
Alignment:
ABCACAGTGGAGCACTG
VDGACTRTGGDD-----
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0291095 |
Alignment:
AGTGCTCCACTGTGDTG
-----VDGACTRTGGDD
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0301919 |
Alignment:
AGTGCTCCACTGTGGTGTCACAGT
-----VDGACTRTGGDD-------
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.032442 |
Alignment:
RGGBCAAAGKYCA
-DDCCAMAGTCHB
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
12 |
| Similarity score: |
0.0360702 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
---DDCCAMAGTCHB---------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
12 |
| Similarity score: |
0.0391228 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
---DDCCAMAGTCHB------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
12 |
| Similarity score: |
0.0393782 |
Alignment:
TGTGGTGTCACAGTGCTCC
-----DDCCAMAGTCHB--
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
12 |
| Similarity score: |
0.0407304 |
Alignment:
VDBHMAGGTCACCCTGACCY
DDCCAMAGTCHB--------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.0463384 |
Alignment:
AAAGGTCAAAGGTCAAC
----DDCCAMAGTCHB-
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 19 |
Motif name: atactttggc |
| Original motif Consensus sequence: ATACTTTGGC |
Reverse complement motif Consensus sequence: GCCAAAGTAT |
|
|
 |
 |
Best Matches for Significant Motif ID 19 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0215909 |
Alignment:
DDCCAMAGTCHB
-GCCAAAGTAT-
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
30 |
| Motif name: |
taaacgatgcc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0375 |
Alignment:
GGCATCGTTTA
GCCAAAGTAT-
| Original motif Consensus sequence: TAAACGATGCC |
Reverse complement motif Consensus sequence: GGCATCGTTTA |
|
|
 |
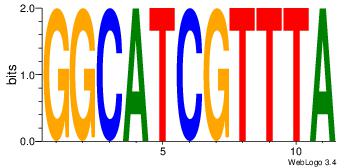 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
10 |
| Similarity score: |
0.0395 |
Alignment:
GTTGACCTTTGACCTTT
----ATACTTTGGC---
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
10 |
| Similarity score: |
0.0472222 |
Alignment:
GGAGCACTGTGACACCACA
------ATACTTTGGC---
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
10 |
| Number of overlap: |
10 |
| Similarity score: |
0.0474206 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
---------GCCAAAGTAT------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.0483209 |
Alignment:
TGMYCTTTGBCCK
--ATACTTTGGC-
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0490385 |
Alignment:
TGAMCTTTGMMCYT
-ATACTTTGGC---
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
10 |
| Similarity score: |
0.0493243 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
----------GCCAAAGTAT----
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
9 |
| Number of overlap: |
10 |
| Similarity score: |
0.0498239 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
--------GCCAAAGTAT------
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0527344 |
Alignment:
AGTGCTCCACTGTGDTG
------ATACTTTGGC-
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 11 |
Motif name: NR4A2 |
| Original motif Consensus sequence: AAGGTCAC |
Reverse complement motif Consensus sequence: GTGACCTT |
|
|
 |
 |
Best Matches for Significant Motif ID 11 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
8 |
| Similarity score: |
0.0167759 |
Alignment:
AAAGGTCAAAGGTCAAC
-AAGGTCAC--------
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
8 |
| Similarity score: |
0.0195235 |
Alignment:
VDBHMAGGTCACCCTGACCY
----AAGGTCAC--------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
8 |
| Similarity score: |
0.0198372 |
Alignment:
BAGGYCABHBTGACCKHV
---------GTGACCTT-
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
8 |
| Similarity score: |
0.0495607 |
Alignment:
BTRGGDCARAGGKCA
-AAGGTCAC------
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
8 |
| Similarity score: |
0.0557243 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
------AAGGTCAC-----------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
8 |
| Similarity score: |
0.0564874 |
Alignment:
AKGYYCAAAGRTCA
------AAGGTCAC
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
8 |
| Similarity score: |
0.0594649 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
----GTGACCTT---------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
8 |
| Similarity score: |
0.060729 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
-------AAGGTCAC---------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
0.0645146 |
Alignment:
ACTGTGGTGTCACART
-----AAGGTCAC---
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
8 |
| Similarity score: |
0.0695512 |
Alignment:
TGTGGTGTCACAGTGCTCC
---AAGGTCAC--------
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 4 |
Motif name: Esrrb |
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
Best Matches for Significant Motif ID 4 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
0.0442611 |
Alignment:
TGAMCTTTGMMCYT
--TGACCTTGMBBB
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
12 |
| Similarity score: |
0.04699 |
Alignment:
BTRGGDCARAGGKCA
---VBBYCAAGGTCA
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
0.0470808 |
Alignment:
RGGBCAAAGKYCA
-VBBYCAAGGTCA
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
12 |
| Similarity score: |
0.0506977 |
Alignment:
AAAGGTCAAAGGTCAAC
---VBBYCAAGGTCA--
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
12 |
| Similarity score: |
0.0628484 |
Alignment:
VDBHMAGGTCACCCTGACCY
TGACCTTGMBBB--------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0694613 |
Alignment:
VHRGGTCABDBTGMCCTB
-----TGACCTTGMBBB-
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
12 |
| Similarity score: |
0.0713236 |
Alignment:
TTCAGCACCATGGACAGCKCC
----VBBYCAAGGTCA-----
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
0.0772556 |
Alignment:
TGTGGTGTCACAGTGCTCC
-------TGACCTTGMBBB
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
12 |
| Number of overlap: |
12 |
| Similarity score: |
0.0785385 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
-----------TGACCTTGMBBB-
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
11 |
| Number of overlap: |
12 |
| Similarity score: |
0.0832722 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
---VBBYCAAGGTCA----------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 1 |
Motif name: HNF4A |
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
Best Matches for Significant Motif ID 1 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
0 |
Alignment:
TGRCCTKTGHCCKAB
TGMYCTTTGBCCK--
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
13 |
| Similarity score: |
0.0227413 |
Alignment:
AKGYYCAAAGRTCA
-RGGBCAAAGKYCA
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
0.0423023 |
Alignment:
AAAGGTCAAAGGTCAAC
--RGGBCAAAGKYCA--
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
13 |
| Similarity score: |
0.0557654 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
-------RGGBCAAAGKYCA-----
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
13 |
| Similarity score: |
0.0656443 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
--------RGGBCAAAGKYCA---
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
0.0671615 |
Alignment:
VHRGGTCABDBTGMCCTB
--RGGBCAAAGKYCA---
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
0.0681338 |
Alignment:
TGTGGTGTCACAGTGCTCC
----RGGBCAAAGKYCA--
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
13 |
| Similarity score: |
0.0711137 |
Alignment:
MGGTCAGGGTGACCTRDBHV
RGGBCAAAGKYCA-------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
13 |
| Similarity score: |
0.0741922 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
-----TGMYCTTTGBCCK---
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
13 |
| Similarity score: |
0.0799096 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
------RGGBCAAAGKYCA-----
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
13 |
| Similarity score: |
0.0850642 |
Alignment:
YDRCCASYAGRKGGCRSYV
---RGGBCAAAGKYCA---
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 12 |
Motif name: RORA_1 |
| Original motif Consensus sequence: AWVDAGGTCA |
Reverse complement motif Consensus sequence: TGACCTDVWT |
|
|
 |
 |
Best Matches for Significant Motif ID 12 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.029401 |
Alignment:
VBBYCAAGGTCA
--AWVDAGGTCA
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0417851 |
Alignment:
MGGTCAGGGTGACCTRDBHV
---------TGACCTDVWT-
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.05425 |
Alignment:
AAAGGTCAAAGGTCAAC
-----AWVDAGGTCA--
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
10 |
| Similarity score: |
0.0654808 |
Alignment:
TGAMCTTTGMMCYT
----TGACCTDVWT
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.06668 |
Alignment:
BTRGGDCARAGGKCA
-----AWVDAGGTCA
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0755608 |
Alignment:
TTCAGCACCATGGACAGCKCC
------AWVDAGGTCA-----
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
10 |
| Similarity score: |
0.0803396 |
Alignment:
RGGBCAAAGKYCA
---AWVDAGGTCA
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0889306 |
Alignment:
GGAGCACTGTGACACCACA
---------TGACCTDVWT
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.09225 |
Alignment:
DDCCAMAGTCHB
-AWVDAGGTCA-
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
10 |
| Similarity score: |
0.0988214 |
Alignment:
BAGGYCABHBTGACCKHV
-----AWVDAGGTCA---
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 6 |
Motif name: Mycn |
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
Best Matches for Significant Motif ID 6 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0 |
Alignment:
DCCACGTGCV
GCCACGTGSD
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0654819 |
Alignment:
BSCCCCGCGBS
-GCCACGTGSD
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.0679017 |
Alignment:
YDRCCASYAGRKGGCRSYV
-----HSCACGTGGC----
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0704988 |
Alignment:
BBSGGCGGGGBS
-HSCACGTGGC-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
10 |
| Similarity score: |
0.0883224 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
------GCCACGTGSD---------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.089863 |
Alignment:
MGGTCAGGGTGACCTRDBHV
--------GCCACGTGSD--
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0943014 |
Alignment:
ABCACAGTGGAGCACTG
-GCCACGTGSD------
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0947124 |
Alignment:
TGMYCTTTGBCCK
-GCCACGTGSD--
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.0949359 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
---------GCCACGTGSD-----
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
10 |
| Similarity score: |
0.0954444 |
Alignment:
TGRCCTKTGHCCKAB
-GCCACGTGSD----
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 8 |
Motif name: Myc |
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
Best Matches for Significant Motif ID 8 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0 |
Alignment:
HSCACGTGGC
VGCACGTGGH
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.064853 |
Alignment:
BSCCCCGCGBS
-DCCACGTGCV
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0728588 |
Alignment:
SBCCCCGCCSBB
-DCCACGTGCV-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
10 |
| Similarity score: |
0.0754257 |
Alignment:
BMSMGCCYMCTKSTGGMHM
-------VGCACGTGGH--
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.0888909 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
----------DCCACGTGCV-----
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0934488 |
Alignment:
TGMYCTTTGBCCK
---VGCACGTGGH
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.0940438 |
Alignment:
MGGTCAGGGTGACCTRDBHV
--------DCCACGTGCV--
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0955234 |
Alignment:
ABCACAGTGGAGCACTG
-DCCACGTGCV------
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
10 |
| Similarity score: |
0.0968598 |
Alignment:
VHRGGTCABDBTGMCCTB
----VGCACGTGGH----
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
11 |
| Number of overlap: |
10 |
| Similarity score: |
0.0971125 |
Alignment:
TGGAGCACTGTGACAVCACAGTGG
----DCCACGTGCV----------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
Best Matches for Each Motif (Highest to Lowest)
| Dataset #: 1 |
Motif ID: 1 |
Motif name: HNF4A |
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
Best Matches for Motif ID 1 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
0 |
Alignment:
TGRCCTKTGHCCKAB
TGMYCTTTGBCCK--
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
13 |
| Similarity score: |
0.0227413 |
Alignment:
AKGYYCAAAGRTCA
-RGGBCAAAGKYCA
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
0.0423023 |
Alignment:
AAAGGTCAAAGGTCAAC
--RGGBCAAAGKYCA--
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
13 |
| Similarity score: |
0.0557654 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
-------RGGBCAAAGKYCA-----
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
13 |
| Similarity score: |
0.0656443 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
--------RGGBCAAAGKYCA---
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
0.0671615 |
Alignment:
VHRGGTCABDBTGMCCTB
--RGGBCAAAGKYCA---
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
0.0681338 |
Alignment:
TGTGGTGTCACAGTGCTCC
----RGGBCAAAGKYCA--
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
13 |
| Similarity score: |
0.0711137 |
Alignment:
MGGTCAGGGTGACCTRDBHV
RGGBCAAAGKYCA-------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
13 |
| Similarity score: |
0.0741922 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
-----TGMYCTTTGBCCK---
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
13 |
| Similarity score: |
0.0799096 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
------RGGBCAAAGKYCA-----
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
13 |
| Similarity score: |
0.0850642 |
Alignment:
YDRCCASYAGRKGGCRSYV
---RGGBCAAAGKYCA---
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 2 |
Motif name: NR2F1 |
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
Best Matches for Motif ID 2 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
14 |
| Similarity score: |
0.0174588 |
Alignment:
GTTGACCTTTGACCTTT
--TGAMCTTTGMMCYT-
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
14 |
| Similarity score: |
0.0278283 |
Alignment:
TGRCCTKTGHCCKAB
TGAMCTTTGMMCYT-
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
14 |
| Similarity score: |
0.0597647 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
-----TGAMCTTTGMMCYT------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
14 |
| Similarity score: |
0.0710475 |
Alignment:
BAGGYCABHBTGACCKHV
---TGAMCTTTGMMCYT-
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
14 |
| Similarity score: |
0.071796 |
Alignment:
TTCAGCACCATGGACAGCKCC
--AKGYYCAAAGRTCA-----
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
8 |
| Number of overlap: |
14 |
| Similarity score: |
0.0737285 |
Alignment:
TGGAGCACTGTGACAVCACAGTGG
---TGAMCTTTGMMCYT-------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
14 |
| Similarity score: |
0.0739698 |
Alignment:
GGAGCACTGTGACACCACA
--TGAMCTTTGMMCYT---
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
13 |
| Similarity score: |
0.514048 |
Alignment:
-TGMYCTTTGBCCK
TGAMCTTTGMMCYT
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
13 |
| Similarity score: |
0.569869 |
Alignment:
VDBHMAGGTCACCCTGACCY-
-------TGAMCTTTGMMCYT
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
1.0303 |
Alignment:
--VBBYCAAGGTCA
AKGYYCAAAGRTCA
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 3 |
Motif name: ESR1 |
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
Best Matches for Motif ID 3 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
20 |
| Similarity score: |
0.0722586 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
---VDBHMAGGTCACCCTGACCY-
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
20 |
| Similarity score: |
0.0818971 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
--VDBHMAGGTCACCCTGACCY---
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
19 |
| Similarity score: |
0.566376 |
Alignment:
-TGTGGTGTCACAGTGCTCC
VDBHMAGGTCACCCTGACCY
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
19 |
| Similarity score: |
0.579928 |
Alignment:
-BMSMGCCYMCTKSTGGMHM
VDBHMAGGTCACCCTGACCY
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
17 |
| Similarity score: |
1.50287 |
Alignment:
---VHRGGTCABDBTGMCCTB
VDBHMAGGTCACCCTGACCY-
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
16 |
| Similarity score: |
2.07409 |
Alignment:
TTCAGCACCATGGACAGCKCC----
-----VDBHMAGGTCACCCTGACCY
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
15 |
| Similarity score: |
2.57517 |
Alignment:
ACTGTGGTGTCACART-----
-VDBHMAGGTCACCCTGACCY
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
15 |
| Similarity score: |
2.57594 |
Alignment:
-----ACTGTGACACCACAGTGGAGCACT
MGGTCAGGGTGACCTRDBHV---------
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
14 |
| Similarity score: |
3.07055 |
Alignment:
------BTRGGDCARAGGKCA
MGGTCAGGGTGACCTRDBHV-
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
13 |
| Similarity score: |
3.5678 |
Alignment:
AAAGGTCAAAGGTCAAC-------
----VDBHMAGGTCACCCTGACCY
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 4 |
Motif name: Esrrb |
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
Best Matches for Motif ID 4 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
0.0442611 |
Alignment:
TGAMCTTTGMMCYT
--TGACCTTGMBBB
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
12 |
| Similarity score: |
0.04699 |
Alignment:
BTRGGDCARAGGKCA
---VBBYCAAGGTCA
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
0.0470808 |
Alignment:
RGGBCAAAGKYCA
-VBBYCAAGGTCA
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
12 |
| Similarity score: |
0.0506977 |
Alignment:
AAAGGTCAAAGGTCAAC
---VBBYCAAGGTCA--
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
12 |
| Similarity score: |
0.0628484 |
Alignment:
VDBHMAGGTCACCCTGACCY
TGACCTTGMBBB--------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0694613 |
Alignment:
VHRGGTCABDBTGMCCTB
-----TGACCTTGMBBB-
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
12 |
| Similarity score: |
0.0713236 |
Alignment:
TTCAGCACCATGGACAGCKCC
----VBBYCAAGGTCA-----
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
0.0772556 |
Alignment:
TGTGGTGTCACAGTGCTCC
-------TGACCTTGMBBB
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
12 |
| Number of overlap: |
12 |
| Similarity score: |
0.0785385 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
-----------TGACCTTGMBBB-
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
11 |
| Number of overlap: |
12 |
| Similarity score: |
0.0832722 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
---VBBYCAAGGTCA----------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 5 |
Motif name: ZEB1 |
| Original motif Consensus sequence: CACCTD |
Reverse complement motif Consensus sequence: HAGGTG |
|
|
 |
 |
Best Matches for Motif ID 5 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
6 |
| Similarity score: |
0 |
Alignment:
YDRCCASYAGRKGGCRSYV
-------HAGGTG------
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
12 |
| Motif name: |
RORA_1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
6 |
| Similarity score: |
0.00688881 |
Alignment:
TGACCTDVWT
-CACCTD---
| Original motif Consensus sequence: AWVDAGGTCA |
Reverse complement motif Consensus sequence: TGACCTDVWT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
6 |
| Similarity score: |
0.00707323 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
----------HAGGTG-----
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
6 |
| Similarity score: |
0.013046 |
Alignment:
VBBYCAAGGTCA
-HAGGTG-----
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
6 |
| Similarity score: |
0.0160352 |
Alignment:
GTTGACCTTTGACCTTT
----------CACCTD-
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
6 |
| Similarity score: |
0.0192435 |
Alignment:
TGRCCTKTGHCCKAB
-CACCTD--------
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
11 |
| Motif name: |
NR4A2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
6 |
| Similarity score: |
0.0216496 |
Alignment:
GTGACCTT
--CACCTD
| Original motif Consensus sequence: AAGGTCAC |
Reverse complement motif Consensus sequence: GTGACCTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
6 |
| Similarity score: |
0.022947 |
Alignment:
MGGTCAGGGTGACCTRDBHV
----CACCTD----------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
6 |
| Similarity score: |
0.0246463 |
Alignment:
HSCACGTGGC
--CACCTD--
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
6 |
| Similarity score: |
0.025129 |
Alignment:
DCCACGTGCV
--HAGGTG--
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 6 |
Motif name: Mycn |
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
Best Matches for Motif ID 6 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0 |
Alignment:
DCCACGTGCV
GCCACGTGSD
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0654819 |
Alignment:
BSCCCCGCGBS
-GCCACGTGSD
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.0679017 |
Alignment:
YDRCCASYAGRKGGCRSYV
-----HSCACGTGGC----
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0704988 |
Alignment:
BBSGGCGGGGBS
-HSCACGTGGC-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
10 |
| Similarity score: |
0.0883224 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
------GCCACGTGSD---------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.089863 |
Alignment:
MGGTCAGGGTGACCTRDBHV
--------GCCACGTGSD--
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0943014 |
Alignment:
ABCACAGTGGAGCACTG
-GCCACGTGSD------
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0947124 |
Alignment:
TGMYCTTTGBCCK
-GCCACGTGSD--
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.0949359 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
---------GCCACGTGSD-----
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
10 |
| Similarity score: |
0.0954444 |
Alignment:
TGRCCTKTGHCCKAB
-GCCACGTGSD----
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 7 |
Motif name: REST |
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
Best Matches for Motif ID 7 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
19 |
| Similarity score: |
0.0605225 |
Alignment:
--AGTGCTCCACTGTGGTGTCACAGT
GGYGCTGTCCATGGTGCTGAA-----
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
16 |
| Similarity score: |
1.56264 |
Alignment:
-----VDBHMAGGTCACCCTGACCY
TTCAGCACCATGGACAGCKCC----
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
14 |
| Similarity score: |
2.57032 |
Alignment:
-------ABCACAGTGGAGCACTG
GGYGCTGTCCATGGTGCTGAA---
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
14 |
| Similarity score: |
2.57054 |
Alignment:
ACTGTGGTGTCACART-------
--GGYGCTGTCCATGGTGCTGAA
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
12 |
| Number of overlap: |
13 |
| Similarity score: |
3.04841 |
Alignment:
TGGAGCACTGTGACAVCACAGTGG--------
-----------TTCAGCACCATGGACAGCKCC
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
13 |
| Similarity score: |
3.05156 |
Alignment:
BAGGYCABHBTGACCKHV--------
-----GGYGCTGTCCATGGTGCTGAA
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
13 |
| Similarity score: |
3.06375 |
Alignment:
TGMYCTTTGBCCK--------
GGYGCTGTCCATGGTGCTGAA
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
3.06574 |
Alignment:
--------BTRGGDCARAGGKCA
TTCAGCACCATGGACAGCKCC--
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
14 |
| Number of overlap: |
12 |
| Similarity score: |
3.55903 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG---------
-------------TTCAGCACCATGGACAGCKCC
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
3.56306 |
Alignment:
VDGACTRTGGDD---------
GGYGCTGTCCATGGTGCTGAA
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 8 |
Motif name: Myc |
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
Best Matches for Motif ID 8 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0 |
Alignment:
HSCACGTGGC
VGCACGTGGH
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.064853 |
Alignment:
BSCCCCGCGBS
-DCCACGTGCV
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0728588 |
Alignment:
SBCCCCGCCSBB
-DCCACGTGCV-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
10 |
| Similarity score: |
0.0754257 |
Alignment:
BMSMGCCYMCTKSTGGMHM
-------VGCACGTGGH--
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.0888909 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
----------DCCACGTGCV-----
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0934488 |
Alignment:
TGMYCTTTGBCCK
---VGCACGTGGH
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.0940438 |
Alignment:
MGGTCAGGGTGACCTRDBHV
--------DCCACGTGCV--
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0955234 |
Alignment:
ABCACAGTGGAGCACTG
-DCCACGTGCV------
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
10 |
| Similarity score: |
0.0968598 |
Alignment:
VHRGGTCABDBTGMCCTB
----VGCACGTGGH----
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
11 |
| Number of overlap: |
10 |
| Similarity score: |
0.0971125 |
Alignment:
TGGAGCACTGTGACAVCACAGTGG
----DCCACGTGCV----------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 9 |
Motif name: ESR2 |
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
Best Matches for Motif ID 9 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
18 |
| Similarity score: |
0.0717757 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
------VHRGGTCABDBTGMCCTB
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
18 |
| Similarity score: |
0.0815903 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
-VHRGGTCABDBTGMCCTB------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
18 |
| Similarity score: |
0.0916961 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
-VHRGGTCABDBTGMCCTB--
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
17 |
| Similarity score: |
0.511812 |
Alignment:
VDBHMAGGTCACCCTGACCY-
---VHRGGTCABDBTGMCCTB
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
17 |
| Similarity score: |
0.569595 |
Alignment:
TGTGGTGTCACAGTGCTCC-
--VHRGGTCABDBTGMCCTB
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
15 |
| Similarity score: |
1.56699 |
Alignment:
BTRGGDCARAGGKCA---
VHRGGTCABDBTGMCCTB
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
14 |
| Similarity score: |
2.08921 |
Alignment:
----TGAMCTTTGMMCYT
VHRGGTCABDBTGMCCTB
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
15 |
| Motif name: |
Tcfcp2l1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
14 |
| Similarity score: |
2.09109 |
Alignment:
----CCAGYYHVADCCRG
BAGGYCABHBTGACCKHV
| Original motif Consensus sequence: CCAGYYHVADCCRG |
Reverse complement motif Consensus sequence: CKGGDTBDMMCTGG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
13 |
| Similarity score: |
2.58379 |
Alignment:
YDRCCASYAGRKGGCRSYV-----
------VHRGGTCABDBTGMCCTB
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
13 |
| Similarity score: |
2.59263 |
Alignment:
-----RGGBCAAAGKYCA
BAGGYCABHBTGACCKHV
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 10 |
Motif name: NR1H2RXRA |
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
Best Matches for Motif ID 10 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
17 |
| Similarity score: |
0.0911991 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
---GTTGACCTTTGACCTTT-
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
15 |
| Similarity score: |
1.05945 |
Alignment:
BTRGGDCARAGGKCA--
AAAGGTCAAAGGTCAAC
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
8 |
| Number of overlap: |
13 |
| Similarity score: |
2.08514 |
Alignment:
----VDBHMAGGTCACCCTGACCY
AAAGGTCAAAGGTCAAC-------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
13 |
| Number of overlap: |
12 |
| Similarity score: |
2.59961 |
Alignment:
AGTGCTCCACTGTGGTGTCACAGT-----
------------AAAGGTCAAAGGTCAAC
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
12 |
| Similarity score: |
2.59967 |
Alignment:
ACTGTGGTGTCACART-----
----AAAGGTCAAAGGTCAAC
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
11 |
| Similarity score: |
3.09731 |
Alignment:
------CAGTGCTCCACTGTGBT
AAAGGTCAAAGGTCAAC------
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
9 |
| Number of overlap: |
10 |
| Similarity score: |
3.56221 |
Alignment:
BAGGYCABHBTGACCKHV-------
--------GTTGACCTTTGACCTTT
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
13 |
| Motif name: |
MIZF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
3.5855 |
Alignment:
-------BAACGTCCGC
AAAGGTCAAAGGTCAAC
| Original motif Consensus sequence: BAACGTCCGC |
Reverse complement motif Consensus sequence: GCGGACGTTV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
9 |
| Similarity score: |
4.07454 |
Alignment:
AKGYYCAAAGRTCA--------
-----AAAGGTCAAAGGTCAAC
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
16 |
| Number of overlap: |
9 |
| Similarity score: |
4.09479 |
Alignment:
--------TGGAGCACTGTGACAVCACAGTGG
GTTGACCTTTGACCTTT---------------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 11 |
Motif name: NR4A2 |
| Original motif Consensus sequence: AAGGTCAC |
Reverse complement motif Consensus sequence: GTGACCTT |
|
|
 |
 |
Best Matches for Motif ID 11 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
8 |
| Similarity score: |
0.0167759 |
Alignment:
AAAGGTCAAAGGTCAAC
-AAGGTCAC--------
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
8 |
| Similarity score: |
0.0195235 |
Alignment:
VDBHMAGGTCACCCTGACCY
----AAGGTCAC--------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
8 |
| Similarity score: |
0.0198372 |
Alignment:
BAGGYCABHBTGACCKHV
---------GTGACCTT-
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
8 |
| Similarity score: |
0.0495607 |
Alignment:
BTRGGDCARAGGKCA
-AAGGTCAC------
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
8 |
| Similarity score: |
0.0557243 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
------AAGGTCAC-----------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
8 |
| Similarity score: |
0.0564874 |
Alignment:
AKGYYCAAAGRTCA
------AAGGTCAC
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
8 |
| Similarity score: |
0.0594649 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
----GTGACCTT---------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
8 |
| Similarity score: |
0.060729 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
-------AAGGTCAC---------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
0.0645146 |
Alignment:
ACTGTGGTGTCACART
-----AAGGTCAC---
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
8 |
| Similarity score: |
0.0695512 |
Alignment:
TGTGGTGTCACAGTGCTCC
---AAGGTCAC--------
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 12 |
Motif name: RORA_1 |
| Original motif Consensus sequence: AWVDAGGTCA |
Reverse complement motif Consensus sequence: TGACCTDVWT |
|
|
 |
 |
Best Matches for Motif ID 12 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.029401 |
Alignment:
VBBYCAAGGTCA
--AWVDAGGTCA
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0417851 |
Alignment:
MGGTCAGGGTGACCTRDBHV
---------TGACCTDVWT-
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
10 |
| Similarity score: |
0.05425 |
Alignment:
AAAGGTCAAAGGTCAAC
-----AWVDAGGTCA--
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
10 |
| Similarity score: |
0.0654808 |
Alignment:
TGAMCTTTGMMCYT
----TGACCTDVWT
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.06668 |
Alignment:
BTRGGDCARAGGKCA
-----AWVDAGGTCA
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0755608 |
Alignment:
TTCAGCACCATGGACAGCKCC
------AWVDAGGTCA-----
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
10 |
| Similarity score: |
0.0803396 |
Alignment:
RGGBCAAAGKYCA
---AWVDAGGTCA
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0889306 |
Alignment:
GGAGCACTGTGACACCACA
---------TGACCTDVWT
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.09225 |
Alignment:
DDCCAMAGTCHB
-AWVDAGGTCA-
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
10 |
| Similarity score: |
0.0988214 |
Alignment:
BAGGYCABHBTGACCKHV
-----AWVDAGGTCA---
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 13 |
Motif name: MIZF |
| Original motif Consensus sequence: BAACGTCCGC |
Reverse complement motif Consensus sequence: GCGGACGTTV |
|
|
 |
 |
Best Matches for Motif ID 13 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0605777 |
Alignment:
BAGGYCABHBTGACCKHV
--------GCGGACGTTV
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
8 |
| Number of overlap: |
10 |
| Similarity score: |
0.0614567 |
Alignment:
VDBHMAGGTCACCCTGACCY
---BAACGTCCGC-------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0646071 |
Alignment:
AAAGGTCAAAGGTCAAC
-------BAACGTCCGC
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0651598 |
Alignment:
GCGCGCSCGCG
BAACGTCCGC-
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
10 |
| Similarity score: |
0.0697767 |
Alignment:
BMSMGCCYMCTKSTGGMHM
BAACGTCCGC---------
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.0708149 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
---------BAACGTCCGC--
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
19 |
| Motif name: |
atactttggc |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.0803571 |
Alignment:
ATACTTTGGC
BAACGTCCGC
| Original motif Consensus sequence: ATACTTTGGC |
Reverse complement motif Consensus sequence: GCCAAAGTAT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
0.555645 |
Alignment:
VGCACGTGGH-
-GCGGACGTTV
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
9 |
| Similarity score: |
0.560913 |
Alignment:
CCGCGGRCACG-
--GCGGACGTTV
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
0.561179 |
Alignment:
HSCACGTGGC-
-GCGGACGTTV
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 14 |
Motif name: PPARGRXRA |
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
Best Matches for Motif ID 14 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
15 |
| Similarity score: |
0.0442974 |
Alignment:
AAAGGTCAAAGGTCAAC
BTRGGDCARAGGKCA--
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
15 |
| Similarity score: |
0.0602408 |
Alignment:
VHRGGTCABDBTGMCCTB
BTRGGDCARAGGKCA---
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
15 |
| Similarity score: |
0.0723092 |
Alignment:
VDBHMAGGTCACCCTGACCY
---BTRGGDCARAGGKCA--
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
15 |
| Similarity score: |
0.0741352 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
-----BTRGGDCARAGGKCA-----
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
15 |
| Similarity score: |
0.0786357 |
Alignment:
YDRCCASYAGRKGGCRSYV
-BTRGGDCARAGGKCA---
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
15 |
| Similarity score: |
0.0796159 |
Alignment:
TTCAGCACCATGGACAGCKCC
-BTRGGDCARAGGKCA-----
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
15 |
| Similarity score: |
0.0813414 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
------BTRGGDCARAGGKCA---
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
15 |
| Similarity score: |
0.0818655 |
Alignment:
TGTGGTGTCACAGTGCTCC
--BTRGGDCARAGGKCA--
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
14 |
| Similarity score: |
0.536521 |
Alignment:
AKGYYCAAAGRTCA-
BTRGGDCARAGGKCA
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
15 |
| Motif name: |
Tcfcp2l1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
14 |
| Similarity score: |
0.584465 |
Alignment:
-CKGGDTBDMMCTGG
TGRCCTKTGHCCKAB
| Original motif Consensus sequence: CCAGYYHVADCCRG |
Reverse complement motif Consensus sequence: CKGGDTBDMMCTGG |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 15 |
Motif name: Tcfcp2l1 |
| Original motif Consensus sequence: CCAGYYHVADCCRG |
Reverse complement motif Consensus sequence: CKGGDTBDMMCTGG |
|
|
 |
 |
Best Matches for Motif ID 15 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
14 |
| Similarity score: |
0.0453398 |
Alignment:
MGGTCAGGGTGACCTRDBHV
---CCAGYYHVADCCRG---
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
14 |
| Similarity score: |
0.0491111 |
Alignment:
YDRCCASYAGRKGGCRSYV
---CCAGYYHVADCCRG--
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
14 |
| Similarity score: |
0.0568174 |
Alignment:
VHRGGTCABDBTGMCCTB
----CKGGDTBDMMCTGG
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
14 |
| Similarity score: |
0.0569399 |
Alignment:
TGRCCTKTGHCCKAB
-CKGGDTBDMMCTGG
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
13 |
| Similarity score: |
0.551995 |
Alignment:
ABCACAGTGGAGCACTG-
----CKGGDTBDMMCTGG
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
14 |
| Number of overlap: |
12 |
| Similarity score: |
1.05046 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT--
-------------CKGGDTBDMMCTGG
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
12 |
| Similarity score: |
1.05111 |
Alignment:
--TTCAGCACCATGGACAGCKCC
CKGGDTBDMMCTGG---------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
1.05387 |
Alignment:
TGMYCTTTGBCCK--
-CKGGDTBDMMCTGG
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
1.05541 |
Alignment:
CAHCACAGTGGAGCACT--
-----CKGGDTBDMMCTGG
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
11 |
| Similarity score: |
1.55702 |
Alignment:
---AKGYYCAAAGRTCA
CKGGDTBDMMCTGG---
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: 1 |
Motif ID: 16 |
Motif name: CTCF |
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
Best Matches for Motif ID 16 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
19 |
| Similarity score: |
0.0655511 |
Alignment:
MGGTCAGGGTGACCTRDBHV
-YDRCCASYAGRKGGCRSYV
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
18 |
| Similarity score: |
0.563463 |
Alignment:
ACTGTGACACCACAGTGGAGCACT-
------YDRCCASYAGRKGGCRSYV
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
8 |
| Number of overlap: |
18 |
| Similarity score: |
0.566264 |
Alignment:
-AGTGSWSCACTGTGAMAMCACAGTG
BMSMGCCYMCTKSTGGMHM-------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
17 |
| Similarity score: |
1.05236 |
Alignment:
--ABCACAGTGGAGCACTG
YDRCCASYAGRKGGCRSYV
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
15 |
| Similarity score: |
2.06184 |
Alignment:
----CAHCACAGTGGAGCACT
YDRCCASYAGRKGGCRSYV--
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
10 |
| Number of overlap: |
15 |
| Similarity score: |
2.06698 |
Alignment:
TGGAGCACTGTGACAVCACAGTGG----
---------YDRCCASYAGRKGGCRSYV
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
15 |
| Motif name: |
Tcfcp2l1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
13 |
| Similarity score: |
3.05613 |
Alignment:
------CCAGYYHVADCCRG
YDRCCASYAGRKGGCRSYV-
| Original motif Consensus sequence: CCAGYYHVADCCRG |
Reverse complement motif Consensus sequence: CKGGDTBDMMCTGG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
13 |
| Similarity score: |
3.06048 |
Alignment:
------VHRGGTCABDBTGMCCTB
YDRCCASYAGRKGGCRSYV-----
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
13 |
| Similarity score: |
3.06565 |
Alignment:
------BTRGGDCARAGGKCA
YDRCCASYAGRKGGCRSYV--
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
3.56185 |
Alignment:
SBCCCCGCCSBB-------
BMSMGCCYMCTKSTGGMHM
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 17 |
Motif name: TgTGgTGTCACAGTGCTCC |
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
Best Matches for Motif ID 17 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
5 |
| Number of overlap: |
19 |
| Similarity score: |
0 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
----TGTGGTGTCACAGTGCTCC-
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
19 |
| Similarity score: |
0.0217259 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
---TGTGGTGTCACAGTGCTCC---
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
19 |
| Similarity score: |
0.074816 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
--TGTGGTGTCACAGTGCTCC---
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
19 |
| Similarity score: |
0.0857262 |
Alignment:
VDBHMAGGTCACCCTGACCY
-TGTGGTGTCACAGTGCTCC
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
17 |
| Similarity score: |
1.08001 |
Alignment:
--VHRGGTCABDBTGMCCTB
TGTGGTGTCACAGTGCTCC-
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
17 |
| Similarity score: |
1.08847 |
Alignment:
CAGTGCTCCACTGTGBT--
TGTGGTGTCACAGTGCTCC
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
16 |
| Similarity score: |
1.59452 |
Alignment:
ACTGTGGTGTCACART---
GGAGCACTGTGACACCACA
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
15 |
| Similarity score: |
2.10007 |
Alignment:
BTRGGDCARAGGKCA----
GGAGCACTGTGACACCACA
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
14 |
| Similarity score: |
2.57554 |
Alignment:
-----CAHCACAGTGGAGCACT
TGTGGTGTCACAGTGCTCC---
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
3.58914 |
Alignment:
-------VBBYCAAGGTCA
GGAGCACTGTGACACCACA
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 18 |
Motif name: sscCCCGCGcs |
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
Best Matches for Motif ID 18 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0 |
Alignment:
SBCCCCGCCSBB
BSCCCCGCGBS-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0417227 |
Alignment:
CGCGSGCGCGC
SBCGCGGGGSB
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
28 |
| Motif name: |
ssGCkTGCssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0523123 |
Alignment:
SSGCKTGCSSK
BSCCCCGCGBS
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
11 |
| Similarity score: |
0.0564004 |
Alignment:
YDRCCASYAGRKGGCRSYV
--SBCGCGGGGSB------
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0734979 |
Alignment:
MGGTCAGGGTGACCTRDBHV
BSCCCCGCGBS---------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0757081 |
Alignment:
CCGCGGRCACG
BSCCCCGCGBS
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
11 |
| Number of overlap: |
11 |
| Similarity score: |
0.0758097 |
Alignment:
TTCAGCACCATGGACAGCKCC
----------BSCCCCGCGBS
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.535606 |
Alignment:
-VGCACGTGGH
SBCGCGGGGSB
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.536235 |
Alignment:
-HSCACGTGGC
SBCGCGGGGSB
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
33 |
| Motif name: |
srACyCCGAyr |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.571697 |
Alignment:
KKTCGGKGTKS-
-SBCGCGGGGSB
| Original motif Consensus sequence: SRACYCCGAYR |
Reverse complement motif Consensus sequence: KKTCGGKGTKS |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 19 |
Motif name: atactttggc |
| Original motif Consensus sequence: ATACTTTGGC |
Reverse complement motif Consensus sequence: GCCAAAGTAT |
|
|
 |
 |
Best Matches for Motif ID 19 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0215909 |
Alignment:
DDCCAMAGTCHB
-GCCAAAGTAT-
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
30 |
| Motif name: |
taaacgatgcc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0375 |
Alignment:
GGCATCGTTTA
GCCAAAGTAT-
| Original motif Consensus sequence: TAAACGATGCC |
Reverse complement motif Consensus sequence: GGCATCGTTTA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
10 |
| Similarity score: |
0.0395 |
Alignment:
GTTGACCTTTGACCTTT
----ATACTTTGGC---
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
10 |
| Similarity score: |
0.0472222 |
Alignment:
GGAGCACTGTGACACCACA
------ATACTTTGGC---
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
10 |
| Number of overlap: |
10 |
| Similarity score: |
0.0474206 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
---------GCCAAAGTAT------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.0483209 |
Alignment:
TGMYCTTTGBCCK
--ATACTTTGGC-
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.0490385 |
Alignment:
TGAMCTTTGMMCYT
-ATACTTTGGC---
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
10 |
| Similarity score: |
0.0493243 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
----------GCCAAAGTAT----
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
9 |
| Number of overlap: |
10 |
| Similarity score: |
0.0498239 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
--------GCCAAAGTAT------
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
10 |
| Similarity score: |
0.0527344 |
Alignment:
AGTGCTCCACTGTGDTG
------ATACTTTGGC-
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 20 |
Motif name: CagTGCTCCACTGTGgT |
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
Best Matches for Motif ID 20 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
17 |
| Similarity score: |
0.0957051 |
Alignment:
BMSMGCCYMCTKSTGGMHM
--CAGTGCTCCACTGTGBT
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
17 |
| Similarity score: |
0.0975871 |
Alignment:
TTCAGCACCATGGACAGCKCC
-ABCACAGTGGAGCACTG---
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
17 |
| Similarity score: |
0.0980868 |
Alignment:
TGTGGTGTCACAGTGCTCC
CAGTGCTCCACTGTGBT--
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
17 |
| Similarity score: |
0.099784 |
Alignment:
AGTGCTCCACTGTGGTGTCACAGT
-----ABCACAGTGGAGCACTG--
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
17 |
| Similarity score: |
0.104226 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
-----ABCACAGTGGAGCACTG---
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
17 |
| Similarity score: |
0.107548 |
Alignment:
TGGAGCACTGTGACAVCACAGTGG
---ABCACAGTGGAGCACTG----
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
16 |
| Similarity score: |
0.5 |
Alignment:
CAHCACAGTGGAGCACT-
-ABCACAGTGGAGCACTG
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
16 |
| Similarity score: |
0.607943 |
Alignment:
-AMTGTGACACCACAGT
CAGTGCTCCACTGTGBT
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
15 |
| Motif name: |
Tcfcp2l1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
13 |
| Similarity score: |
2.1063 |
Alignment:
----CKGGDTBDMMCTGG
ABCACAGTGGAGCACTG-
| Original motif Consensus sequence: CCAGYYHVADCCRG |
Reverse complement motif Consensus sequence: CKGGDTBDMMCTGG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
2.58308 |
Alignment:
VDGACTRTGGDD-----
ABCACAGTGGAGCACTG
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 21 |
Motif name: AmTGTGACACCACAGT |
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
Best Matches for Motif ID 21 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
16 |
| Similarity score: |
0 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
AMTGTGACACCACAGT--------
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
16 |
| Similarity score: |
0.00171837 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
--ACTGTGGTGTCACART------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
16 |
| Similarity score: |
0.0197279 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
-ACTGTGGTGTCACART--------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
16 |
| Similarity score: |
0.0950937 |
Alignment:
GGAGCACTGTGACACCACA
ACTGTGGTGTCACART---
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
16 |
| Similarity score: |
0.0988969 |
Alignment:
CAGTGCTCCACTGTGBT
-AMTGTGACACCACAGT
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
16 |
| Similarity score: |
0.101875 |
Alignment:
AGTGCTCCACTGTGDTG
AMTGTGACACCACAGT-
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
15 |
| Similarity score: |
0.595092 |
Alignment:
-MGGTCAGGGTGACCTRDBHV
AMTGTGACACCACAGT-----
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
8 |
| Number of overlap: |
14 |
| Similarity score: |
1.1019 |
Alignment:
--GGYGCTGTCCATGGTGCTGAA
ACTGTGGTGTCACART-------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
12 |
| Similarity score: |
2.07799 |
Alignment:
----BAGGYCABHBTGACCKHV
AMTGTGACACCACAGT------
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
2.10225 |
Alignment:
----GTTGACCTTTGACCTTT
AMTGTGACACCACAGT-----
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 22 |
Motif name: CACTGTGrYrtCACAGTGswsCAcT |
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
Best Matches for Motif ID 22 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
24 |
| Similarity score: |
0.0247168 |
Alignment:
-ACTGTGACACCACAGTGGAGCACT
CACTGTGRTRTCACAGTGSWSCACT
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
23 |
| Similarity score: |
0.500966 |
Alignment:
--TGGAGCACTGTGACAVCACAGTGG
AGTGSWSCACTGTGAMAMCACAGTG-
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
18 |
| Similarity score: |
3.08246 |
Alignment:
MGGTCAGGGTGACCTRDBHV-------
--CACTGTGRTRTCACAGTGSWSCACT
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
18 |
| Similarity score: |
3.08842 |
Alignment:
BMSMGCCYMCTKSTGGMHM-------
-AGTGSWSCACTGTGAMAMCACAGTG
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
18 |
| Similarity score: |
3.08984 |
Alignment:
GGAGCACTGTGACACCACA-------
-CACTGTGRTRTCACAGTGSWSCACT
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
17 |
| Similarity score: |
3.52449 |
Alignment:
--------CAHCACAGTGGAGCACT
CACTGTGRTRTCACAGTGSWSCACT
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
16 |
| Similarity score: |
4.02265 |
Alignment:
---------CAGTGCTCCACTGTGBT
AGTGSWSCACTGTGAMAMCACAGTG-
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
13 |
| Similarity score: |
5.58301 |
Alignment:
VHRGGTCABDBTGMCCTB------------
-----AGTGSWSCACTGTGAMAMCACAGTG
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
13 |
| Similarity score: |
5.58597 |
Alignment:
ACTGTGGTGTCACART------------
---CACTGTGRTRTCACAGTGSWSCACT
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
12 |
| Similarity score: |
6.07826 |
Alignment:
-------------GGYGCTGTCCATGGTGCTGAA
CACTGTGRTRTCACAGTGSWSCACT---------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 23 |
Motif name: TGGAGCACTGTGACAcCACAGTGg |
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
Best Matches for Motif ID 23 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
23 |
| Similarity score: |
0.0136536 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT-
--CCACTGTGVTGTCACAGTGCTCCA
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
22 |
| Similarity score: |
0.560082 |
Alignment:
--ACTGTGACACCACAGTGGAGCACT
CCACTGTGVTGTCACAGTGCTCCA--
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
18 |
| Similarity score: |
2.5833 |
Alignment:
------BAGGYCABHBTGACCKHV
TGGAGCACTGTGACAVCACAGTGG
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
17 |
| Similarity score: |
3.10239 |
Alignment:
MGGTCAGGGTGACCTRDBHV-------
---CCACTGTGVTGTCACAGTGCTCCA
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
16 |
| Similarity score: |
3.56015 |
Alignment:
GGAGCACTGTGACACCACA--------
---CCACTGTGVTGTCACAGTGCTCCA
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
16 |
| Similarity score: |
3.58845 |
Alignment:
ABCACAGTGGAGCACTG--------
-CCACTGTGVTGTCACAGTGCTCCA
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
15 |
| Similarity score: |
4.07113 |
Alignment:
AGTGCTCCACTGTGDTG---------
--TGGAGCACTGTGACAVCACAGTGG
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
15 |
| Similarity score: |
4.10183 |
Alignment:
---------YDRCCASYAGRKGGCRSYV
TGGAGCACTGTGACAVCACAGTGG----
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
14 |
| Similarity score: |
4.60325 |
Alignment:
BTRGGDCARAGGKCA----------
-TGGAGCACTGTGACAVCACAGTGG
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
13 |
| Similarity score: |
5.08033 |
Alignment:
-----------TTCAGCACCATGGACAGCKCC
TGGAGCACTGTGACAVCACAGTGG--------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 24 |
Motif name: ssCCCCGCSssk |
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
Best Matches for Motif ID 24 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.063076 |
Alignment:
MGGTCAGGGTGACCTRDBHV
-------BBSGGCGGGGBS-
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.0642087 |
Alignment:
YDRCCASYAGRKGGCRSYV
-BBSGGCGGGGBS------
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.0671548 |
Alignment:
TGRCCTKTGHCCKAB
-SBCCCCGCCSBB--
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.0734481 |
Alignment:
BAGGYCABHBTGACCKHV
-SBCCCCGCCSBB-----
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
12 |
| Similarity score: |
0.0751084 |
Alignment:
TTCAGCACCATGGACAGCKCC
BBSGGCGGGGBS---------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.5 |
Alignment:
BSCCCCGCGBS-
SBCCCCGCCSBB
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.548902 |
Alignment:
-CGCGSGCGCGC
BBSGGCGGGGBS
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
28 |
| Motif name: |
ssGCkTGCssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.550852 |
Alignment:
SSGCKTGCSSK-
SBCCCCGCCSBB
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.56634 |
Alignment:
CGTGKCCGCGG-
BBSGGCGGGGBS
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.571364 |
Alignment:
VBBYCAAGGTCA-
-BBSGGCGGGGBS
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 25 |
Motif name: wwCCAmAGTCmt |
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
Best Matches for Motif ID 25 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0263648 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
--------DDCCAMAGTCHB-----
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0282828 |
Alignment:
ABCACAGTGGAGCACTG
VDGACTRTGGDD-----
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0291095 |
Alignment:
AGTGCTCCACTGTGDTG
-----VDGACTRTGGDD
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
12 |
| Similarity score: |
0.0301919 |
Alignment:
AGTGCTCCACTGTGGTGTCACAGT
-----VDGACTRTGGDD-------
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.032442 |
Alignment:
RGGBCAAAGKYCA
-DDCCAMAGTCHB
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
12 |
| Similarity score: |
0.0360702 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
---DDCCAMAGTCHB---------
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
12 |
| Similarity score: |
0.0391228 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
---DDCCAMAGTCHB------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
12 |
| Similarity score: |
0.0393782 |
Alignment:
TGTGGTGTCACAGTGCTCC
-----DDCCAMAGTCHB--
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
12 |
| Similarity score: |
0.0407304 |
Alignment:
VDBHMAGGTCACCCTGACCY
DDCCAMAGTCHB--------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
0.0463384 |
Alignment:
AAAGGTCAAAGGTCAAC
----DDCCAMAGTCHB-
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 26 |
Motif name: CAcCACAGTGGAGCAct |
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
Best Matches for Motif ID 26 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
17 |
| Similarity score: |
0.00742502 |
Alignment:
ACTGTGACACCACAGTGGAGCACT
-------CAHCACAGTGGAGCACT
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
9 |
| Number of overlap: |
17 |
| Similarity score: |
0.0456803 |
Alignment:
CACTGTGRTRTCACAGTGSWSCACT
--------CAHCACAGTGGAGCACT
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
17 |
| Similarity score: |
0.0894884 |
Alignment:
TTCAGCACCATGGACAGCKCC
--CAHCACAGTGGAGCACT--
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
17 |
| Similarity score: |
0.0962536 |
Alignment:
GGAGCACTGTGACACCACA
-CAHCACAGTGGAGCACT-
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
17 |
| Similarity score: |
0.0978126 |
Alignment:
YDRCCASYAGRKGGCRSYV
-CAHCACAGTGGAGCACT-
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
17 |
| Similarity score: |
0.106708 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
-----AGTGCTCCACTGTGDTG--
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
16 |
| Similarity score: |
0.5 |
Alignment:
ABCACAGTGGAGCACTG-
-CAHCACAGTGGAGCACT
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
16 |
| Similarity score: |
0.610921 |
Alignment:
ACTGTGGTGTCACART-
CAHCACAGTGGAGCACT
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
12 |
| Similarity score: |
2.5839 |
Alignment:
-----VDGACTRTGGDD
AGTGCTCCACTGTGDTG
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
12 |
| Similarity score: |
2.60332 |
Alignment:
-----TGMYCTTTGBCCK
AGTGCTCCACTGTGDTG-
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 27 |
Motif name: AgTGCTCCACTGTGgTGTCACAgT |
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
Best Matches for Motif ID 27 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
24 |
| Similarity score: |
0.0409586 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
-AGTGCTCCACTGTGGTGTCACAGT
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
22 |
| Similarity score: |
1.06364 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA--
--ACTGTGACACCACAGTGGAGCACT
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
19 |
| Similarity score: |
2.59599 |
Alignment:
TTCAGCACCATGGACAGCKCC-----
--ACTGTGACACCACAGTGGAGCACT
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
18 |
| Similarity score: |
3.10123 |
Alignment:
------GGAGCACTGTGACACCACA
ACTGTGACACCACAGTGGAGCACT-
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
18 |
| Similarity score: |
3.10186 |
Alignment:
------BMSMGCCYMCTKSTGGMHM
AGTGCTCCACTGTGGTGTCACAGT-
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
26 |
| Motif name: |
CAcCACAGTGGAGCAct |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
17 |
| Similarity score: |
3.50248 |
Alignment:
CAHCACAGTGGAGCACT-------
ACTGTGACACCACAGTGGAGCACT
| Original motif Consensus sequence: CAHCACAGTGGAGCACT |
Reverse complement motif Consensus sequence: AGTGCTCCACTGTGDTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
16 |
| Similarity score: |
4 |
Alignment:
--------ABCACAGTGGAGCACTG
ACTGTGACACCACAGTGGAGCACT-
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
16 |
| Similarity score: |
4.0041 |
Alignment:
ACTGTGGTGTCACART--------
AGTGCTCCACTGTGGTGTCACAGT
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
15 |
| Similarity score: |
4.59996 |
Alignment:
MGGTCAGGGTGACCTRDBHV---------
-----ACTGTGACACCACAGTGGAGCACT
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
12 |
| Similarity score: |
6.08041 |
Alignment:
------------VHRGGTCABDBTGMCCTB
AGTGCTCCACTGTGGTGTCACAGT------
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 28 |
Motif name: ssGCkTGCssk |
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
Best Matches for Motif ID 28 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.00456539 |
Alignment:
GCGCGCSCGCG
SSGCKTGCSSK
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0176347 |
Alignment:
BBSGGCGGGGBS
-YSSGCAYGCSS
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0190954 |
Alignment:
SBCGCGGGGSB
YSSGCAYGCSS
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0276515 |
Alignment:
CCGCGGRCACG
SSGCKTGCSSK
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
11 |
| Similarity score: |
0.0349439 |
Alignment:
YDRCCASYAGRKGGCRSYV
-------SSGCKTGCSSK-
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
4 |
| Motif name: |
Esrrb |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0358701 |
Alignment:
VBBYCAAGGTCA
-YSSGCAYGCSS
| Original motif Consensus sequence: VBBYCAAGGTCA |
Reverse complement motif Consensus sequence: TGACCTTGMBBB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.0379074 |
Alignment:
VHRGGTCABDBTGMCCTB
-----YSSGCAYGCSS--
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
10 |
| Number of overlap: |
11 |
| Similarity score: |
0.0409322 |
Alignment:
TTCAGCACCATGGACAGCKCC
-YSSGCAYGCSS---------
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.041419 |
Alignment:
BTRGGDCARAGGKCA
--YSSGCAYGCSS--
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
1 |
| Motif name: |
HNF4A |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.0437981 |
Alignment:
RGGBCAAAGKYCA
--YSSGCAYGCSS
| Original motif Consensus sequence: RGGBCAAAGKYCA |
Reverse complement motif Consensus sequence: TGMYCTTTGBCCK |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 29 |
Motif name: cCGCGGrCACG |
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
Best Matches for Motif ID 29 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.010473 |
Alignment:
GCGCGCSCGCG
CCGCGGRCACG
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
11 |
| Similarity score: |
0.0484441 |
Alignment:
TTCAGCACCATGGACAGCKCC
-------CCGCGGRCACG---
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.0486581 |
Alignment:
MGGTCAGGGTGACCTRDBHV
-------CGTGKCCGCGG--
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
28 |
| Motif name: |
ssGCkTGCssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0503056 |
Alignment:
SSGCKTGCSSK
CCGCGGRCACG
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.055777 |
Alignment:
BBSGGCGGGGBS
CGTGKCCGCGG-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
8 |
| Number of overlap: |
11 |
| Similarity score: |
0.0588663 |
Alignment:
YDRCCASYAGRKGGCRSYV
-------CCGCGGRCACG-
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
14 |
| Motif name: |
PPARGRXRA |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0629476 |
Alignment:
TGRCCTKTGHCCKAB
---CGTGKCCGCGG-
| Original motif Consensus sequence: BTRGGDCARAGGKCA |
Reverse complement motif Consensus sequence: TGRCCTKTGHCCKAB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0651451 |
Alignment:
BSCCCCGCGBS
CCGCGGRCACG
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
9 |
| Number of overlap: |
10 |
| Similarity score: |
0.551597 |
Alignment:
BAGGYCABHBTGACCKHV-
--------CGTGKCCGCGG
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
1.0488 |
Alignment:
VGCACGTGGH--
-CGTGKCCGCGG
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 30 |
Motif name: taaacgatgcc |
| Original motif Consensus sequence: TAAACGATGCC |
Reverse complement motif Consensus sequence: GGCATCGTTTA |
|
|
 |
 |
Best Matches for Motif ID 30 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
10 |
| Motif name: |
NR1H2RXRA |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
11 |
| Similarity score: |
0.0511364 |
Alignment:
GTTGACCTTTGACCTTT
----TAAACGATGCC--
| Original motif Consensus sequence: AAAGGTCAAAGGTCAAC |
Reverse complement motif Consensus sequence: GTTGACCTTTGACCTTT |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
2 |
| Motif name: |
NR2F1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0563811 |
Alignment:
TGAMCTTTGMMCYT
TAAACGATGCC---
| Original motif Consensus sequence: TGAMCTTTGMMCYT |
Reverse complement motif Consensus sequence: AKGYYCAAAGRTCA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
19 |
| Motif name: |
atactttggc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.5375 |
Alignment:
GCCAAAGTAT-
GGCATCGTTTA
| Original motif Consensus sequence: ATACTTTGGC |
Reverse complement motif Consensus sequence: GCCAAAGTAT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
31 |
| Motif name: |
ctatacggacg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.5625 |
Alignment:
CTATACGGACG-
-TAAACGATGCC
| Original motif Consensus sequence: CTATACGGACG |
Reverse complement motif Consensus sequence: CGTCCGTATAG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
27 |
| Motif name: |
AgTGCTCCACTGTGgTGTCACAgT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
17 |
| Number of overlap: |
8 |
| Similarity score: |
1.55502 |
Alignment:
ACTGTGACACCACAGTGGAGCACT---
----------------TAAACGATGCC
| Original motif Consensus sequence: AGTGCTCCACTGTGGTGTCACAGT |
Reverse complement motif Consensus sequence: ACTGTGACACCACAGTGGAGCACT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
21 |
| Motif name: |
AmTGTGACACCACAGT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
9 |
| Number of overlap: |
8 |
| Similarity score: |
1.55908 |
Alignment:
---AMTGTGACACCACAGT
TAAACGATGCC--------
| Original motif Consensus sequence: AMTGTGACACCACAGT |
Reverse complement motif Consensus sequence: ACTGTGGTGTCACART |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
1.5625 |
Alignment:
CGTGKCCGCGG---
---GGCATCGTTTA
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
33 |
| Motif name: |
srACyCCGAyr |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
7 |
| Similarity score: |
2.04464 |
Alignment:
----SRACYCCGAYR
TAAACGATGCC----
| Original motif Consensus sequence: SRACYCCGAYR |
Reverse complement motif Consensus sequence: KKTCGGKGTKS |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
13 |
| Number of overlap: |
7 |
| Similarity score: |
2.05416 |
Alignment:
BMSMGCCYMCTKSTGGMHM----
------------TAAACGATGCC
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
20 |
| Motif name: |
CagTGCTCCACTGTGgT |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
11 |
| Number of overlap: |
7 |
| Similarity score: |
2.05952 |
Alignment:
ABCACAGTGGAGCACTG----
----------GGCATCGTTTA
| Original motif Consensus sequence: CAGTGCTCCACTGTGBT |
Reverse complement motif Consensus sequence: ABCACAGTGGAGCACTG |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 31 |
Motif name: ctatacggacg |
| Original motif Consensus sequence: CTATACGGACG |
Reverse complement motif Consensus sequence: CGTCCGTATAG |
|
|
 |
 |
Best Matches for Motif ID 31 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
33 |
| Motif name: |
srACyCCGAyr |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0803571 |
Alignment:
SRACYCCGAYR
CTATACGGACG
| Original motif Consensus sequence: SRACYCCGAYR |
Reverse complement motif Consensus sequence: KKTCGGKGTKS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
30 |
| Motif name: |
taaacgatgcc |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.580357 |
Alignment:
-TAAACGATGCC
CTATACGGACG-
| Original motif Consensus sequence: TAAACGATGCC |
Reverse complement motif Consensus sequence: GGCATCGTTTA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
13 |
| Motif name: |
MIZF |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
1.0623 |
Alignment:
GCGGACGTTV--
-CGTCCGTATAG
| Original motif Consensus sequence: BAACGTCCGC |
Reverse complement motif Consensus sequence: GCGGACGTTV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
19 |
| Motif name: |
atactttggc |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
9 |
| Similarity score: |
1.06647 |
Alignment:
GCCAAAGTAT--
-CGTCCGTATAG
| Original motif Consensus sequence: ATACTTTGGC |
Reverse complement motif Consensus sequence: GCCAAAGTAT |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
3 |
| Number of overlap: |
9 |
| Similarity score: |
1.07758 |
Alignment:
--CCGCGGRCACG
CTATACGGACG--
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
28 |
| Motif name: |
ssGCkTGCssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
1.56473 |
Alignment:
---SSGCKTGCSSK
CGTCCGTATAG---
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
32 |
| Motif name: |
gCGCGCsCgsG |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
1.57871 |
Alignment:
GCGCGCSCGCG---
---CGTCCGTATAG
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
7 |
| Similarity score: |
2.06779 |
Alignment:
BBSGGCGGGGBS----
-----CTATACGGACG
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
6 |
| Similarity score: |
2.54768 |
Alignment:
-----VGCACGTGGH
CTATACGGACG----
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
6 |
| Similarity score: |
2.55363 |
Alignment:
-----HSCACGTGGC
CTATACGGACG----
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 32 |
Motif name: gCGCGCsCgsG |
| Original motif Consensus sequence: GCGCGCSCGCG |
Reverse complement motif Consensus sequence: CGCGSGCGCGC |
|
|
 |
 |
Best Matches for Motif ID 32 (Highest to Lowest)
| Dataset #: |
2 |
| Motif ID: |
29 |
| Motif name: |
cCGCGGrCACG |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0131931 |
Alignment:
CCGCGGRCACG
GCGCGCSCGCG
| Original motif Consensus sequence: CCGCGGRCACG |
Reverse complement motif Consensus sequence: CGTGKCCGCGG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
28 |
| Motif name: |
ssGCkTGCssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0299395 |
Alignment:
SSGCKTGCSSK
GCGCGCSCGCG
| Original motif Consensus sequence: SSGCKTGCSSK |
Reverse complement motif Consensus sequence: YSSGCAYGCSS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0338798 |
Alignment:
BSCCCCGCGBS
GCGCGCSCGCG
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0410595 |
Alignment:
SBCCCCGCCSBB
GCGCGCSCGCG-
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
16 |
| Motif name: |
CTCF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
11 |
| Similarity score: |
0.0537298 |
Alignment:
BMSMGCCYMCTKSTGGMHM
------GCGCGCSCGCG--
| Original motif Consensus sequence: YDRCCASYAGRKGGCRSYV |
Reverse complement motif Consensus sequence: BMSMGCCYMCTKSTGGMHM |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
4 |
| Number of overlap: |
11 |
| Similarity score: |
0.0651602 |
Alignment:
MGGTCAGGGTGACCTRDBHV
---GCGCGCSCGCG------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
13 |
| Motif name: |
MIZF |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.55601 |
Alignment:
BAACGTCCGC-
GCGCGCSCGCG
| Original motif Consensus sequence: BAACGTCCGC |
Reverse complement motif Consensus sequence: GCGGACGTTV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
8 |
| Motif name: |
Myc |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.561823 |
Alignment:
DCCACGTGCV-
GCGCGCSCGCG
| Original motif Consensus sequence: VGCACGTGGH |
Reverse complement motif Consensus sequence: DCCACGTGCV |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
6 |
| Motif name: |
Mycn |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
1 |
| Number of overlap: |
10 |
| Similarity score: |
0.561904 |
Alignment:
-HSCACGTGGC
GCGCGCSCGCG
| Original motif Consensus sequence: HSCACGTGGC |
Reverse complement motif Consensus sequence: GCCACGTGSD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
31 |
| Motif name: |
ctatacggacg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Backward |
| Position number: |
4 |
| Number of overlap: |
8 |
| Similarity score: |
1.56956 |
Alignment:
---CTATACGGACG
CGCGSGCGCGC---
| Original motif Consensus sequence: CTATACGGACG |
Reverse complement motif Consensus sequence: CGTCCGTATAG |
|
|
 |
 |
| Dataset #: 2 |
Motif ID: 33 |
Motif name: srACyCCGAyr |
| Original motif Consensus sequence: SRACYCCGAYR |
Reverse complement motif Consensus sequence: KKTCGGKGTKS |
|
|
 |
 |
Best Matches for Motif ID 33 (Highest to Lowest)
| Dataset #: |
1 |
| Motif ID: |
3 |
| Motif name: |
ESR1 |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0287254 |
Alignment:
MGGTCAGGGTGACCTRDBHV
-KKTCGGKGTKS--------
| Original motif Consensus sequence: VDBHMAGGTCACCCTGACCY |
Reverse complement motif Consensus sequence: MGGTCAGGGTGACCTRDBHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
23 |
| Motif name: |
TGGAGCACTGTGACAcCACAGTGg |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
11 |
| Similarity score: |
0.0316919 |
Alignment:
CCACTGTGVTGTCACAGTGCTCCA
---------KKTCGGKGTKS----
| Original motif Consensus sequence: TGGAGCACTGTGACAVCACAGTGG |
Reverse complement motif Consensus sequence: CCACTGTGVTGTCACAGTGCTCCA |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
9 |
| Motif name: |
ESR2 |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
5 |
| Number of overlap: |
11 |
| Similarity score: |
0.032424 |
Alignment:
BAGGYCABHBTGACCKHV
---SRACYCCGAYR----
| Original motif Consensus sequence: VHRGGTCABDBTGMCCTB |
Reverse complement motif Consensus sequence: BAGGYCABHBTGACCKHV |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
17 |
| Motif name: |
TgTGgTGTCACAGTGCTCC |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
6 |
| Number of overlap: |
11 |
| Similarity score: |
0.0326389 |
Alignment:
GGAGCACTGTGACACCACA
-----SRACYCCGAYR---
| Original motif Consensus sequence: TGTGGTGTCACAGTGCTCC |
Reverse complement motif Consensus sequence: GGAGCACTGTGACACCACA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
22 |
| Motif name: |
CACTGTGrYrtCACAGTGswsCAcT |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
7 |
| Number of overlap: |
11 |
| Similarity score: |
0.0342172 |
Alignment:
AGTGSWSCACTGTGAMAMCACAGTG
--------SRACYCCGAYR------
| Original motif Consensus sequence: CACTGTGRTRTCACAGTGSWSCACT |
Reverse complement motif Consensus sequence: AGTGSWSCACTGTGAMAMCACAGTG |
|
|
 |
 |
| Dataset #: |
1 |
| Motif ID: |
7 |
| Motif name: |
REST |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
7 |
| Number of overlap: |
11 |
| Similarity score: |
0.0347246 |
Alignment:
GGYGCTGTCCATGGTGCTGAA
------KKTCGGKGTKS----
| Original motif Consensus sequence: TTCAGCACCATGGACAGCKCC |
Reverse complement motif Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
31 |
| Motif name: |
ctatacggacg |
| Matching format of first motif: |
Original Motif |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
1 |
| Number of overlap: |
11 |
| Similarity score: |
0.0430556 |
Alignment:
CTATACGGACG
SRACYCCGAYR
| Original motif Consensus sequence: CTATACGGACG |
Reverse complement motif Consensus sequence: CGTCCGTATAG |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
25 |
| Motif name: |
wwCCAmAGTCmt |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Original Motif |
| Direction: |
Forward |
| Position number: |
2 |
| Number of overlap: |
11 |
| Similarity score: |
0.0430556 |
Alignment:
DDCCAMAGTCHB
-KKTCGGKGTKS
| Original motif Consensus sequence: DDCCAMAGTCHB |
Reverse complement motif Consensus sequence: VDGACTRTGGDD |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
24 |
| Motif name: |
ssCCCCGCSssk |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Forward |
| Position number: |
3 |
| Number of overlap: |
10 |
| Similarity score: |
0.513889 |
Alignment:
BBSGGCGGGGBS-
--KKTCGGKGTKS
| Original motif Consensus sequence: SBCCCCGCCSBB |
Reverse complement motif Consensus sequence: BBSGGCGGGGBS |
|
|
 |
 |
| Dataset #: |
2 |
| Motif ID: |
18 |
| Motif name: |
sscCCCGCGcs |
| Matching format of first motif: |
Reverse Complement |
| Matching format of second motif: |
Reverse Complement |
| Direction: |
Backward |
| Position number: |
2 |
| Number of overlap: |
10 |
| Similarity score: |
0.535703 |
Alignment:
-SBCGCGGGGSB
KKTCGGKGTKS-
| Original motif Consensus sequence: BSCCCCGCGBS |
Reverse complement motif Consensus sequence: SBCGCGGGGSB |
|
|
 |
 |































































































































































































































