Best Matches in Database for Each Motif (Highest to Lowest)
| Dataset #: 1 | Motif ID: 1 | Motif name: Motif 1 |
| Original motif | Reverse complement motif |
| Consensus sequence: RGGAAR | Consensus sequence: MTTCCK |
 |
 |
Best Matches for Motif ID 1 (Highest to Lowest)
| Motif ID: | UP00090 |
| Motif name: | Elf3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
TDAMTTCCYYDTD
---MTTCCK----
| Original motif | Reverse complement motif |
| Consensus sequence: DAHMMGGAARTDA | Consensus sequence: TDAMTTCCYYDTD |
 |
 |
| Motif ID: | UP00085 |
| Motif name: | Sfpi1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.00351709 |
Alignment:
HDACTTCCBCWTDV
---MTTCCK-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDAWGVGGAAGTDH | Consensus sequence: HDACTTCCBCWTDV |
 |
 |
| Motif ID: | UP00414 |
| Motif name: | Ets1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.00356242 |
Alignment:
DDBAMTTCCGGTDHDHB
----MTTCCK-------
| Original motif | Reverse complement motif |
| Consensus sequence: BDDDHACCGGAARTVDD | Consensus sequence: DDBAMTTCCGGTDHDHB |
 |
 |
| Motif ID: | UP00409 |
| Motif name: | Elf5 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 6 |
| Similarity score: | 0.00439981 |
Alignment:
BWAHSMGGAAGTWD
-----RGGAAR---
| Original motif | Reverse complement motif |
| Consensus sequence: BWAHSMGGAAGTWD | Consensus sequence: DWACTTCCRSDTWV |
 |
 |
| Motif ID: | UP00015 |
| Motif name: | Ehf_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 6 |
| Similarity score: | 0.00655644 |
Alignment:
DDACTTCCGGBTHBB
---MTTCCK------
| Original motif | Reverse complement motif |
| Consensus sequence: VBDABCCGGAAGTDD | Consensus sequence: DDACTTCCGGBTHBB |
 |
 |
| Motif ID: | UP00013 |
| Motif name: | Gabpa_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.00663687 |
Alignment:
BHHHACTTCCGGTHHBD
-----MTTCCK------
| Original motif | Reverse complement motif |
| Consensus sequence: HVHHACCGGAAGTDHHV | Consensus sequence: BHHHACTTCCGGTHHBD |
 |
 |
| Motif ID: | UP00413 |
| Motif name: | Elf4 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.00896416 |
Alignment:
DBBHACTTCCGGKWTH
-----MTTCCK-----
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHACTTCCGGKWTH | Consensus sequence: HAWYCCGGAAGTHBBD |
 |
 |
| Motif ID: | UP00419 |
| Motif name: | Spic |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 6 |
| Similarity score: | 0.00975526 |
Alignment:
VAAAGMGGAAGTWD
-----RGGAAR---
| Original motif | Reverse complement motif |
| Consensus sequence: VAAAGMGGAAGTWD | Consensus sequence: DWACTTCCRCTTTB |
 |
 |
| Motif ID: | UP00418 |
| Motif name: | Etv6 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0.0103447 |
Alignment:
DVHBTACTTCCGGHTHB
------MTTCCK-----
| Original motif | Reverse complement motif |
| Consensus sequence: DVHBTACTTCCGGHTHB | Consensus sequence: BHAHCCGGAAGTABDVD |
 |
 |
| Motif ID: | UP00404 |
| Motif name: | Elf2 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.010491 |
Alignment:
DBBHACTTCCGGDWDB
-----MTTCCK-----
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHACTTCCGGDWDB | Consensus sequence: BDWDCCGGAAGTHBBD |
 |
 |
| Dataset #: 1 | Motif ID: 2 | Motif name: Motif 2 |
| Original motif | Reverse complement motif |
| Consensus sequence: AMACA | Consensus sequence: TGTRT |
 |
 |
Best Matches for Motif ID 2 (Highest to Lowest)
| Motif ID: | UP00042 |
| Motif name: | Gm397_second |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 5 |
| Similarity score: | 0 |
Alignment:
DVMVGCACACACKCVH
------AMACA-----
| Original motif | Reverse complement motif |
| Consensus sequence: DVMVGCACACACKCVH | Consensus sequence: HBGYGTGTGTGCVRVD |
 |
 |
| Motif ID: | UP00025 |
| Motif name: | Foxk1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 5 |
| Similarity score: | 0.00605402 |
Alignment:
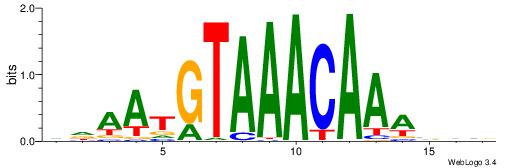
DDAATGTAAACAAAVVB
-------AMACA-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDAATGTAAACAAAVVB | Consensus sequence: BBVTTTGTTTACATTDD |
 |
 |
| Motif ID: | UP00041 |
| Motif name: | Foxj1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 5 |
| Similarity score: | 0.00753267 |
Alignment:
HDHRTAAACAAAHDDD
-----AMACA------
| Original motif | Reverse complement motif |
| Consensus sequence: HDHRTAAACAAAHDDD | Consensus sequence: DDDHTTTGTTTAMHDH |
 |
 |
| Motif ID: | UP00039 |
| Motif name: | Foxj3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 5 |
| Similarity score: | 0.00965634 |
Alignment:
HDHADGTAAACAAAVVH
-------AMACA-----
| Original motif | Reverse complement motif |
| Consensus sequence: HDHADGTAAACAAAVVH | Consensus sequence: DVVTTTGTTTACDTHDH |
 |
 |
| Motif ID: | UP00061 |
| Motif name: | Foxl1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 5 |
| Similarity score: | 0.00989492 |
Alignment:
DWAATRTAAACAAWVVB
-------AMACA-----
| Original motif | Reverse complement motif |
| Consensus sequence: DWAATRTAAACAAWVVB | Consensus sequence: VVVWTTGTTTAMATTWD |
 |
 |
| Motif ID: | UP00004 |
| Motif name: | Sox14_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 5 |
| Similarity score: | 0.0116453 |
Alignment:
BBDCMATTGTSTBRB
-------TGTRT---
| Original motif | Reverse complement motif |
| Consensus sequence: BKBASACAATRGHBB | Consensus sequence: BBDCMATTGTSTBRB |
 |
 |
| Motif ID: | UP00073 |
| Motif name: | Foxa2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 5 |
| Similarity score: | 0.0143122 |
Alignment:
HHWAWGTAAAYAAAVDB
-------AMACA-----
| Original motif | Reverse complement motif |
| Consensus sequence: HHWAWGTAAAYAAAVDB | Consensus sequence: BDVTTTKTTTACWTWHH |
 |
 |
| Motif ID: | UP00026 |
| Motif name: | Zscan4_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 5 |
| Similarity score: | 0.0153228 |
Alignment:
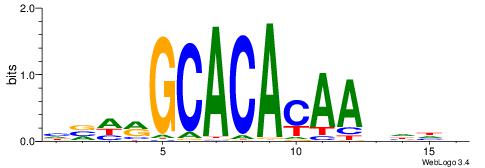
BHHHTTGTGTGCKHVD
-----TGTRT------
| Original motif | Reverse complement motif |
| Consensus sequence: HVHRGCACACAAHHHV | Consensus sequence: BHHHTTGTGTGCKHVD |
 |
 |
| Motif ID: | UP00011 |
| Motif name: | Irf6_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 5 |
| Similarity score: | 0.024195 |
Alignment:
DHDBTKGTTTCGHTHDD
-----TGTRT-------
| Original motif | Reverse complement motif |
| Consensus sequence: HDDAHCGAAACYAVDDD | Consensus sequence: DHDBTKGTTTCGHTHDD |
 |
 |
| Motif ID: | UP00000 |
| Motif name: | Smad3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 5 |
| Similarity score: | 0.0255974 |
Alignment:
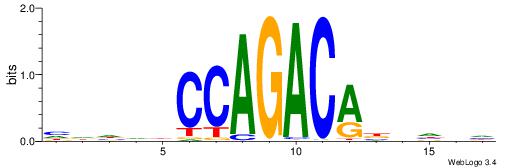
HDHDDCCAGACABBHVH
-------AMACA-----
| Original motif | Reverse complement motif |
| Consensus sequence: HDHDDCCAGACABBHVH | Consensus sequence: HVHBVTGTCTGGDDHDD |
 |
 |
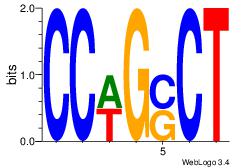
| Dataset #: 1 | Motif ID: 3 | Motif name: Motif 3 |
| Original motif | Reverse complement motif |
| Consensus sequence: AGSCWGG | Consensus sequence: CCWGSCT |
 |
 |
Best Matches for Motif ID 3 (Highest to Lowest)
| Motif ID: | UP00035 |
| Motif name: | Hic1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 7 |
| Similarity score: | 0 |
Alignment:
DDBHGGTTGGCATVBD
---AGSCWGG------
| Original motif | Reverse complement motif |
| Consensus sequence: DBBATGCCAACCHVHH | Consensus sequence: DDBHGGTTGGCATVBD |
 |
 |
| Motif ID: | UP00070 |
| Motif name: | Gcm1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 7 |
| Similarity score: | 0.00560414 |
Alignment:
DHDVATGCGGGTHVVH
----AGSCWGG-----
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHACCCGCATVDHD | Consensus sequence: DHDVATGCGGGTHVVH |
 |
 |
| Motif ID: | UP00095 |
| Motif name: | Zfp691_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 7 |
| Similarity score: | 0.0109149 |
Alignment:
DABBWGACTCCTHDVHV
--CCWGSCT--------
| Original motif | Reverse complement motif |
| Consensus sequence: DABBWGACTCCTHDVHV | Consensus sequence: VHBDDAGGAGTCWBBTD |
 |
 |
| Motif ID: | UP00006 |
| Motif name: | Zic3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 7 |
| Similarity score: | 0.0109242 |
Alignment:
DVHCYTGCTGTGHDH
--CCWGSCT------
| Original motif | Reverse complement motif |
| Consensus sequence: DDDCACAGCAKGHVD | Consensus sequence: DVHCYTGCTGTGHDH |
 |
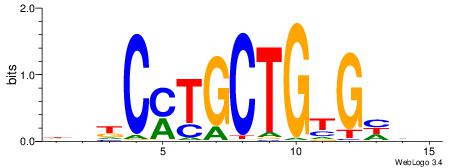 |
| Motif ID: | UP00000 |
| Motif name: | Smad3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 7 |
| Similarity score: | 0.0114503 |
Alignment:
HDHDDCCAGACABBHVH
-----CCWGSCT-----
| Original motif | Reverse complement motif |
| Consensus sequence: HDHDDCCAGACABBHVH | Consensus sequence: HVHBVTGTCTGGDDHDD |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 7 |
| Similarity score: | 0.0115258 |
Alignment:
CBDMCMGGGTGGTCCHVBVBAH
-----AGSCWGG----------
| Original motif | Reverse complement motif |
| Consensus sequence: HTBVVVDGGACCACCCRGRDBG | Consensus sequence: CBDMCMGGGTGGTCCHVBVBAH |
 |
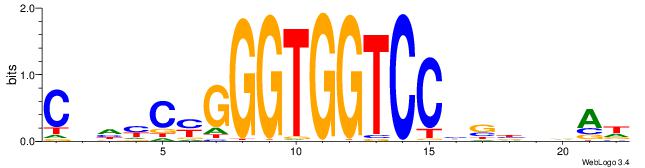 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 7 |
| Similarity score: | 0.0117486 |
Alignment:
HGDHVHKGACCACCCGGBGBGDB
---------CCWGSCT-------
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
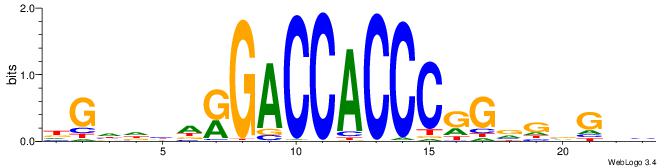 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 7 |
| Similarity score: | 0.0131676 |
Alignment:
CVHBSCGGGTGGTCMHVHHCDH
-----AGSCWGG----------
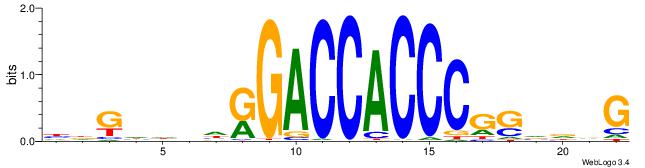
| Original motif | Reverse complement motif |
| Consensus sequence: HDGHHBHRGACCACCCGSVDVG | Consensus sequence: CVHBSCGGGTGGTCMHVHHCDH |
 |
 |
| Motif ID: | UP00081 |
| Motif name: | Mybl1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 7 |
| Similarity score: | 0.0136026 |
Alignment:
BHCGGCAGTTGGBBB
--AGSCWGG------
| Original motif | Reverse complement motif |
| Consensus sequence: BBVCCAACTGCCGHB | Consensus sequence: BHCGGCAGTTGGBBB |
 |
 |
| Motif ID: | UP00057 |
| Motif name: | Zic2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 7 |
| Similarity score: | 0.0139558 |
Alignment:
DVBCYTGCTGBGDDD
--CCWGSCT------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDCVCAGCAKGVVD | Consensus sequence: DVBCYTGCTGBGDDD |
 |
 |
| Dataset #: 1 | Motif ID: 4 | Motif name: Motif 4 |
| Original motif | Reverse complement motif |
| Consensus sequence: CTCTGY | Consensus sequence: KCAGAG |
 |
 |
Best Matches for Motif ID 4 (Highest to Lowest)
| Motif ID: | UP00031 |
| Motif name: | Zbtb3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
BBBAVTGCAGTGBBVDD
------KCAGAG-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDBBBCACTGCABTBBB | Consensus sequence: BBBAVTGCAGTGBBVDD |
 |
 |
| Motif ID: | UP00018 |
| Motif name: | Irf4_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0.00610915 |
Alignment:
BHHWCCGAGAMTVVB
---KCAGAG------
| Original motif | Reverse complement motif |
| Consensus sequence: VVBAYTCTCGGWHDB | Consensus sequence: BHHWCCGAGAMTVVB |
 |
 |
| Motif ID: | UP00095 |
| Motif name: | Zfp691_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.00648219 |
Alignment:
VHBDDAGGAGTCWBBTD
----KCAGAG-------
| Original motif | Reverse complement motif |
| Consensus sequence: DABBWGACTCCTHDVHV | Consensus sequence: VHBDDAGGAGTCWBBTD |
 |
 |
| Motif ID: | UP00006 |
| Motif name: | Zic3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 6 |
| Similarity score: | 0.00853125 |
Alignment:
DVHCYTGCTGTGHDH
-------CTCTGY--
| Original motif | Reverse complement motif |
| Consensus sequence: DDDCACAGCAKGHVD | Consensus sequence: DVHCYTGCTGTGHDH |
 |
 |
| Motif ID: | UP00007 |
| Motif name: | Egr1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.00895004 |
Alignment:
BHHDTCCCACTCBDBV
---------CTCTGY-
| Original motif | Reverse complement motif |
| Consensus sequence: BBHBGAGTGGGAHHDB | Consensus sequence: BHHDTCCCACTCBDBV |
 |
 |
| Motif ID: | UP00095 |
| Motif name: | Zfp691_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.00955889 |
Alignment:
DDHBYGAGCACTGBHHB
--------CTCTGY---
| Original motif | Reverse complement motif |
| Consensus sequence: BDHVCAGTGCTCMBHDD | Consensus sequence: DDHBYGAGCACTGBHHB |
 |
 |
| Motif ID: | UP00067 |
| Motif name: | Lef1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.0106184 |
Alignment:
VDDAGATCAAAGGKDDD
------KCAGAG-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATCTDDV | Consensus sequence: VDDAGATCAAAGGKDDD |
 |
 |
| Motif ID: | UP00102 |
| Motif name: | Zic1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0.0122635 |
Alignment:
BHDCVCAGCAGGHVD
--KCAGAG-------
| Original motif | Reverse complement motif |
| Consensus sequence: BHDCVCAGCAGGHVD | Consensus sequence: DVHCCTGCTGBGDDB |
 |
 |
| Motif ID: | UP00083 |
| Motif name: | Tcf7l2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.0131237 |
Alignment:
BDDASATCAAAGGMDDD
------KCAGAG-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATSTDDV | Consensus sequence: BDDASATCAAAGGMDDD |
 |
 |
| Motif ID: | UP00057 |
| Motif name: | Zic2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 6 |
| Similarity score: | 0.0136526 |
Alignment:
HHDCVCAGCAKGVVD
--KCAGAG-------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDCVCAGCAKGVVD | Consensus sequence: DVBCYTGCTGBGDDD |
 |
 |
| Dataset #: 1 | Motif ID: 5 | Motif name: Motif 5 |
| Original motif | Reverse complement motif |
| Consensus sequence: ATKTM | Consensus sequence: RARAT |
 |
 |
Best Matches for Motif ID 5 (Highest to Lowest)
| Motif ID: | UP00080 |
| Motif name: | Gata5_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 5 |
| Similarity score: | 0 |
Alignment:
VVBHRAGATATCWDHBB
----RARAT--------
| Original motif | Reverse complement motif |
| Consensus sequence: VVBHRAGATATCWDHBB | Consensus sequence: VVHHWGATATCTMHBBV |
 |
 |
| Motif ID: | UP00148 |
| Motif name: | Hdx |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 5 |
| Similarity score: | 0.00279496 |
Alignment:
DDKDHGAAATCAHHBHV
-----RARAT-------
| Original motif | Reverse complement motif |
| Consensus sequence: DDKDHGAAATCAHHBHV | Consensus sequence: BDBDHTGATTTCDHYDD |
 |
 |
| Motif ID: | UP00032 |
| Motif name: | Gata3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 5 |
| Similarity score: | 0.00967655 |
Alignment:
BDHDDADAGATAAGADHBDHVD
------RARAT-----------
| Original motif | Reverse complement motif |
| Consensus sequence: BDHDDADAGATAAGADHBDHVD | Consensus sequence: HBHDVHDTCTTATCTHTDDHDV |
 |
 |
| Motif ID: | UP00250 |
| Motif name: | Irx5 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 5 |
| Similarity score: | 0.0116566 |
Alignment:
DDDHWACATGTWHDABW
-------ATKTM-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDHWACATGTWHDABW | Consensus sequence: WVTDHWACATGTWHDDD |
 |
 |
| Motif ID: | UP00236 |
| Motif name: | Irx2 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 5 |
| Similarity score: | 0.0120196 |
Alignment:
WDDDTACATGTADDWBW
-------ATKTM-----
| Original motif | Reverse complement motif |
| Consensus sequence: WDDDTACATGTADDWBW | Consensus sequence: WVWDDTACATGTADDDW |
 |
 |
| Motif ID: | UP00150 |
| Motif name: | Irx6 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 5 |
| Similarity score: | 0.0120601 |
Alignment:
HAHDTACATGTAVDWHT
-------ATKTM-----
| Original motif | Reverse complement motif |
| Consensus sequence: HAHDTACATGTAVDWHT | Consensus sequence: AHWDBTACATGTADHTH |
 |
 |
| Motif ID: | UP00223 |
| Motif name: | Irx3_2226.1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 5 |
| Similarity score: | 0.0124884 |
Alignment:
DDDHTACATGTAVWDHD
-------ATKTM-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDHTACATGTAVWDHD | Consensus sequence: DHDWBTACATGTAHDDD |
 |
 |
| Motif ID: | UP00033 |
| Motif name: | Zfp410_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 5 |
| Similarity score: | 0.0150044 |
Alignment:
DVDDATGGGATGKMDDV
---------ATKTM---
| Original motif | Reverse complement motif |
| Consensus sequence: DVDDATGGGATGKMDDV | Consensus sequence: BDDYYCATCCCATDDBD |
 |
 |
| Motif ID: | UP00223 |
| Motif name: | Irx3_0920.1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 5 |
| Similarity score: | 0.0150192 |
Alignment:
DDDDTACATGTAVWWHD
-------ATKTM-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDDTACATGTAVWWHD | Consensus sequence: DDWWBTACATGTADDDD |
 |
 |
| Motif ID: | UP00528 |
| Motif name: | Foxm1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 12 |
| Number of overlap: | 5 |
| Similarity score: | 0.0166075 |
Alignment:
MWSMBAARRATGCDCABHATGD
------RARAT-----------
| Original motif | Reverse complement motif |
| Consensus sequence: MWSMBAARRATGCDCABHATGD | Consensus sequence: DCATDBTGHGCATKMTTBRSWR |
 |
 |
| Dataset #: 1 | Motif ID: 6 | Motif name: Motif 6 |
| Original motif | Reverse complement motif |
| Consensus sequence: CMGCRGC | Consensus sequence: GCKGCYG |
 |
 |
Best Matches for Motif ID 6 (Highest to Lowest)
| Motif ID: | UP00052 |
| Motif name: | Osr2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 7 |
| Similarity score: | 0 |
Alignment:
HHHHGCTACKGTHVHH
----GCKGCYG-----
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHACRGTAGCHHHD | Consensus sequence: HHHHGCTACKGTHVHH |
 |
 |
| Motif ID: | UP00027 |
| Motif name: | Osr1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 7 |
| Similarity score: | 0.00108105 |
Alignment:
DHBHGCTACKGTHDHH
----GCKGCYG-----
| Original motif | Reverse complement motif |
| Consensus sequence: HHDHACRGTAGCHVHD | Consensus sequence: DHBHGCTACKGTHDHH |
 |
 |
| Motif ID: | UP00099 |
| Motif name: | Ascl2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 7 |
| Similarity score: | 0.00905497 |
Alignment:
BBYVVCAGCTGCBVHHD
-------GCKGCYG---
| Original motif | Reverse complement motif |
| Consensus sequence: BBYVVCAGCTGCBVHHD | Consensus sequence: HHDVVGCAGCTGVBKVB |
 |
 |
| Motif ID: | UP00102 |
| Motif name: | Zic1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 7 |
| Similarity score: | 0.0111061 |
Alignment:
DVHCCTGCTGBGDDB
---GCKGCYG-----
| Original motif | Reverse complement motif |
| Consensus sequence: BHDCVCAGCAGGHVD | Consensus sequence: DVHCCTGCTGBGDDB |
 |
 |
| Motif ID: | UP00057 |
| Motif name: | Zic2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 7 |
| Similarity score: | 0.0126897 |
Alignment:
HHDCVCAGCAKGVVD
-----CMGCRGC---
| Original motif | Reverse complement motif |
| Consensus sequence: HHDCVCAGCAKGVVD | Consensus sequence: DVBCYTGCTGBGDDD |
 |
 |
| Motif ID: | UP00006 |
| Motif name: | Zic3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 7 |
| Similarity score: | 0.0137402 |
Alignment:
DVHCYTGCTGTGHDH
---GCKGCYG-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDCACAGCAKGHVD | Consensus sequence: DVHCYTGCTGTGHDH |
 |
 |
| Motif ID: | UP00045 |
| Motif name: | Mafb_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 7 |
| Similarity score: | 0.0253076 |
Alignment:
DMTWDGTCAGCADWTTH
-------CMGCRGC---
| Original motif | Reverse complement motif |
| Consensus sequence: HAAWDTGCTGACDWARH | Consensus sequence: DMTWDGTCAGCADWTTH |
 |
 |
| Motif ID: | UP00193 |
| Motif name: | Rhox11_1765.2 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 7 |
| Similarity score: | 0.0292668 |
Alignment:
WVVHWTWACAGCGBHBT
--------CMGCRGC--
| Original motif | Reverse complement motif |
| Consensus sequence: AVDVCGCTGTWAWDVVW | Consensus sequence: WVVHWTWACAGCGBHBT |
 |
 |
| Motif ID: | UP00193 |
| Motif name: | Rhox11_2205.1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 7 |
| Similarity score: | 0.0298083 |
Alignment:
WVVHWTWACAGCGHHVT
--------CMGCRGC--
| Original motif | Reverse complement motif |
| Consensus sequence: AVDHCGCTGTWAWDVVW | Consensus sequence: WVVHWTWACAGCGHHVT |
 |
 |
| Motif ID: | UP00122 |
| Motif name: | Tgif1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 7 |
| Similarity score: | 0.0332976 |
Alignment:
DABWWTGACAGCTVVBD
--------CMGCRGC--
| Original motif | Reverse complement motif |
| Consensus sequence: DABWWTGACAGCTVVBD | Consensus sequence: DBVVAGCTGTCAWWVTH |
 |
 |
| Dataset #: 1 | Motif ID: 7 | Motif name: Motif 7 |
| Original motif | Reverse complement motif |
| Consensus sequence: CCCRCCCC | Consensus sequence: GGGGMGGG |
 |
 |
Best Matches for Motif ID 7 (Highest to Lowest)
| Motif ID: | UP00093 |
| Motif name: | Klf7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
HBDRCCMCGCCCHHHD
-----CCCRCCCC---
| Original motif | Reverse complement motif |
| Consensus sequence: HBDRCCMCGCCCHHHD | Consensus sequence: DHHDGGGCGRGGKHBH |
 |
 |
| Motif ID: | UP00002 |
| Motif name: | Sp4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.00490878 |
Alignment:
BVVHAGGGGGCGRDHHB
------GGGGMGGG---
| Original motif | Reverse complement motif |
| Consensus sequence: BDHHMCGCCCCCTHVBB | Consensus sequence: BVVHAGGGGGCGRDHHB |
 |
 |
| Motif ID: | UP00007 |
| Motif name: | Egr1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0101441 |
Alignment:
HCCGCCCCCGCAHB
CCCRCCCC------
| Original motif | Reverse complement motif |
| Consensus sequence: HCCGCCCCCGCAHB | Consensus sequence: VHTGCGGGGGCGGH |
 |
 |
| Motif ID: | UP00021 |
| Motif name: | Zfp281_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0106908 |
Alignment:
DBCCCCCCCCCCMYC
---CCCRCCCC----
| Original motif | Reverse complement motif |
| Consensus sequence: DBCCCCCCCCCCMYC | Consensus sequence: GKRGGGGGGGGGGBD |
 |
 |
| Motif ID: | UP00043 |
| Motif name: | Bcl6b_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0152681 |
Alignment:
HBBBCCGCCCCWHHHV
---CCCRCCCC-----
| Original motif | Reverse complement motif |
| Consensus sequence: HBBBCCGCCCCWHHHV | Consensus sequence: BHHHWGGGGCGGBBVH |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0.0252184 |
Alignment:
MHHHWHYTMCCHCCCACVAABVH
--------CCCRCCCC-------
| Original motif | Reverse complement motif |
| Consensus sequence: MHHHWHYTMCCHCCCACVAABVH | Consensus sequence: DVVTTVGTGGGHGGYAMHWHHHY |
 |
 |
| Motif ID: | UP00022 |
| Motif name: | Zfp740_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0258528 |
Alignment:
BHHDYGGGGGGGGBVD
------GGGGMGGG--
| Original motif | Reverse complement motif |
| Consensus sequence: HVBCCCCCCCCMHHHB | Consensus sequence: BHHDYGGGGGGGGBVD |
 |
 |
| Motif ID: | UP00007 |
| Motif name: | Egr1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.0318239 |
Alignment:
BHHDTCCCACTCBDBV
-----CCCRCCCC---
| Original motif | Reverse complement motif |
| Consensus sequence: BBHBGAGTGGGAHHDB | Consensus sequence: BHHDTCCCACTCBDBV |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.0333599 |
Alignment:
HGDHVHKGACCACCCGGBGBGDB
--------CCCRCCCC-------
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00033 |
| Motif name: | Zfp410_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.0336185 |
Alignment:
HHHHDGGGGCGGDBHDV
-----GGGGMGGG----
| Original motif | Reverse complement motif |
| Consensus sequence: BHHBHCCGCCCCDHHHH | Consensus sequence: HHHHDGGGGCGGDBHDV |
 |
 |
| Dataset #: 1 | Motif ID: 8 | Motif name: Motif 8 |
| Original motif | Reverse complement motif |
| Consensus sequence: CTGGGA | Consensus sequence: TCCCAG |
 |
 |
Best Matches for Motif ID 8 (Highest to Lowest)
| Motif ID: | UP00033 |
| Motif name: | Zfp410_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
DVDDATGGGATGKMDDV
----CTGGGA-------
| Original motif | Reverse complement motif |
| Consensus sequence: DVDDATGGGATGKMDDV | Consensus sequence: BDDYYCATCCCATDDBD |
 |
 |
| Motif ID: | UP00216 |
| Motif name: | Obox1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.000600574 |
Alignment:
DHDGTTAATCCCCKTDD
--------TCCCAG---
| Original motif | Reverse complement motif |
| Consensus sequence: DDARGGGGATTAACDHH | Consensus sequence: DHDGTTAATCCCCKTDD |
 |
 |
| Motif ID: | UP00229 |
| Motif name: | Otx1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.0140775 |
Alignment:
VHHWATTAATCCCMDHH
---------TCCCAG--
| Original motif | Reverse complement motif |
| Consensus sequence: DDDRGGGATTAATWHHB | Consensus sequence: VHHWATTAATCCCMDHH |
 |
 |
| Motif ID: | UP00160 |
| Motif name: | Obox3 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0142192 |
Alignment:
HHWGTTAATCCCCCBBH
--------TCCCAG---
| Original motif | Reverse complement motif |
| Consensus sequence: HBVGGGGGATTAACWHH | Consensus sequence: HHWGTTAATCCCCCBBH |
 |
 |
| Motif ID: | UP00239 |
| Motif name: | Obox2 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 6 |
| Similarity score: | 0.0184594 |
Alignment:
HTARTTAATCCCCCKBD
--------TCCCAG---
| Original motif | Reverse complement motif |
| Consensus sequence: DBRGGGGGATTAAMTAH | Consensus sequence: HTARTTAATCCCCCKBD |
 |
 |
| Motif ID: | UP00416 |
| Motif name: | Fli1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0.018604 |
Alignment:
DHDACCGGAAKTBBVV
----CTGGGA------
| Original motif | Reverse complement motif |
| Consensus sequence: VBBVAYTTCCGGTDDH | Consensus sequence: DHDACCGGAAKTBBVV |
 |
 |
| Motif ID: | UP00411 |
| Motif name: | Erg |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.0196433 |
Alignment:
BDDACCGGAAKTVBVB
----CTGGGA------
| Original motif | Reverse complement motif |
| Consensus sequence: VBBVAYTTCCGGTDDB | Consensus sequence: BDDACCGGAAKTVBVB |
 |
 |
| Motif ID: | UP00413 |
| Motif name: | Elf4 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.0196806 |
Alignment:
HAWYCCGGAAGTHBBD
----CTGGGA------
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHACTTCCGGKWTH | Consensus sequence: HAWYCCGGAAGTHBBD |
 |
 |
| Motif ID: | UP00143 |
| Motif name: | Dobox5 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.0199632 |
Alignment:
DDHVGGGATTAATHDTH
--CTGGGA---------
| Original motif | Reverse complement motif |
| Consensus sequence: DDHVGGGATTAATHDTH | Consensus sequence: DADHATTAATCCCBHHH |
 |
 |
| Motif ID: | UP00265 |
| Motif name: | Pitx3 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.0210625 |
Alignment:
GBHDGCTAATCCCCCB
---------TCCCAG-
| Original motif | Reverse complement motif |
| Consensus sequence: VGGGGGATTAGCDDBC | Consensus sequence: GBHDGCTAATCCCCCB |
 |
 |
| Dataset #: 1 | Motif ID: 9 | Motif name: Motif 9 |
| Original motif | Reverse complement motif |
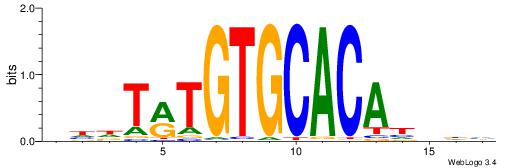
| Consensus sequence: CACACAVRCACACA | Consensus sequence: TGTGTGKVTGTGTG |
 |
 |
Best Matches for Motif ID 9 (Highest to Lowest)
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 14 |
| Similarity score: | 0.0487588 |
Alignment:
VYWVVHCRCCACMCACGAHSBHH
-CACACAVRCACACA--------
| Original motif | Reverse complement motif |
| Consensus sequence: VYWVVHCRCCACMCACGAHSBHH | Consensus sequence: HHBSHTCGTGRGTGGKGDBVWMV |
 |
 |
| Motif ID: | UP00042 |
| Motif name: | Gm397_second |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0496974 |
Alignment:
HBGYGTGTGTGCVRVD
-TGTGTGKVTGTGTG-
| Original motif | Reverse complement motif |
| Consensus sequence: DVMVGCACACACKCVH | Consensus sequence: HBGYGTGTGTGCVRVD |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 14 |
| Similarity score: | 0.0503392 |
Alignment:
HVBDAGCGBGGGTGGCRDBDDDB
TGTGTGKVTGTGTG---------
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Motif ID: | UP00025 |
| Motif name: | Foxk1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0506792 |
Alignment:
HDDTGTTKTTKTHHD
-TGTGTGKVTGTGTG
| Original motif | Reverse complement motif |
| Consensus sequence: HHHAYAAYAACAHHH | Consensus sequence: HDDTGTTKTTKTHHD |
 |
 |
| Motif ID: | UP00036 |
| Motif name: | Myf6_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0516113 |
Alignment:
VVVRACAGVCBCDCC
-CACACAVRCACACA
| Original motif | Reverse complement motif |
| Consensus sequence: VVVRACAGVCBCDCC | Consensus sequence: GGDGBGVCTGTKVVB |
 |
 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0523703 |
Alignment:
GTVBDAATTGTVTGHGDDHKVB
CACACAVRCACACA--------
| Original motif | Reverse complement motif |
| Consensus sequence: GTVBDAATTGTVTGHGDDHKVB | Consensus sequence: VVYDDDCHCAVACAATTDBVAC |
 |
 |
| Motif ID: | UP00026 |
| Motif name: | Zscan4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0557355 |
Alignment:
DHVDTGTGCACAYAHDH
CACACAVRCACACA---
| Original motif | Reverse complement motif |
| Consensus sequence: DHVDTGTGCACAYAHDH | Consensus sequence: HDHTMTGTGCACADVHD |
 |
 |
| Motif ID: | UP00526 |
| Motif name: | Foxn1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0578572 |
Alignment:
DDRMHBACGCGYGCGTDGBBDB
-CACACAVRCACACA-------
| Original motif | Reverse complement motif |
| Consensus sequence: DDRMHBACGCGYGCGTDGBBDB | Consensus sequence: BDVBCDACGCKCGCGTBHRKHD |
 |
 |
| Motif ID: | UP00400 |
| Motif name: | Zif268 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0580046 |
Alignment:
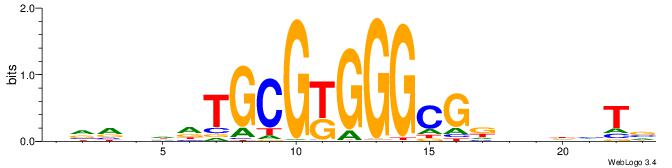
VVVDHHTGCGYGGGCGDHVHHTB
------TGTGTGKVTGTGTG---
| Original motif | Reverse complement motif |
| Consensus sequence: BADHBDHCGCCCMCGCAHHDBBV | Consensus sequence: VVVDHHTGCGYGGGCGDHVHHTB |
 |
 |
| Motif ID: | UP00014 |
| Motif name: | Sox17_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0581238 |
Alignment:
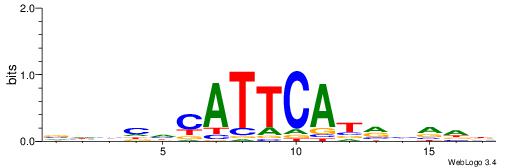
DDBDKATGAATGHDBHV
--TGTGTGKVTGTGTG-
| Original motif | Reverse complement motif |
| Consensus sequence: VHBHHCATTCATRHVDD | Consensus sequence: DDBDKATGAATGHDBHV |
 |
 |
| Dataset #: 1 | Motif ID: 10 | Motif name: Motif 10 |
| Original motif | Reverse complement motif |
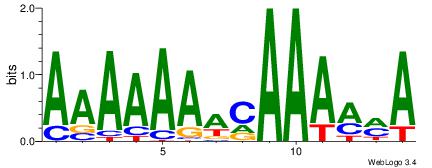
| Consensus sequence: TKKTTTGTTTTTTT | Consensus sequence: AAAAAAACAAARRA |
 |
 |
Best Matches for Motif ID 10 (Highest to Lowest)
| Motif ID: | UP00061 |
| Motif name: | Foxl1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0319341 |
Alignment:
HDTYTTGTTWTKDDDT
TKKTTTGTTTTTTT--
| Original motif | Reverse complement motif |
| Consensus sequence: ADDDYAWAACAAMAHH | Consensus sequence: HDTYTTGTTWTKDDDT |
 |
 |
| Motif ID: | UP00062 |
| Motif name: | Sox4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0368361 |
Alignment:
HBDDDAACAAAGRVBHH
AAAAAAACAAARRA---
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDAACAAAGRVBHH | Consensus sequence: HHBBMCTTTGTTHDDBH |
 |
 |
| Motif ID: | UP00030 |
| Motif name: | Sox11_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0370295 |
Alignment:
HBDDRAACAAAGRABHH
AAAAAAACAAARRA---
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDRAACAAAGRABHH | Consensus sequence: HHBTMCTTTGTTMDDVH |
 |
 |
| Motif ID: | UP00077 |
| Motif name: | Srf_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.038882 |
Alignment:
HHVBYTTTTTTTTDDVV
-TKKTTTGTTTTTTT--
| Original motif | Reverse complement motif |
| Consensus sequence: VBDDAAAAAAAAMVBHH | Consensus sequence: HHVBYTTTTTTTTDDVV |
 |
 |
| Motif ID: | UP00101 |
| Motif name: | Sox12_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0402601 |
Alignment:
HDTMCTTTGTSTVDBB
AAAAAAACAAARRA--
| Original motif | Reverse complement motif |
| Consensus sequence: VVDBASACAAAGRADH | Consensus sequence: HDTMCTTTGTSTVDBB |
 |
 |
| Motif ID: | UP00073 |
| Motif name: | Foxa2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0402844 |
Alignment:
VVDAATAACAAAVVB
-AAAAAAACAAARRA
| Original motif | Reverse complement motif |
| Consensus sequence: VVDAATAACAAAVVB | Consensus sequence: BVVTTTGTTATTDBB |
 |
 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0409353 |
Alignment:
HHBDHDDAACAATDRHHHHHBW
--AAAAAAACAAARRA------
| Original motif | Reverse complement motif |
| Consensus sequence: HHBDHDDAACAATDRHHHHHBW | Consensus sequence: WBHHHHHMDATTGTTHDHDVHH |
 |
 |
| Motif ID: | UP00025 |
| Motif name: | Foxk1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0416172 |
Alignment:
HDDTGTTKTTKTHHD
-TKKTTTGTTTTTTT
| Original motif | Reverse complement motif |
| Consensus sequence: HHHAYAAYAACAHHH | Consensus sequence: HDDTGTTKTTKTHHD |
 |
 |
| Motif ID: | UP00039 |
| Motif name: | Foxj3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0419017 |
Alignment:
DVVTTTGTTTACDTHDH
---AAAAAAACAAARRA
| Original motif | Reverse complement motif |
| Consensus sequence: HDHADGTAAACAAAVVH | Consensus sequence: DVVTTTGTTTACDTHDH |
 |
 |
| Motif ID: | UP00073 |
| Motif name: | Foxa2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0439168 |
Alignment:
BDVTTTKTTTACWTWHH
---AAAAAAACAAARRA
| Original motif | Reverse complement motif |
| Consensus sequence: HHWAWGTAAAYAAAVDB | Consensus sequence: BDVTTTKTTTACWTWHH |
 |
 |
| Dataset #: 1 | Motif ID: 11 | Motif name: Motif 11 |
| Original motif | Reverse complement motif |
| Consensus sequence: CAYACACACACACA | Consensus sequence: TGTGTGTGTGTKTG |
 |
 |
Best Matches for Motif ID 11 (Highest to Lowest)
| Motif ID: | UP00042 |
| Motif name: | Gm397_second |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0235042 |
Alignment:
DVMVGCACACACKCVH
-CAYACACACACACA-
| Original motif | Reverse complement motif |
| Consensus sequence: DVMVGCACACACKCVH | Consensus sequence: HBGYGTGTGTGCVRVD |
 |
 |
| Motif ID: | UP00026 |
| Motif name: | Zscan4_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0372612 |
Alignment:
BHHHTTGTGTGCKHVD
-TGTGTGTGTGTKTG-
| Original motif | Reverse complement motif |
| Consensus sequence: HVHRGCACACAAHHHV | Consensus sequence: BHHHTTGTGTGCKHVD |
 |
 |
| Motif ID: | UP00000 |
| Motif name: | Smad3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0446373 |
Alignment:
HVHBVTGTCTGGDDHDD
---TGTGTGTGTGTKTG
| Original motif | Reverse complement motif |
| Consensus sequence: HDHDDCCAGACABBHVH | Consensus sequence: HVHBVTGTCTGGDDHDD |
 |
 |
| Motif ID: | UP00400 |
| Motif name: | Zif268 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0457169 |
Alignment:
BADHBDHCGCCCMCGCAHHDBBV
---CAYACACACACACA------
| Original motif | Reverse complement motif |
| Consensus sequence: BADHBDHCGCCCMCGCAHHDBBV | Consensus sequence: VVVDHHTGCGYGGGCGDHVHHTB |
 |
 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0477221 |
Alignment:
GTVBDAATTGTVTGHGDDHKVB
CAYACACACACACA--------
| Original motif | Reverse complement motif |
| Consensus sequence: GTVBDAATTGTVTGHGDDHKVB | Consensus sequence: VVYDDDCHCAVACAATTDBVAC |
 |
 |
| Motif ID: | UP00025 |
| Motif name: | Foxk1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0485483 |
Alignment:
HDDTGTTKTTKTHHD
-TGTGTGTGTGTKTG
| Original motif | Reverse complement motif |
| Consensus sequence: HHHAYAAYAACAHHH | Consensus sequence: HDDTGTTKTTKTHHD |
 |
 |
| Motif ID: | UP00041 |
| Motif name: | Foxj1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0488633 |
Alignment:
HDHRTAAACAAAHDDD
CAYACACACACACA--
| Original motif | Reverse complement motif |
| Consensus sequence: HDHRTAAACAAAHDDD | Consensus sequence: DDDHTTTGTTTAMHDH |
 |
 |
| Motif ID: | UP00026 |
| Motif name: | Zscan4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0489596 |
Alignment:
DHVDTGTGCACAYAHDH
--CAYACACACACACA-
| Original motif | Reverse complement motif |
| Consensus sequence: DHVDTGTGCACAYAHDH | Consensus sequence: HDHTMTGTGCACADVHD |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 14 |
| Similarity score: | 0.0501133 |
Alignment:
HVBDAGCGBGGGTGGCRDBDDDB
TGTGTGTGTGTKTG---------
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Motif ID: | UP00025 |
| Motif name: | Foxk1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0518949 |
Alignment:
DDAATGTAAACAAAVVB
--CAYACACACACACA-
| Original motif | Reverse complement motif |
| Consensus sequence: DDAATGTAAACAAAVVB | Consensus sequence: BBVTTTGTTTACATTDD |
 |
 |
| Dataset #: 2 | Motif ID: 12 | Motif name: Zfx |
| Original motif | Reverse complement motif |
| Consensus sequence: BBVGCCBVGGCCTV | Consensus sequence: VAGGCCBBGGCVBB |
 |
 |
Best Matches for Motif ID 12 (Highest to Lowest)
| Motif ID: | UP00005 |
| Motif name: | Tcfap2a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0596136 |
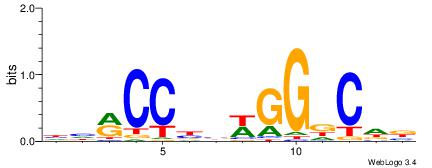
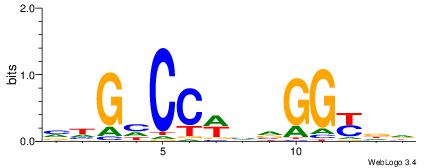
Alignment:
HHRCCBBWGGDCDB
BBVGCCBVGGCCTV
| Original motif | Reverse complement motif |
| Consensus sequence: HHRCCBBWGGDCDB | Consensus sequence: BDGHCCWBVGGKDH |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 14 |
| Similarity score: | 0.0603516 |
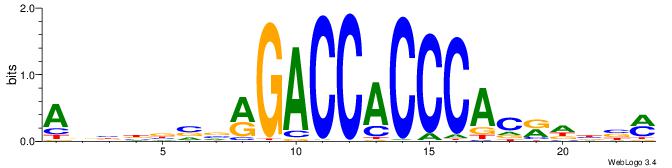
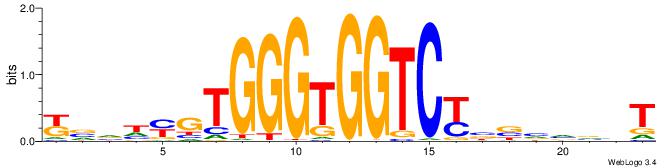
Alignment:
ABBBBVVRGACCACCCACRDBBM
BBVGCCBVGGCCTV---------
| Original motif | Reverse complement motif |
| Consensus sequence: ABBBBVVRGACCACCCACRDBBM | Consensus sequence: YBVDMGTGGGTGGTCKVVBVBBT |
 |
 |
| Motif ID: | UP00002 |
| Motif name: | Sp4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0604694 |
Alignment:
BDHHMCGCCCCCTHVBB
---VAGGCCBBGGCVBB
| Original motif | Reverse complement motif |
| Consensus sequence: BDHHMCGCCCCCTHVBB | Consensus sequence: BVVHAGGGGGCGRDHHB |
 |
 |
| Motif ID: | UP00093 |
| Motif name: | Klf7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0606067 |
Alignment:
HBDRCCMCGCCCHHHD
BBVGCCBVGGCCTV--
| Original motif | Reverse complement motif |
| Consensus sequence: HBDRCCMCGCCCHHHD | Consensus sequence: DHHDGGGCGRGGKHBH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 14 |
| Similarity score: | 0.0606878 |
Alignment:
HVGVSAGGAGGGTCYHHHHYVH
--------VAGGCCBBGGCVBB
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |
| Motif ID: | UP00087 |
| Motif name: | Tcfap2c_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0608063 |
Alignment:
HWGCCCTDGGGCDB
VAGGCCBBGGCVBB
| Original motif | Reverse complement motif |
| Consensus sequence: BHGCCCDAGGGCWD | Consensus sequence: HWGCCCTDGGGCDB |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0615001 |
Alignment:
DVTTVVCVYGGHGGYCHHDMYB
----BBVGCCBVGGCCTV----
| Original motif | Reverse complement motif |
| Consensus sequence: DVTTVVCVYGGHGGYCHHDMYB | Consensus sequence: VMYDHDGMCCHCCKBGVVAAVH |
 |
 |
| Motif ID: | UP00010 |
| Motif name: | Tcfap2b_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0624288 |
Alignment:
HHKGCCYSRGGCWAD
-BBVGCCBVGGCCTV
| Original motif | Reverse complement motif |
| Consensus sequence: DTWGCCKSMGGCRHH | Consensus sequence: HHKGCCYSRGGCWAD |
 |
 |
| Motif ID: | UP00082 |
| Motif name: | Zfp187_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0627664 |
Alignment:
VHMDGGACVAGGRCDH
VAGGCCBBGGCVBB--
| Original motif | Reverse complement motif |
| Consensus sequence: DDGMCCTBGTCCHYHV | Consensus sequence: VHMDGGACVAGGRCDH |
 |
 |
| Motif ID: | UP00009 |
| Motif name: | Nr2f2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0636529 |
Alignment:
HVBVTGACCCSGBVBV
--VAGGCCBBGGCVBB
| Original motif | Reverse complement motif |
| Consensus sequence: VBVBCSGGGTCAVBBH | Consensus sequence: HVBVTGACCCSGBVBV |
 |
 |
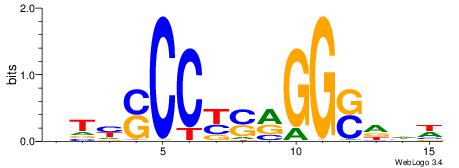
| Dataset #: 2 | Motif ID: 13 | Motif name: Zfp423 |
| Original motif | Reverse complement motif |
| Consensus sequence: GSMMCCYARGGKKKC | Consensus sequence: GYRYCCMTKGGYRSC |
 |
 |
Best Matches for Motif ID 13 (Highest to Lowest)
| Motif ID: | UP00057 |
| Motif name: | Zic2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.0178077 |
Alignment:
BMCCCCCGGGGGGGB
GSMMCCYARGGKKKC
| Original motif | Reverse complement motif |
| Consensus sequence: BMCCCCCGGGGGGGB | Consensus sequence: VCCCCCCCGGGGGRB |
 |
 |
| Motif ID: | UP00006 |
| Motif name: | Zic3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.0213948 |
Alignment:
BMCCCCCGGGGGGGB
GSMMCCYARGGKKKC
| Original motif | Reverse complement motif |
| Consensus sequence: BMCCCCCGGGGGGGB | Consensus sequence: VCCCCCCCGGGGGRB |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0255241 |
Alignment:
BDDHVDKGCCACCCVCGCTDBVH
-------GSMMCCYARGGKKKC-
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Motif ID: | UP00087 |
| Motif name: | Tcfap2c_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.0261399 |
Alignment:
WHHSCCYSRGGSDAD
GSMMCCYARGGKKKC
| Original motif | Reverse complement motif |
| Consensus sequence: WHHSCCYSRGGSDAD | Consensus sequence: DTHSCCKSMGGSHHW |
 |
 |
| Motif ID: | UP00005 |
| Motif name: | Tcfap2a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.0266201 |
Alignment:
WDHSCCYSRGGSRAD
GSMMCCYARGGKKKC
| Original motif | Reverse complement motif |
| Consensus sequence: WDHSCCYSRGGSRAD | Consensus sequence: DTMSCCKSMGGSHDW |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 15 |
| Similarity score: | 0.027503 |
Alignment:
RTHBSYCGCCMCMYVCGBTVDH
-------GSMMCCYARGGKKKC
| Original motif | Reverse complement motif |
| Consensus sequence: RTHBSYCGCCMCMYVCGBTVDH | Consensus sequence: HDBABCGBKRGYGGCGMSBHAK |
 |
 |
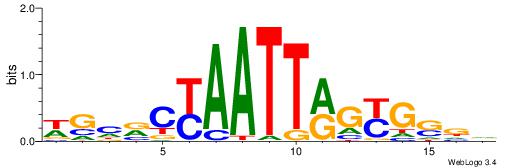
| Motif ID: | UP00220 |
| Motif name: | Nkx1-1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0276935 |
Alignment:
WGHRCYAATTRGYGSDV
-GYRYCCMTKGGYRSC-
| Original motif | Reverse complement motif |
| Consensus sequence: WGHRCYAATTRGYGSDV | Consensus sequence: BHSCMCKAATTMGMDCW |
 |
 |
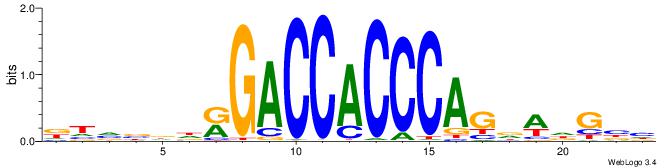
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.0280481 |
Alignment:
DTVBDDRGACCACCCAGDWDGBH
--------GSMMCCYARGGKKKC
| Original motif | Reverse complement motif |
| Consensus sequence: DTVBDDRGACCACCCAGDWDGBH | Consensus sequence: DBCDWHCTGGGTGGTCMDHBBAH |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 15 |
| Similarity score: | 0.0288797 |
Alignment:
HGDHVHKGACCACCCGGBGBGDB
--------GSMMCCYARGGKKKC
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00028 |
| Motif name: | Tcfap2e_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.029693 |
Alignment:
HHHGCCTSAGGCDAD
GSMMCCYARGGKKKC
| Original motif | Reverse complement motif |
| Consensus sequence: HHHGCCTSAGGCDAD | Consensus sequence: DTHGCCTSAGGCHHH |
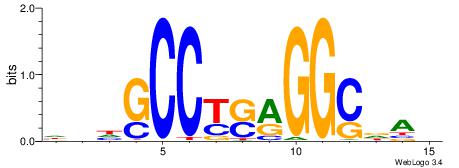 |
 |
| Dataset #: 2 | Motif ID: 14 | Motif name: ZEB1 |
| Original motif | Reverse complement motif |
| Consensus sequence: CACCTD | Consensus sequence: HAGGTG |
 |
 |
Best Matches for Motif ID 14 (Highest to Lowest)
| Motif ID: | UP00068 |
| Motif name: | Eomes_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
VVBVCKACACCTHVBV
-------CACCTD---
| Original motif | Reverse complement motif |
| Consensus sequence: VBVDAGGTGTYGVBBV | Consensus sequence: VVBVCKACACCTHVBV |
 |
 |
| Motif ID: | UP00199 |
| Motif name: | Six4 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 6 |
| Similarity score: | 0.00433068 |
Alignment:
BBVHVGGTGTCATHDDT
---HAGGTG--------
| Original motif | Reverse complement motif |
| Consensus sequence: ADDHATGACACCBHBBV | Consensus sequence: BBVHVGGTGTCATHDDT |
 |
 |
| Motif ID: | UP00068 |
| Motif name: | Eomes_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.00524081 |
Alignment:
HHHTTTCACACCTDHBV
--------CACCTD---
| Original motif | Reverse complement motif |
| Consensus sequence: BVHDAGGTGTGAAAHHH | Consensus sequence: HHHTTTCACACCTDHBV |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.00911301 |
Alignment:
VBHHVCAGGTGCDVDHD
-----HAGGTG------
| Original motif | Reverse complement motif |
| Consensus sequence: VBHHVCAGGTGCDVDHD | Consensus sequence: DHDBHGCACCTGBDDVB |
 |
 |
| Motif ID: | UP00047 |
| Motif name: | Zbtb7b_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0233556 |
Alignment:
BDDRVGACCACCHBDVB
--------CACCTD---
| Original motif | Reverse complement motif |
| Consensus sequence: BDDRVGACCACCHBDVB | Consensus sequence: BBDVHGGTGGTCBKDDB |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 6 |
| Similarity score: | 0.0233928 |
Alignment:
BHCBCBCCGGGTGGTCYHVHDCH
-------HAGGTG----------
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 12 |
| Number of overlap: | 6 |
| Similarity score: | 0.0254338 |
Alignment:
HTBVVVDGGACCACCCRGRDBG
-----------CACCTD-----
| Original motif | Reverse complement motif |
| Consensus sequence: HTBVVVDGGACCACCCRGRDBG | Consensus sequence: CBDMCMGGGTGGTCCHVBVBAH |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 12 |
| Number of overlap: | 6 |
| Similarity score: | 0.0290143 |
Alignment:
HDGHHBHRGACCACCCGSVDVG
-----------CACCTD-----
| Original motif | Reverse complement motif |
| Consensus sequence: HDGHHBHRGACCACCCGSVDVG | Consensus sequence: CVHBSCGGGTGGTCMHVHHCDH |
 |
 |
| Motif ID: | UP00036 |
| Motif name: | Myf6_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.0309673 |
Alignment:
VVDBAACAGBTGBYHB
------HAGGTG----
| Original motif | Reverse complement motif |
| Consensus sequence: VVDBAACAGBTGBYHB | Consensus sequence: BDKVCABCTGTTBDBV |
 |
 |
| Motif ID: | UP00249 |
| Motif name: | Nkx2-5 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.031502 |
Alignment:
HDADCCACTTRAAWTT
-----CACCTD-----
| Original motif | Reverse complement motif |
| Consensus sequence: HDADCCACTTRAAWTT | Consensus sequence: AAWTTMAAGTGGHTDH |
 |
 |
| Dataset #: 2 | Motif ID: 15 | Motif name: TFAP2A |
| Original motif | Reverse complement motif |
| Consensus sequence: GCCBBVRGS | Consensus sequence: SCMVBBGGC |
 |
 |
Best Matches for Motif ID 15 (Highest to Lowest)
| Motif ID: | UP00010 |
| Motif name: | Tcfap2b_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 9 |
| Similarity score: | 0.0115546 |
Alignment:
HHKGCCYSRGGCWAD
---GCCBBVRGS---
| Original motif | Reverse complement motif |
| Consensus sequence: DTWGCCKSMGGCRHH | Consensus sequence: HHKGCCYSRGGCWAD |
 |
 |
| Motif ID: | UP00010 |
| Motif name: | Tcfap2b_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 9 |
| Similarity score: | 0.0150447 |
Alignment:
DYGCCYBARGGCAH
--GCCBBVRGS---
| Original motif | Reverse complement motif |
| Consensus sequence: DYGCCYBARGGCAH | Consensus sequence: HTGCCMTVKGGCMD |
 |
 |
| Motif ID: | UP00005 |
| Motif name: | Tcfap2a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 9 |
| Similarity score: | 0.0151527 |
Alignment:
WDHSCCYSRGGSRAD
---GCCBBVRGS---
| Original motif | Reverse complement motif |
| Consensus sequence: WDHSCCYSRGGSRAD | Consensus sequence: DTMSCCKSMGGSHDW |
 |
 |
| Motif ID: | UP00087 |
| Motif name: | Tcfap2c_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 9 |
| Similarity score: | 0.016909 |
Alignment:
WHHSCCYSRGGSDAD
---GCCBBVRGS---
| Original motif | Reverse complement motif |
| Consensus sequence: WHHSCCYSRGGSDAD | Consensus sequence: DTHSCCKSMGGSHHW |
 |
 |
| Motif ID: | UP00087 |
| Motif name: | Tcfap2c_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 9 |
| Similarity score: | 0.0170259 |
Alignment:
BHGCCCDAGGGCWD
--GCCBBVRGS---
| Original motif | Reverse complement motif |
| Consensus sequence: BHGCCCDAGGGCWD | Consensus sequence: HWGCCCTDGGGCDB |
 |
 |
| Motif ID: | UP00028 |
| Motif name: | Tcfap2e_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 9 |
| Similarity score: | 0.0198854 |
Alignment:
HHHGCCTSAGGCDAD
---GCCBBVRGS---
| Original motif | Reverse complement motif |
| Consensus sequence: HHHGCCTSAGGCDAD | Consensus sequence: DTHGCCTSAGGCHHH |
 |
 |
| Motif ID: | UP00000 |
| Motif name: | Smad3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 9 |
| Similarity score: | 0.0203681 |
Alignment:
BHBHDTGGCGGGGBDHD
SCMVBBGGC--------
| Original motif | Reverse complement motif |
| Consensus sequence: DHHBCCCCGCCAHHBHB | Consensus sequence: BHBHDTGGCGGGGBDHD |
 |
 |
| Motif ID: | UP00003 |
| Motif name: | E2F3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 9 |
| Similarity score: | 0.0217064 |
Alignment:
BBBDYTGGCGCCKWDBD
SCMVBBGGC--------
| Original motif | Reverse complement motif |
| Consensus sequence: HBHWYGGCGCCAMDVBB | Consensus sequence: BBBDYTGGCGCCKWDBD |
 |
 |
| Motif ID: | UP00005 |
| Motif name: | Tcfap2a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 9 |
| Similarity score: | 0.0234775 |
Alignment:
HHRCCBBWGGDCDB
--GCCBBVRGS---
| Original motif | Reverse complement motif |
| Consensus sequence: HHRCCBBWGGDCDB | Consensus sequence: BDGHCCWBVGGKDH |
 |
 |
| Motif ID: | UP00093 |
| Motif name: | Klf7_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 9 |
| Similarity score: | 0.026642 |
Alignment:
VVDBATACGCCCHAHHD
--------GCCBBVRGS
| Original motif | Reverse complement motif |
| Consensus sequence: VVDBATACGCCCHAHHD | Consensus sequence: DHDTHGGGCGTATBHBB |
 |
 |
| Dataset #: 2 | Motif ID: 16 | Motif name: SP1 |
| Original motif | Reverse complement motif |
| Consensus sequence: CCCCKCCCCC | Consensus sequence: GGGGGYGGGG |
 |
 |
Best Matches for Motif ID 16 (Highest to Lowest)
| Motif ID: | UP00021 |
| Motif name: | Zfp281_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0231018 |
Alignment:
DBCCCCCCCCCCMYC
--CCCCKCCCCC---
| Original motif | Reverse complement motif |
| Consensus sequence: DBCCCCCCCCCCMYC | Consensus sequence: GKRGGGGGGGGGGBD |
 |
 |
| Motif ID: | UP00093 |
| Motif name: | Klf7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0303207 |
Alignment:
HBDRCCMCGCCCHHHD
----CCCCKCCCCC--
| Original motif | Reverse complement motif |
| Consensus sequence: HBDRCCMCGCCCHHHD | Consensus sequence: DHHDGGGCGRGGKHBH |
 |
 |
| Motif ID: | UP00033 |
| Motif name: | Zfp410_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0323007 |
Alignment:
HHHHDGGGGCGGDBHDV
----GGGGGYGGGG---
| Original motif | Reverse complement motif |
| Consensus sequence: BHHBHCCGCCCCDHHHH | Consensus sequence: HHHHDGGGGCGGDBHDV |
 |
 |
| Motif ID: | UP00099 |
| Motif name: | Ascl2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0340973 |
Alignment:
HVDVGGGHGGGGDBHB
--GGGGGYGGGG----
| Original motif | Reverse complement motif |
| Consensus sequence: BHVDCCCCDCCCBDBH | Consensus sequence: HVDVGGGHGGGGDBHB |
 |
 |
| Motif ID: | UP00043 |
| Motif name: | Bcl6b_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0351051 |
Alignment:
BHHHWGGGGCGGBBVH
----GGGGGYGGGG--
| Original motif | Reverse complement motif |
| Consensus sequence: HBBBCCGCCCCWHHHV | Consensus sequence: BHHHWGGGGCGGBBVH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0390437 |
Alignment:
HDBABCGBKRGYGGCGMSBHAK
------GGGGGYGGGG------
| Original motif | Reverse complement motif |
| Consensus sequence: RTHBSYCGCCMCMYVCGBTVDH | Consensus sequence: HDBABCGBKRGYGGCGMSBHAK |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0393358 |
Alignment:
HHBSHTCGTGRGTGGKGDBVWMV
-------GGGGGYGGGG------
| Original motif | Reverse complement motif |
| Consensus sequence: VYWVVHCRCCACMCACGAHSBHH | Consensus sequence: HHBSHTCGTGRGTGGKGDBVWMV |
 |
 |
| Motif ID: | UP00000 |
| Motif name: | Smad3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0413863 |
Alignment:
DHHBCCCCGCCAHHBHB
----CCCCKCCCCC---
| Original motif | Reverse complement motif |
| Consensus sequence: DHHBCCCCGCCAHHBHB | Consensus sequence: BHBHDTGGCGGGGBDHD |
 |
 |
| Motif ID: | UP00022 |
| Motif name: | Zfp740_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0418947 |
Alignment:
HVBCCCCCCCCMHHHB
---CCCCKCCCCC---
| Original motif | Reverse complement motif |
| Consensus sequence: HVBCCCCCCCCMHHHB | Consensus sequence: BHHDYGGGGGGGGBVD |
 |
 |
| Motif ID: | UP00002 |
| Motif name: | Sp4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0420708 |
Alignment:
BDHHMCGCCCCCTHVBB
--CCCCKCCCCC-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDHHMCGCCCCCTHVBB | Consensus sequence: BVVHAGGGGGCGRDHHB |
 |
 |
| Dataset #: 2 | Motif ID: 17 | Motif name: RXRAVDR |
| Original motif | Reverse complement motif |
| Consensus sequence: GGGTCAWBGRGTTCA | Consensus sequence: TGAACKCBWTGACCC |
 |
 |
Best Matches for Motif ID 17 (Highest to Lowest)
| Motif ID: | UP00037 |
| Motif name: | Zfp105_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.0631638 |
Alignment:
DYGVKTCAATAABTYDD
--GGGTCAWBGRGTTCA
| Original motif | Reverse complement motif |
| Consensus sequence: DYGVKTCAATAABTYDD | Consensus sequence: HDMAVTTATTGARVCMH |
 |
 |
| Motif ID: | UP00044 |
| Motif name: | Mafk_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.0639668 |
Alignment:
DWAAAWTGCTGACTH
TGAACKCBWTGACCC
| Original motif | Reverse complement motif |
| Consensus sequence: DWAAAWTGCTGACTH | Consensus sequence: HAGTCAGCAWTTTWD |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 15 |
| Similarity score: | 0.0682816 |
Alignment:
BHCBCBCCGGGTGGTCYHVHDCH
--------GGGTCAWBGRGTTCA
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.06995 |
Alignment:
HTBVVVDGGACCACCCRGRDBG
-TGAACKCBWTGACCC------
| Original motif | Reverse complement motif |
| Consensus sequence: HTBVVVDGGACCACCCRGRDBG | Consensus sequence: CBDMCMGGGTGGTCCHVBVBAH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.0711699 |
Alignment:
DVMHHDHKGACCCTCCTSVCBH
-------TGAACKCBWTGACCC
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.072162 |
Alignment:
CVHBSCGGGTGGTCMHVHHCDH
------GGGTCAWBGRGTTCA-
| Original motif | Reverse complement motif |
| Consensus sequence: HDGHHBHRGACCACCCGSVDVG | Consensus sequence: CVHBSCGGGTGGTCMHVHHCDH |
 |
 |
| Motif ID: | UP00045 |
| Motif name: | Mafb_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.562034 |
Alignment:
-HAAWDTGCTGACDWARH
TGAACKCBWTGACCC---
| Original motif | Reverse complement motif |
| Consensus sequence: HAAWDTGCTGACDWARH | Consensus sequence: DMTWDGTCAGCADWTTH |
 |
 |
| Motif ID: | UP00012 |
| Motif name: | Bbx_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.562788 |
Alignment:
HWBTTCAWTGAADTH-
-GGGTCAWBGRGTTCA
| Original motif | Reverse complement motif |
| Consensus sequence: HADTTCAWTGAABWD | Consensus sequence: HWBTTCAWTGAADTH |
 |
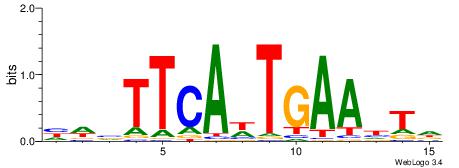 |
| Motif ID: | UP00005 |
| Motif name: | Tcfap2a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.56549 |
Alignment:
-HHRCCBBWGGDCDB
GGGTCAWBGRGTTCA-
| Original motif | Reverse complement motif |
| Consensus sequence: HHRCCBBWGGDCDB | Consensus sequence: BDGHCCWBVGGKDH |
 |
 |
| Motif ID: | UP00067 |
| Motif name: | Lef1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.565777 |
Alignment:
VDDAGATCAAAGGKDDD-
---GGGTCAWBGRGTTCA
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATCTDDV | Consensus sequence: VDDAGATCAAAGGKDDD |
 |
 |
| Dataset #: 2 | Motif ID: 18 | Motif name: RREB1 |
| Original motif | Reverse complement motif |
| Consensus sequence: CCCCMAAMCAMCCMCMMMCV | Consensus sequence: BGRRRGRGGRTGRTTYGGGG |
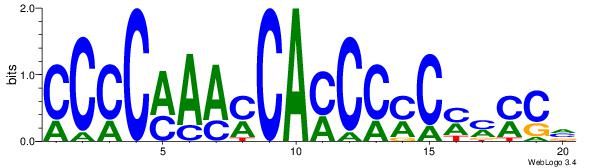 |
 |
Best Matches for Motif ID 18 (Highest to Lowest)
| Motif ID: | UP00540 |
| Motif name: | Gli3_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 20 |
| Similarity score: | 0.0443369 |
Alignment:
YBVDMGTGGGTGGTCKVVBVBBT
BGRRRGRGGRTGRTTYGGGG---
| Original motif | Reverse complement motif |
| Consensus sequence: ABBBBVVRGACCACCCACRDBBM | Consensus sequence: YBVDMGTGGGTGGTCKVVBVBBT |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0490569 |
Alignment:
MHHHWHYTMCCHCCCACVAABVH
--CCCCMAAMCAMCCMCMMMCV-
| Original motif | Reverse complement motif |
| Consensus sequence: MHHHWHYTMCCHCCCACVAABVH | Consensus sequence: DVVTTVGTGGGHGGYAMHWHHHY |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0546172 |
Alignment:
BTBVTCVTGGGTGGTCMVVDVBB
-BGRRRGRGGRTGRTTYGGGG--
| Original motif | Reverse complement motif |
| Consensus sequence: BBBDVVRGACCACCCAVGABBAB | Consensus sequence: BTBVTCVTGGGTGGTCMVVDVBB |
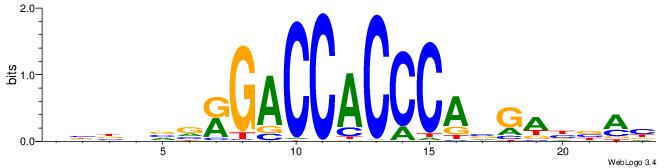 |
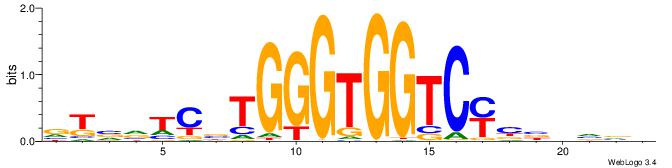 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.055783 |
Alignment:
VYWVVHCRCCACMCACGAHSBHH
-CCCCMAAMCAMCCMCMMMCV--
| Original motif | Reverse complement motif |
| Consensus sequence: VYWVVHCRCCACMCACGAHSBHH | Consensus sequence: HHBSHTCGTGRGTGGKGDBVWMV |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.057969 |
Alignment:
HVBDAGCGBGGGTGGCRDBDDDB
--BGRRRGRGGRTGRTTYGGGG-
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0595101 |
Alignment:
DTVBDDRGACCACCCAGDWDGBH
--CCCCMAAMCAMCCMCMMMCV-
| Original motif | Reverse complement motif |
| Consensus sequence: DTVBDDRGACCACCCAGDWDGBH | Consensus sequence: DBCDWHCTGGGTGGTCMDHBBAH |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0622003 |
Alignment:
BHCBCBCCGGGTGGTCYHVHDCH
-BGRRRGRGGRTGRTTYGGGG--
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0626951 |
Alignment:
RTHBSYCGCCMCMYVCGBTVDH
-CCCCMAAMCAMCCMCMMMCV-
| Original motif | Reverse complement motif |
| Consensus sequence: RTHBSYCGCCMCMYVCGBTVDH | Consensus sequence: HDBABCGBKRGYGGCGMSBHAK |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0631901 |
Alignment:
DVMHHDHKGACCCTCCTSVCBH
CCCCMAAMCAMCCMCMMMCV--
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |
| Motif ID: | UP00528 |
| Motif name: | Foxm1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0673848 |
Alignment:
DCATDBTGHGCATKMTTBRSWR
--BGRRRGRGGRTGRTTYGGGG
| Original motif | Reverse complement motif |
| Consensus sequence: MWSMBAARRATGCDCABHATGD | Consensus sequence: DCATDBTGHGCATKMTTBRSWR |
 |
 |
| Dataset #: 2 | Motif ID: 19 | Motif name: REST |
| Original motif | Reverse complement motif |
| Consensus sequence: TTCAGCACCATGGACAGCKCC | Consensus sequence: GGYGCTGTCCATGGTGCTGAA |
 |
 |
Best Matches for Motif ID 19 (Highest to Lowest)
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 21 |
| Similarity score: | 0.0699178 |
Alignment:
BBBDVVRGACCACCCAVGABBAB
--TTCAGCACCATGGACAGCKCC
| Original motif | Reverse complement motif |
| Consensus sequence: BBBDVVRGACCACCCAVGABBAB | Consensus sequence: BTBVTCVTGGGTGGTCMVVDVBB |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 21 |
| Similarity score: | 0.0734685 |
Alignment:
VMYDHDGMCCHCCKBGVVAAVH
GGYGCTGTCCATGGTGCTGAA-
| Original motif | Reverse complement motif |
| Consensus sequence: DVTTVVCVYGGHGGYCHHDMYB | Consensus sequence: VMYDHDGMCCHCCKBGVVAAVH |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 21 |
| Similarity score: | 0.0740652 |
Alignment:
VYWVVHCRCCACMCACGAHSBHH
-TTCAGCACCATGGACAGCKCC-
| Original motif | Reverse complement motif |
| Consensus sequence: VYWVVHCRCCACMCACGAHSBHH | Consensus sequence: HHBSHTCGTGRGTGGKGDBVWMV |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 21 |
| Similarity score: | 0.0750152 |
Alignment:
MHHHWHYTMCCHCCCACVAABVH
--TTCAGCACCATGGACAGCKCC
| Original motif | Reverse complement motif |
| Consensus sequence: MHHHWHYTMCCHCCCACVAABVH | Consensus sequence: DVVTTVGTGGGHGGYAMHWHHHY |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 21 |
| Similarity score: | 0.0751449 |
Alignment:
BHCBCBCCGGGTGGTCYHVHDCH
GGYGCTGTCCATGGTGCTGAA--
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 21 |
| Similarity score: | 0.075832 |
Alignment:
BDDHVDKGCCACCCVCGCTDBVH
-TTCAGCACCATGGACAGCKCC-
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 21 |
| Similarity score: | 0.0773446 |
Alignment:
DTVBDDRGACCACCCAGDWDGBH
--TTCAGCACCATGGACAGCKCC
| Original motif | Reverse complement motif |
| Consensus sequence: DTVBDDRGACCACCCAGDWDGBH | Consensus sequence: DBCDWHCTGGGTGGTCMDHBBAH |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 21 |
| Similarity score: | 0.0775367 |
Alignment:
YDYBDHTMCACGTGGADDBMDGT
--TTCAGCACCATGGACAGCKCC
| Original motif | Reverse complement motif |
| Consensus sequence: YDYBDHTMCACGTGGADDBMDGT | Consensus sequence: ACHRBHDTCCACGTGYAHHBMHM |
 |
 |
| Motif ID: | UP00528 |
| Motif name: | Foxm1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 21 |
| Similarity score: | 0.077977 |
Alignment:
DCATDBTGHGCATKMTTBRSWR
-GGYGCTGTCCATGGTGCTGAA
| Original motif | Reverse complement motif |
| Consensus sequence: MWSMBAARRATGCDCABHATGD | Consensus sequence: DCATDBTGHGCATKMTTBRSWR |
 |
 |
| Motif ID: | UP00400 |
| Motif name: | Zif268 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 21 |
| Similarity score: | 0.0785969 |
Alignment:
BADHBDHCGCCCMCGCAHHDBBV
--GGYGCTGTCCATGGTGCTGAA
| Original motif | Reverse complement motif |
| Consensus sequence: BADHBDHCGCCCMCGCAHHDBBV | Consensus sequence: VVVDHHTGCGYGGGCGDHVHHTB |
 |
 |
| Dataset #: 2 | Motif ID: 20 | Motif name: PPARGRXRA |
| Original motif | Reverse complement motif |
| Consensus sequence: BTRGGDCARAGGKCA | Consensus sequence: TGRCCTKTGHCCKAB |
 |
 |
Best Matches for Motif ID 20 (Highest to Lowest)
| Motif ID: | UP00067 |
| Motif name: | Lef1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0304954 |
Alignment:
VDDAGATCAAAGGKDDD
-BTRGGDCARAGGKCA-
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATCTDDV | Consensus sequence: VDDAGATCAAAGGKDDD |
 |
 |
| Motif ID: | UP00058 |
| Motif name: | Tcf3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0319475 |
Alignment:
HHHASATCAAAGVRHHH
-BTRGGDCARAGGKCA-
| Original motif | Reverse complement motif |
| Consensus sequence: HHHASATCAAAGVRHHH | Consensus sequence: HHHKVCTTTGATSTHHH |
 |
 |
| Motif ID: | UP00054 |
| Motif name: | Tcf7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.032779 |
Alignment:
BHHASATCAAAGGAHHH
-BTRGGDCARAGGKCA-
| Original motif | Reverse complement motif |
| Consensus sequence: BHHASATCAAAGGAHHH | Consensus sequence: HHHTCCTTTGATSTHHV |
 |
 |
| Motif ID: | UP00082 |
| Motif name: | Zfp187_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0334185 |
Alignment:
VHMDGGACVAGGRCDH
BTRGGDCARAGGKCA-
| Original motif | Reverse complement motif |
| Consensus sequence: DDGMCCTBGTCCHYHV | Consensus sequence: VHMDGGACVAGGRCDH |
 |
 |
| Motif ID: | UP00062 |
| Motif name: | Sox4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0352539 |
Alignment:
HBDDDAACAAAGRVBHH
-BTRGGDCARAGGKCA-
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDAACAAAGRVBHH | Consensus sequence: HHBBMCTTTGTTHDDBH |
 |
 |
| Motif ID: | UP00030 |
| Motif name: | Sox11_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0358782 |
Alignment:
HBDDRAACAAAGRABHH
-BTRGGDCARAGGKCA-
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDRAACAAAGRABHH | Consensus sequence: HHBTMCTTTGTTMDDVH |
 |
 |
| Motif ID: | UP00101 |
| Motif name: | Sox12_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0374729 |
Alignment:
HDTMCTTTGTSTVDBB
TGRCCTKTGHCCKAB-
| Original motif | Reverse complement motif |
| Consensus sequence: VVDBASACAAAGRADH | Consensus sequence: HDTMCTTTGTSTVDBB |
 |
 |
| Motif ID: | UP00083 |
| Motif name: | Tcf7l2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0379757 |
Alignment:
BDDASATCAAAGGMDDD
-BTRGGDCARAGGKCA-
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATSTDDV | Consensus sequence: BDDASATCAAAGGMDDD |
 |
 |
| Motif ID: | UP00093 |
| Motif name: | Klf7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0382733 |
Alignment:
DHHDGGGCGRGGKHBH
BTRGGDCARAGGKCA-
| Original motif | Reverse complement motif |
| Consensus sequence: HBDRCCMCGCCCHHHD | Consensus sequence: DHHDGGGCGRGGKHBH |
 |
 |
| Motif ID: | UP00055 |
| Motif name: | Hbp1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 15 |
| Similarity score: | 0.0397168 |
Alignment:
HBTHVACAATGGGMDHD
BTRGGDCARAGGKCA--
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCCATTGTVHABH | Consensus sequence: HBTHVACAATGGGMDHD |
 |
 |
| Dataset #: 2 | Motif ID: 21 | Motif name: PLAG1 |
| Original motif | Reverse complement motif |
| Consensus sequence: GGGGCCCAAGGGGG | Consensus sequence: CCCCCTTGGGCCCC |
 |
 |
Best Matches for Motif ID 21 (Highest to Lowest)
| Motif ID: | UP00102 |
| Motif name: | Zic1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0499525 |
Alignment:
CCCCCCCGGGGGHB
CCCCCTTGGGCCCC
| Original motif | Reverse complement motif |
| Consensus sequence: BHCCCCCGGGGGGG | Consensus sequence: CCCCCCCGGGGGHB |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0540613 |
Alignment:
BTBVTCVTGGGTGGTCMVVDVBB
---------CCCCCTTGGGCCCC
| Original motif | Reverse complement motif |
| Consensus sequence: BBBDVVRGACCACCCAVGABBAB | Consensus sequence: BTBVTCVTGGGTGGTCMVVDVBB |
 |
 |
| Motif ID: | UP00057 |
| Motif name: | Zic2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0548411 |
Alignment:
BMCCCCCGGGGGGGB
-CCCCCTTGGGCCCC
| Original motif | Reverse complement motif |
| Consensus sequence: BMCCCCCGGGGGGGB | Consensus sequence: VCCCCCCCGGGGGRB |
 |
 |
| Motif ID: | UP00005 |
| Motif name: | Tcfap2a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0556785 |
Alignment:
BDGHCCWBVGGKDH
GGGGCCCAAGGGGG
| Original motif | Reverse complement motif |
| Consensus sequence: HHRCCBBWGGDCDB | Consensus sequence: BDGHCCWBVGGKDH |
 |
 |
| Motif ID: | UP00006 |
| Motif name: | Zic3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.056976 |
Alignment:
VCCCCCCCGGGGGRB
-CCCCCTTGGGCCCC
| Original motif | Reverse complement motif |
| Consensus sequence: BMCCCCCGGGGGGGB | Consensus sequence: VCCCCCCCGGGGGRB |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 14 |
| Similarity score: | 0.0573495 |
Alignment:
HDGHHBHRGACCACCCGSVDVG
CCCCCTTGGGCCCC--------
| Original motif | Reverse complement motif |
| Consensus sequence: HDGHHBHRGACCACCCGSVDVG | Consensus sequence: CVHBSCGGGTGGTCMHVHHCDH |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 14 |
| Similarity score: | 0.0579022 |
Alignment:
VMYDHDGMCCHCCKBGVVAAVH
--------CCCCCTTGGGCCCC
| Original motif | Reverse complement motif |
| Consensus sequence: DVTTVVCVYGGHGGYCHHDMYB | Consensus sequence: VMYDHDGMCCHCCKBGVVAAVH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 14 |
| Similarity score: | 0.0580662 |
Alignment:
DVMHHDHKGACCCTCCTSVCBH
CCCCCTTGGGCCCC--------
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0597453 |
Alignment:
HGDHVHKGACCACCCGGBGBGDB
---------CCCCCTTGGGCCCC
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00021 |
| Motif name: | Zfp281_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0598088 |
Alignment:
DBCCCCCCCCCCMYC
-CCCCCTTGGGCCCC
| Original motif | Reverse complement motif |
| Consensus sequence: DBCCCCCCCCCCMYC | Consensus sequence: GKRGGGGGGGGGGBD |
 |
 |
| Dataset #: 2 | Motif ID: 22 | Motif name: NR2F1 |
| Original motif | Reverse complement motif |
| Consensus sequence: TGAMCTTTGMMCYT | Consensus sequence: AKGYYCAAAGRTCA |
 |
 |
Best Matches for Motif ID 22 (Highest to Lowest)
| Motif ID: | UP00083 |
| Motif name: | Tcf7l2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0452364 |
Alignment:
DDDYCCTTTGATSTDDV
-TGAMCTTTGMMCYT--
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATSTDDV | Consensus sequence: BDDASATCAAAGGMDDD |
 |
 |
| Motif ID: | UP00067 |
| Motif name: | Lef1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0453449 |
Alignment:
VDDAGATCAAAGGKDDD
--AKGYYCAAAGRTCA-
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATCTDDV | Consensus sequence: VDDAGATCAAAGGKDDD |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0461358 |
Alignment:
DVMHHDHKGACCCTCCTSVCBH
-------TGAMCTTTGMMCYT-
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |
| Motif ID: | UP00054 |
| Motif name: | Tcf7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0461768 |
Alignment:
BHHASATCAAAGGAHHH
--AKGYYCAAAGRTCA-
| Original motif | Reverse complement motif |
| Consensus sequence: BHHASATCAAAGGAHHH | Consensus sequence: HHHTCCTTTGATSTHHV |
 |
 |
| Motif ID: | UP00058 |
| Motif name: | Tcf3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0473628 |
Alignment:
HHHASATCAAAGVRHHH
--AKGYYCAAAGRTCA-
| Original motif | Reverse complement motif |
| Consensus sequence: HHHASATCAAAGVRHHH | Consensus sequence: HHHKVCTTTGATSTHHH |
 |
 |
| Motif ID: | UP00082 |
| Motif name: | Zfp187_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0480188 |
Alignment:
DDGMCCTBGTCCHYHV
TGAMCTTTGMMCYT--
| Original motif | Reverse complement motif |
| Consensus sequence: DDGMCCTBGTCCHYHV | Consensus sequence: VHMDGGACVAGGRCDH |
 |
 |
| Motif ID: | UP00062 |
| Motif name: | Sox4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0495447 |
Alignment:
HHBBMCTTTGTTHDDBH
-TGAMCTTTGMMCYT--
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDAACAAAGRVBHH | Consensus sequence: HHBBMCTTTGTTHDDBH |
 |
 |
| Motif ID: | UP00030 |
| Motif name: | Sox11_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0499033 |
Alignment:
HHBTMCTTTGTTMDDVH
-TGAMCTTTGMMCYT--
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDRAACAAAGRABHH | Consensus sequence: HHBTMCTTTGTTMDDVH |
 |
 |
| Motif ID: | UP00023 |
| Motif name: | Sox30_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0511534 |
Alignment:
DHDBAACAATDRHHHH
--TGAMCTTTGMMCYT
| Original motif | Reverse complement motif |
| Consensus sequence: DHDBAACAATDRHHHH | Consensus sequence: HHHHMHATTGTTBDHD |
 |
 |
| Motif ID: | UP00005 |
| Motif name: | Tcfap2a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0516973 |
Alignment:
BDGHCCWBVGGKDH
AKGYYCAAAGRTCA
| Original motif | Reverse complement motif |
| Consensus sequence: HHRCCBBWGGDCDB | Consensus sequence: BDGHCCWBVGGKDH |
 |
 |
| Dataset #: 2 | Motif ID: 23 | Motif name: MZF1_1-4 |
| Original motif | Reverse complement motif |
| Consensus sequence: BGGGGA | Consensus sequence: TCCCCV |
 |
 |
Best Matches for Motif ID 23 (Highest to Lowest)
| Motif ID: | UP00176 |
| Motif name: | Crx |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
YBWDGGGGATTARCCD
---BGGGGA-------
| Original motif | Reverse complement motif |
| Consensus sequence: YBWDGGGGATTARCCD | Consensus sequence: DGGMTAATCCCCDWBK |
 |
 |
| Motif ID: | UP00265 |
| Motif name: | Pitx3 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 6 |
| Similarity score: | 0.00193291 |
Alignment:
GBHDGCTAATCCCCCB
---------TCCCCV-
| Original motif | Reverse complement motif |
| Consensus sequence: VGGGGGATTAGCDDBC | Consensus sequence: GBHDGCTAATCCCCCB |
 |
 |
| Motif ID: | UP00239 |
| Motif name: | Obox2 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.00284735 |
Alignment:
HTARTTAATCCCCCKBD
--------TCCCCV---
| Original motif | Reverse complement motif |
| Consensus sequence: DBRGGGGGATTAAMTAH | Consensus sequence: HTARTTAATCCCCCKBD |
 |
 |
| Motif ID: | UP00216 |
| Motif name: | Obox1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 6 |
| Similarity score: | 0.00461976 |
Alignment:
DHDGTTAATCCCCKTDD
--------TCCCCV---
| Original motif | Reverse complement motif |
| Consensus sequence: DDARGGGGATTAACDHH | Consensus sequence: DHDGTTAATCCCCKTDD |
 |
 |
| Motif ID: | UP00160 |
| Motif name: | Obox3 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.00641818 |
Alignment:
HHWGTTAATCCCCCBBH
--------TCCCCV---
| Original motif | Reverse complement motif |
| Consensus sequence: HBVGGGGGATTAACWHH | Consensus sequence: HHWGTTAATCCCCCBBH |
 |
 |
| Motif ID: | UP00208 |
| Motif name: | Obox5_2284.1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 6 |
| Similarity score: | 0.0110256 |
Alignment:
HDVRGGGATTAAHTHHH
--BGGGGA---------
| Original motif | Reverse complement motif |
| Consensus sequence: HDVRGGGATTAAHTHHH | Consensus sequence: DHHAHTTAATCCCKVDH |
 |
 |
| Motif ID: | UP00143 |
| Motif name: | Dobox5 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.0136568 |
Alignment:
DDHVGGGATTAATHDTH
--BGGGGA---------
| Original motif | Reverse complement motif |
| Consensus sequence: DDHVGGGATTAATHDTH | Consensus sequence: DADHATTAATCCCBHHH |
 |
 |
| Motif ID: | UP00229 |
| Motif name: | Otx1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.0138939 |
Alignment:
DDDRGGGATTAATWHHB
--BGGGGA---------
| Original motif | Reverse complement motif |
| Consensus sequence: DDDRGGGATTAATWHHB | Consensus sequence: VHHWATTAATCCCMDHH |
 |
 |
| Motif ID: | UP00267 |
| Motif name: | Otx2 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 6 |
| Similarity score: | 0.016387 |
Alignment:
DDDRGGGATTAATHDDH
--BGGGGA---------
| Original motif | Reverse complement motif |
| Consensus sequence: DDDRGGGATTAATHDDH | Consensus sequence: DDHHATTAATCCCKDHD |
 |
 |
| Motif ID: | UP00153 |
| Motif name: | Pitx1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0164951 |
Alignment:
HBVDRGGGATTAAMDHH
---BGGGGA--------
| Original motif | Reverse complement motif |
| Consensus sequence: HBVDRGGGATTAAMDHH | Consensus sequence: HHDRTTAATCCCKHBVH |
 |
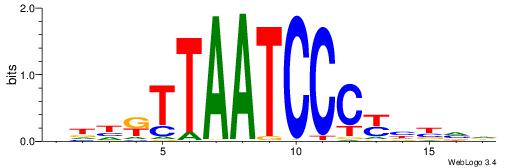 |
| Dataset #: 2 | Motif ID: 24 | Motif name: Ar |
| Original motif | Reverse complement motif |
| Consensus sequence: HWDAGHACRHHVTGTHCCHVMV | Consensus sequence: VRVDGGHACAVDDKGTHCTDWH |
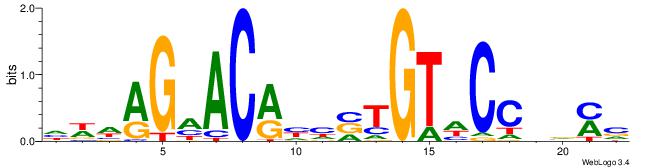 |
 |
Best Matches for Motif ID 24 (Highest to Lowest)
| Motif ID: | UP00032 |
| Motif name: | Gata3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 22 |
| Similarity score: | 0.0676991 |
Alignment:
HBHDVBWGATHBTATCRVBHHH
HWDAGHACRHHVTGTHCCHVMV
| Original motif | Reverse complement motif |
| Consensus sequence: HBHDVBWGATHBTATCRVBHHH | Consensus sequence: HHHBBMGATAVHATCWVVDHVH |
 |
 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 22 |
| Similarity score: | 0.0692146 |
Alignment:
VVYDDDCHCAVACAATTDBVAC
VRVDGGHACAVDDKGTHCTDWH
| Original motif | Reverse complement motif |
| Consensus sequence: GTVBDAATTGTVTGHGDDHKVB | Consensus sequence: VVYDDDCHCAVACAATTDBVAC |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 22 |
| Similarity score: | 0.0695612 |
Alignment:
DDHBHBSTCACGTGAHBBDHHM
VRVDGGHACAVDDKGTHCTDWH
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 22 |
| Similarity score: | 0.0771346 |
Alignment:
YDYBDHTMCACGTGGADDBMDGT
-HWDAGHACRHHVTGTHCCHVMV
| Original motif | Reverse complement motif |
| Consensus sequence: YDYBDHTMCACGTGGADDBMDGT | Consensus sequence: ACHRBHDTCCACGTGYAHHBMHM |
 |
 |
| Motif ID: | UP00526 |
| Motif name: | Foxn1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 22 |
| Similarity score: | 0.0772045 |
Alignment:
HDWHMHGACGCGCGTRBVVBHW
VRVDGGHACAVDDKGTHCTDWH
| Original motif | Reverse complement motif |
| Consensus sequence: WHVBVVMACGCGCGTCDYHWDH | Consensus sequence: HDWHMHGACGCGCGTRBVVBHW |
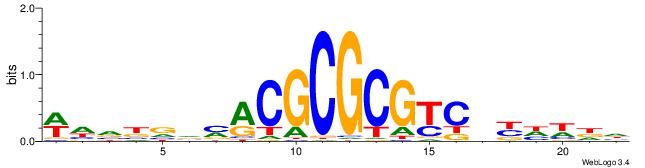 |
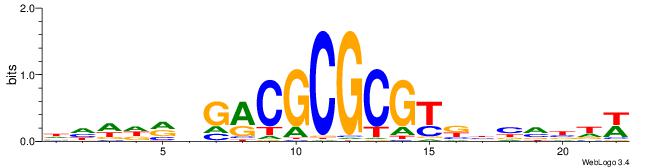 |
| Motif ID: | UP00526 |
| Motif name: | Foxn1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 22 |
| Similarity score: | 0.077433 |
Alignment:
DDRMHBACGCGYGCGTDGBBDB
VRVDGGHACAVDDKGTHCTDWH
| Original motif | Reverse complement motif |
| Consensus sequence: DDRMHBACGCGYGCGTDGBBDB | Consensus sequence: BDVBCDACGCKCGCGTBHRKHD |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 22 |
| Similarity score: | 0.0776897 |
Alignment:
VYWVVHCRCCACMCACGAHSBHH
-HWDAGHACRHHVTGTHCCHVMV
| Original motif | Reverse complement motif |
| Consensus sequence: VYWVVHCRCCACMCACGAHSBHH | Consensus sequence: HHBSHTCGTGRGTGGKGDBVWMV |
 |
 |
| Motif ID: | UP00528 |
| Motif name: | Foxm1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 22 |
| Similarity score: | 0.0787977 |
Alignment:
MWSMBAARRATGCDCABHATGD
HWDAGHACRHHVTGTHCCHVMV
| Original motif | Reverse complement motif |
| Consensus sequence: MWSMBAARRATGCDCABHATGD | Consensus sequence: DCATDBTGHGCATKMTTBRSWR |
 |
 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 22 |
| Similarity score: | 0.0792723 |
Alignment:
HHBDHDDAACAATDRHHHHHBW
HWDAGHACRHHVTGTHCCHVMV
| Original motif | Reverse complement motif |
| Consensus sequence: HHBDHDDAACAATDRHHHHHBW | Consensus sequence: WBHHHHHMDATTGTTHDHDVHH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 22 |
| Similarity score: | 0.0799314 |
Alignment:
HVGVSAGGAGGGTCYHHHHYVH
HWDAGHACRHHVTGTHCCHVMV
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |
| Dataset #: 2 | Motif ID: 25 | Motif name: Arnt |
| Original motif | Reverse complement motif |
| Consensus sequence: CACGTG | Consensus sequence: CACGTG |
 |
 |
Best Matches for Motif ID 25 (Highest to Lowest)
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
RDHDBVDTCACGTGASBHVHDH
--------CACGTG--------
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0026425 |
Alignment:
YDYBDHTMCACGTGGADDBMDGT
--------CACGTG---------
| Original motif | Reverse complement motif |
| Consensus sequence: YDYBDHTMCACGTGGADDBMDGT | Consensus sequence: ACHRBHDTCCACGTGYAHHBMHM |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.0044422 |
Alignment:
HHVBABCACGTGSTHD
------CACGTG----
| Original motif | Reverse complement motif |
| Consensus sequence: DDASCACGTGBTBVDD | Consensus sequence: HHVBABCACGTGSTHD |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0.0342153 |
Alignment:
DHDBHGCACCTGBDDVB
------CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: VBHHVCAGGTGCDVDHD | Consensus sequence: DHDBHGCACCTGBDDVB |
 |
 |
| Motif ID: | UP00097 |
| Motif name: | Mtf1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0471928 |
Alignment:
HDYTTGCACACGDHBH
--------CACGTG--
| Original motif | Reverse complement motif |
| Consensus sequence: DBDHCGTGTGCAAMDH | Consensus sequence: HDYTTGCACACGDHBH |
 |
 |
| Motif ID: | UP00249 |
| Motif name: | Nkx2-5 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.0482749 |
Alignment:
HDADCCACTTRAAWTT
-----CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: HDADCCACTTRAAWTT | Consensus sequence: AAWTTMAAGTGGHTDH |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.0485437 |
Alignment:
BBGHCACGCGACDD
----CACGTG----
| Original motif | Reverse complement motif |
| Consensus sequence: BBGHCACGCGACDD | Consensus sequence: HDGTCGCGTGDCVB |
 |
 |
| Motif ID: | UP00231 |
| Motif name: | Nkx2-2 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0.0497234 |
Alignment:
DVDTKTCAAGTGGKBDH
------CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: HDVRCCACTTGARADBD | Consensus sequence: DVDTKTCAAGTGGKBDH |
 |
 |
| Motif ID: | UP00165 |
| Motif name: | Titf1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.0498919 |
Alignment:
AAHYTCAAGTGGBTBV
-----CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: BVABCCACTTGAMHTT | Consensus sequence: AAHYTCAAGTGGBTBV |
 |
 |
| Motif ID: | UP00147 |
| Motif name: | Nkx2-6 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.0544601 |
Alignment:
DDADCCACTTAAVHTT
-----CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDADCCACTTAAVHTT | Consensus sequence: AAHVTTAAGTGGHTDD |
 |
 |
| Dataset #: 2 | Motif ID: 26 | Motif name: ArntAhr |
| Original motif | Reverse complement motif |
| Consensus sequence: YGCGTG | Consensus sequence: CACGCM |
 |
 |
Best Matches for Motif ID 26 (Highest to Lowest)
| Motif ID: | UP00400 |
| Motif name: | Zif268 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
VVVDHHTGCGYGGGCGDHVHHTB
------YGCGTG-----------
| Original motif | Reverse complement motif |
| Consensus sequence: BADHBDHCGCCCMCGCAHHDBBV | Consensus sequence: VVVDHHTGCGYGGGCGDHVHHTB |
 |
 |
| Motif ID: | UP00002 |
| Motif name: | Sp4_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.00183587 |
Alignment:
HHDGBCACGCCTWDB
-----CACGCM----
| Original motif | Reverse complement motif |
| Consensus sequence: BDWAGGCGTGBCHHD | Consensus sequence: HHDGBCACGCCTWDB |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.00348048 |
Alignment:
HDGTCGCGTGDCVB
----YGCGTG----
| Original motif | Reverse complement motif |
| Consensus sequence: BBGHCACGCGACDD | Consensus sequence: HDGTCGCGTGDCVB |
 |
 |
| Motif ID: | UP00527 |
| Motif name: | Foxn4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0.00526158 |
Alignment:
TCDAMVBGACGCMAKDDDVBBD
-------CACGCM---------
| Original motif | Reverse complement motif |
| Consensus sequence: DBBVDDDRTRGCGTCBBYTHGA | Consensus sequence: TCDAMVBGACGCMAKDDDVBBD |
 |
 |
| Motif ID: | UP00042 |
| Motif name: | Gm397_second |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0.00558171 |
Alignment:
HBGYGTGTGTGCVRVD
---YGCGTG-------
| Original motif | Reverse complement motif |
| Consensus sequence: DVMVGCACACACKCVH | Consensus sequence: HBGYGTGTGTGCVRVD |
 |
 |
| Motif ID: | UP00097 |
| Motif name: | Mtf1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.00702416 |
Alignment:
HDYTTGCACACGDHBH
--------CACGCM--
| Original motif | Reverse complement motif |
| Consensus sequence: DBDHCGTGTGCAAMDH | Consensus sequence: HDYTTGCACACGDHBH |
 |
 |
| Motif ID: | UP00526 |
| Motif name: | Foxn1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0.0123233 |
Alignment:
DDRMHBACGCGYGCGTDGBBDB
-------YGCGTG---------
| Original motif | Reverse complement motif |
| Consensus sequence: DDRMHBACGCGYGCGTDGBBDB | Consensus sequence: BDVBCDACGCKCGCGTBHRKHD |
 |
 |
| Motif ID: | UP00007 |
| Motif name: | Egr1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 6 |
| Similarity score: | 0.0141579 |
Alignment:
HCCGCCCCCGCAHB
------CACGCM--
| Original motif | Reverse complement motif |
| Consensus sequence: HCCGCCCCCGCAHB | Consensus sequence: VHTGCGGGGGCGGH |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 6 |
| Similarity score: | 0.0144485 |
Alignment:
BHCBCBCCGGGTGGTCYHVHDCH
-------YGCGTG----------
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00527 |
| Motif name: | Foxn4_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0.0172109 |
Alignment:
VHYTCCDWCCGCGTCMMTHWHV
---------YGCGTG-------
| Original motif | Reverse complement motif |
| Consensus sequence: VDWDARRGACGCGGWHGGAKHV | Consensus sequence: VHYTCCDWCCGCGTCMMTHWHV |
 |
 |
| Dataset #: 2 | Motif ID: 27 | Motif name: E2F1 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTTSGCGC | Consensus sequence: GCGCSAAA |
 |
 |
Best Matches for Motif ID 27 (Highest to Lowest)
| Motif ID: | UP00001 |
| Motif name: | E2F2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
BBBDTTGGCGCCKWVVD
---TTTSGCGC------
| Original motif | Reverse complement motif |
| Consensus sequence: HVBWYGGCGCCAADVBB | Consensus sequence: BBBDTTGGCGCCKWVVD |
 |
 |
| Motif ID: | UP00003 |
| Motif name: | E2F3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.00182028 |
Alignment:
HBHWYGGCGCCAMDVBB
------GCGCSAAA---
| Original motif | Reverse complement motif |
| Consensus sequence: HBHWYGGCGCCAMDVBB | Consensus sequence: BBBDYTGGCGCCKWDBD |
 |
 |
| Motif ID: | UP00072 |
| Motif name: | IRC900814_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0237639 |
Alignment:
DDHTACGACAAAWDDB
----GCGCSAAA----
| Original motif | Reverse complement motif |
| Consensus sequence: DDHTACGACAAAWDDB | Consensus sequence: BHDWTTTGTCGTAHDD |
 |
 |
| Motif ID: | UP00003 |
| Motif name: | E2F3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 8 |
| Similarity score: | 0.0275636 |
Alignment:
WHSGCGCGCCHTDHB
-----GCGCSAAA--
| Original motif | Reverse complement motif |
| Consensus sequence: VHDADGGCGCGCSHW | Consensus sequence: WHSGCGCGCCHTDHB |
 |
 |
| Motif ID: | UP00065 |
| Motif name: | Zfp161_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0.028109 |
Alignment:
DMMDGMGCGCGCGCHD
--------GCGCSAAA
| Original motif | Reverse complement motif |
| Consensus sequence: DDGCGCGCGCRCHYRD | Consensus sequence: DMMDGMGCGCGCGCHD |
 |
 |
| Motif ID: | UP00001 |
| Motif name: | E2F2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.0296302 |
Alignment:
VHDARGGCGCGCVHH
--TTTSGCGC-----
| Original motif | Reverse complement motif |
| Consensus sequence: VHDARGGCGCGCVHH | Consensus sequence: HHVGCGCGCCKTDHB |
 |
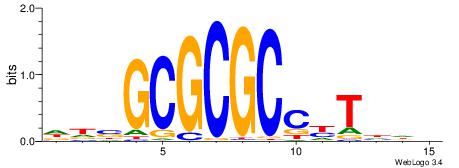 |
| Motif ID: | UP00068 |
| Motif name: | Eomes_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0348969 |
Alignment:
BVHDAGGTGTGAAAHHH
------GCGCSAAA---
| Original motif | Reverse complement motif |
| Consensus sequence: BVHDAGGTGTGAAAHHH | Consensus sequence: HHHTTTCACACCTDHBV |
 |
 |
| Motif ID: | UP00011 |
| Motif name: | Irf6_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.0368278 |
Alignment:
DHDBTKGTTTCGHTHDD
-------TTTSGCGC--
| Original motif | Reverse complement motif |
| Consensus sequence: HDDAHCGAAACYAVDDD | Consensus sequence: DHDBTKGTTTCGHTHDD |
 |
 |
| Motif ID: | UP00049 |
| Motif name: | Sp100_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.0383813 |
Alignment:
DWTTTCCGHHWAWD
-----GCGCSAAA-
| Original motif | Reverse complement motif |
| Consensus sequence: DWTWHHCGGAAAWD | Consensus sequence: DWTTTCCGHHWAWD |
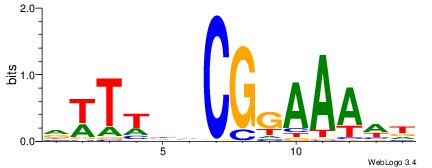 |
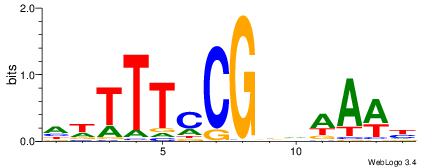 |
| Motif ID: | UP00527 |
| Motif name: | Foxn4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0.0406002 |
Alignment:
TCDAMVBGACGCMAKDDDVBBD
--------GCGCSAAA------
| Original motif | Reverse complement motif |
| Consensus sequence: DBBVDDDRTRGCGTCBBYTHGA | Consensus sequence: TCDAMVBGACGCMAKDDDVBBD |
 |
 |
| Dataset #: 2 | Motif ID: 28 | Motif name: Egr1 |
| Original motif | Reverse complement motif |
| Consensus sequence: HGCGTGGGCGK | Consensus sequence: YCGCCCACGCH |
 |
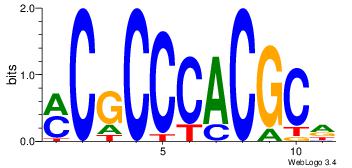 |
Best Matches for Motif ID 28 (Highest to Lowest)
| Motif ID: | UP00400 |
| Motif name: | Zif268 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0 |
Alignment:
VVVDHHTGCGYGGGCGDHVHHTB
------HGCGTGGGCGK------
| Original motif | Reverse complement motif |
| Consensus sequence: BADHBDHCGCCCMCGCAHHDBBV | Consensus sequence: VVVDHHTGCGYGGGCGDHVHHTB |
 |
 |
| Motif ID: | UP00007 |
| Motif name: | Egr1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.00727533 |
Alignment:
HCCGCCCCCGCAHB
-YCGCCCACGCH--
| Original motif | Reverse complement motif |
| Consensus sequence: HCCGCCCCCGCAHB | Consensus sequence: VHTGCGGGGGCGGH |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0262579 |
Alignment:
MHHHWHYTMCCHCCCACVAABVH
---------YCGCCCACGCH---
| Original motif | Reverse complement motif |
| Consensus sequence: MHHHWHYTMCCHCCCACVAABVH | Consensus sequence: DVVTTVGTGGGHGGYAMHWHHHY |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0297053 |
Alignment:
YBVDMGTGGGTGGTCKVVBVBBT
--HGCGTGGGCGK----------
| Original motif | Reverse complement motif |
| Consensus sequence: ABBBBVVRGACCACCCACRDBBM | Consensus sequence: YBVDMGTGGGTGGTCKVVBVBBT |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0318144 |
Alignment:
HVBDAGCGBGGGTGGCRDBDDDB
----HGCGTGGGCGK--------
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Motif ID: | UP00002 |
| Motif name: | Sp4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0345009 |
Alignment:
BVVHAGGGGGCGRDHHB
--HGCGTGGGCGK----
| Original motif | Reverse complement motif |
| Consensus sequence: BDHHMCGCCCCCTHVBB | Consensus sequence: BVVHAGGGGGCGRDHHB |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0353502 |
Alignment:
BTBVTCVTGGGTGGTCMVVDVBB
---HGCGTGGGCGK---------
| Original motif | Reverse complement motif |
| Consensus sequence: BBBDVVRGACCACCCAVGABBAB | Consensus sequence: BTBVTCVTGGGTGGTCMVVDVBB |
 |
 |
| Motif ID: | UP00007 |
| Motif name: | Egr1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0360229 |
Alignment:
BHHDTCCCACTCBDBV
--YCGCCCACGCH---
| Original motif | Reverse complement motif |
| Consensus sequence: BBHBGAGTGGGAHHDB | Consensus sequence: BHHDTCCCACTCBDBV |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0362743 |
Alignment:
VYWVVHCRCCACMCACGAHSBHH
--------YCGCCCACGCH----
| Original motif | Reverse complement motif |
| Consensus sequence: VYWVVHCRCCACMCACGAHSBHH | Consensus sequence: HHBSHTCGTGRGTGGKGDBVWMV |
 |
 |
| Motif ID: | UP00097 |
| Motif name: | Mtf1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0386944 |
Alignment:
HDYTTGCACACGDHBH
---YCGCCCACGCH--
| Original motif | Reverse complement motif |
| Consensus sequence: DBDHCGTGTGCAAMDH | Consensus sequence: HDYTTGCACACGDHBH |
 |
 |
| Dataset #: 2 | Motif ID: 29 | Motif name: ESR1 |
| Original motif | Reverse complement motif |
| Consensus sequence: VDBHMAGGTCACCCTGACCY | Consensus sequence: MGGTCAGGGTGACCTRDBHV |
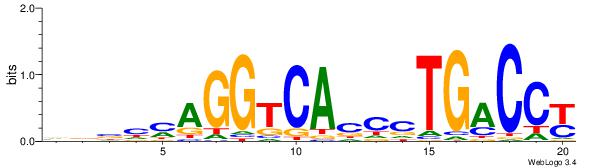 |
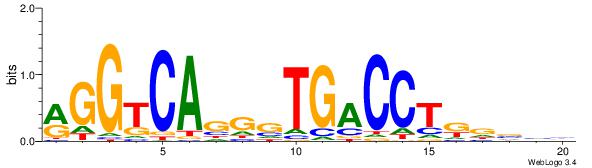 |
Best Matches for Motif ID 29 (Highest to Lowest)
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 20 |
| Similarity score: | 0.0658405 |
Alignment:
BDDHVDKGCCACCCVCGCTDBVH
VDBHMAGGTCACCCTGACCY---
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Motif ID: | UP00098 |
| Motif name: | Rfx3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0689748 |
Alignment:
VCTBHBVCTTGGWTACVVVVVDB
VDBHMAGGTCACCCTGACCY---
| Original motif | Reverse complement motif |
| Consensus sequence: VCTBHBVCTTGGWTACVVVVVDB | Consensus sequence: BDBBVBVGTAWCCAAGVBHBAGB |
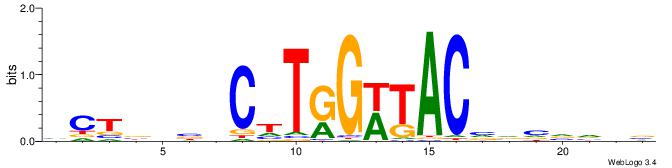 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0701247 |
Alignment:
CBDMCMGGGTGGTCCHVBVBAH
MGGTCAGGGTGACCTRDBHV--
| Original motif | Reverse complement motif |
| Consensus sequence: HTBVVVDGGACCACCCRGRDBG | Consensus sequence: CBDMCMGGGTGGTCCHVBVBAH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0711267 |
Alignment:
HVGVSAGGAGGGTCYHHHHYVH
MGGTCAGGGTGACCTRDBHV--
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0742427 |
Alignment:
CVHBSCGGGTGGTCMHVHHCDH
-VDBHMAGGTCACCCTGACCY-
| Original motif | Reverse complement motif |
| Consensus sequence: HDGHHBHRGACCACCCGSVDVG | Consensus sequence: CVHBSCGGGTGGTCMHVHHCDH |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0747053 |
Alignment:
ABBBBVVRGACCACCCACRDBBM
--VDBHMAGGTCACCCTGACCY-
| Original motif | Reverse complement motif |
| Consensus sequence: ABBBBVVRGACCACCCACRDBBM | Consensus sequence: YBVDMGTGGGTGGTCKVVBVBBT |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 20 |
| Similarity score: | 0.0749165 |
Alignment:
BHCBCBCCGGGTGGTCYHVHDCH
---VDBHMAGGTCACCCTGACCY
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0759307 |
Alignment:
BBBDVVRGACCACCCAVGABBAB
-VDBHMAGGTCACCCTGACCY--
| Original motif | Reverse complement motif |
| Consensus sequence: BBBDVVRGACCACCCAVGABBAB | Consensus sequence: BTBVTCVTGGGTGGTCMVVDVBB |
 |
 |
| Motif ID: | UP00400 |
| Motif name: | Zif268 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0762689 |
Alignment:
VVVDHHTGCGYGGGCGDHVHHTB
-MGGTCAGGGTGACCTRDBHV--
| Original motif | Reverse complement motif |
| Consensus sequence: BADHBDHCGCCCMCGCAHHDBBV | Consensus sequence: VVVDHHTGCGYGGGCGDHVHHTB |
 |
 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0780113 |
Alignment:
GTVBDAATTGTVTGHGDDHKVB
VDBHMAGGTCACCCTGACCY--
| Original motif | Reverse complement motif |
| Consensus sequence: GTVBDAATTGTVTGHGDDHKVB | Consensus sequence: VVYDDDCHCAVACAATTDBVAC |
 |
 |
| Dataset #: 2 | Motif ID: 30 | Motif name: ESR2 |
| Original motif | Reverse complement motif |
| Consensus sequence: VHRGGTCABDBTGMCCTB | Consensus sequence: BAGGYCABHBTGACCKHV |
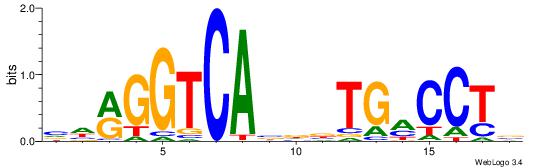 |
 |
Best Matches for Motif ID 30 (Highest to Lowest)
| Motif ID: | UP00034 |
| Motif name: | Sox7_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 18 |
| Similarity score: | 0.0443894 |
Alignment:
GTVBDAATTGTVTGHGDDHKVB
---VHRGGTCABDBTGMCCTB-
| Original motif | Reverse complement motif |
| Consensus sequence: GTVBDAATTGTVTGHGDDHKVB | Consensus sequence: VVYDDDCHCAVACAATTDBVAC |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 18 |
| Similarity score: | 0.049219 |
Alignment:
BDDHVDKGCCACCCVCGCTDBVH
---VHRGGTCABDBTGMCCTB--
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0509182 |
Alignment:
DDHBHBSTCACGTGAHBBDHHM
--VHRGGTCABDBTGMCCTB--
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 18 |
| Similarity score: | 0.0527075 |
Alignment:
DVTTVVCVYGGHGGYCHHDMYB
-BAGGYCABHBTGACCKHV---
| Original motif | Reverse complement motif |
| Consensus sequence: DVTTVVCVYGGHGGYCHHDMYB | Consensus sequence: VMYDHDGMCCHCCKBGVVAAVH |
 |
 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 18 |
| Similarity score: | 0.0541044 |
Alignment:
WBHHHHHMDATTGTTHDHDVHH
-BAGGYCABHBTGACCKHV---
| Original motif | Reverse complement motif |
| Consensus sequence: HHBDHDDAACAATDRHHHHHBW | Consensus sequence: WBHHHHHMDATTGTTHDHDVHH |
 |
 |
| Motif ID: | UP00098 |
| Motif name: | Rfx3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0541108 |
Alignment:
VCTBHBVCTTGGWTACVVVVVDB
--BAGGYCABHBTGACCKHV---
| Original motif | Reverse complement motif |
| Consensus sequence: VCTBHBVCTTGGWTACVVVVVDB | Consensus sequence: BDBBVBVGTAWCCAAGVBHBAGB |
 |
 |
| Motif ID: | UP00527 |
| Motif name: | Foxn4_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 18 |
| Similarity score: | 0.0543733 |
Alignment:
VDWDARRGACGCGGWHGGAKHV
---VHRGGTCABDBTGMCCTB-
| Original motif | Reverse complement motif |
| Consensus sequence: VDWDARRGACGCGGWHGGAKHV | Consensus sequence: VHYTCCDWCCGCGTCMMTHWHV |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 18 |
| Similarity score: | 0.0548087 |
Alignment:
BHCBCBCCGGGTGGTCYHVHDCH
-BAGGYCABHBTGACCKHV----
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00528 |
| Motif name: | Foxm1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 18 |
| Similarity score: | 0.055453 |
Alignment:
DCATDBTGHGCATKMTTBRSWR
----VHRGGTCABDBTGMCCTB
| Original motif | Reverse complement motif |
| Consensus sequence: MWSMBAARRATGCDCABHATGD | Consensus sequence: DCATDBTGHGCATKMTTBRSWR |
 |
 |
| Motif ID: | UP00032 |
| Motif name: | Gata3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 18 |
| Similarity score: | 0.0566426 |
Alignment:
HBHDVBWGATHBTATCRVBHHH
-VHRGGTCABDBTGMCCTB---
| Original motif | Reverse complement motif |
| Consensus sequence: HBHDVBWGATHBTATCRVBHHH | Consensus sequence: HHHBBMGATAVHATCWVVDHVH |
 |
 |
| Dataset #: 2 | Motif ID: 31 | Motif name: Esrrb |
| Original motif | Reverse complement motif |
| Consensus sequence: CACGTG | Consensus sequence: CACGTG |
 |
 |
Best Matches for Motif ID 31 (Highest to Lowest)
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
RDHDBVDTCACGTGASBHVHDH
--------CACGTG--------
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0026425 |
Alignment:
ACHRBHDTCCACGTGYAHHBMHM
---------CACGTG--------
| Original motif | Reverse complement motif |
| Consensus sequence: YDYBDHTMCACGTGGADDBMDGT | Consensus sequence: ACHRBHDTCCACGTGYAHHBMHM |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.0044422 |
Alignment:
DDASCACGTGBTBVDD
----CACGTG------
| Original motif | Reverse complement motif |
| Consensus sequence: DDASCACGTGBTBVDD | Consensus sequence: HHVBABCACGTGSTHD |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0.0342153 |
Alignment:
DHDBHGCACCTGBDDVB
------CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: VBHHVCAGGTGCDVDHD | Consensus sequence: DHDBHGCACCTGBDDVB |
 |
 |
| Motif ID: | UP00097 |
| Motif name: | Mtf1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0471928 |
Alignment:
HDYTTGCACACGDHBH
--------CACGTG--
| Original motif | Reverse complement motif |
| Consensus sequence: DBDHCGTGTGCAAMDH | Consensus sequence: HDYTTGCACACGDHBH |
 |
 |
| Motif ID: | UP00249 |
| Motif name: | Nkx2-5 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.0482749 |
Alignment:
HDADCCACTTRAAWTT
-----CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: HDADCCACTTRAAWTT | Consensus sequence: AAWTTMAAGTGGHTDH |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.0485437 |
Alignment:
BBGHCACGCGACDD
----CACGTG----
| Original motif | Reverse complement motif |
| Consensus sequence: BBGHCACGCGACDD | Consensus sequence: HDGTCGCGTGDCVB |
 |
 |
| Motif ID: | UP00231 |
| Motif name: | Nkx2-2 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0.0497234 |
Alignment:
DVDTKTCAAGTGGKBDH
------CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: HDVRCCACTTGARADBD | Consensus sequence: DVDTKTCAAGTGGKBDH |
 |
 |
| Motif ID: | UP00165 |
| Motif name: | Titf1 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.0498919 |
Alignment:
AAHYTCAAGTGGBTBV
-----CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: BVABCCACTTGAMHTT | Consensus sequence: AAHYTCAAGTGGBTBV |
 |
 |
| Motif ID: | UP00147 |
| Motif name: | Nkx2-6 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 6 |
| Similarity score: | 0.0544601 |
Alignment:
DDADCCACTTAAVHTT
-----CACGTG-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDADCCACTTAAVHTT | Consensus sequence: AAHVTTAAGTGGHTDD |
 |
 |
| Dataset #: 2 | Motif ID: 32 | Motif name: EWSR1-FLI1 |
| Original motif | Reverse complement motif |
| Consensus sequence: GGAAGGAAGGAAGGAAGG | Consensus sequence: CCTTCCTTCCTTCCTTCC |
 |
 |
Best Matches for Motif ID 32 (Highest to Lowest)
| Motif ID: | UP00419 |
| Motif name: | Spic |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0442834 |
Alignment:
----VAAAGMGGAAGTWD
GGAAGGAAGGAAGGAAGG--
| Original motif | Reverse complement motif |
| Consensus sequence: VAAAGMGGAAGTWD | Consensus sequence: DWACTTCCRCTTTB |
 |
 |
| Motif ID: | UP00055 |
| Motif name: | Hbp1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0466226 |
Alignment:
----VDHATTCATTSAWDDH
CCTTCCTTCCTTCCTTCC----
| Original motif | Reverse complement motif |
| Consensus sequence: HHDWTSAATGAATDDB | Consensus sequence: VDHATTCATTSAWDDH |
 |
 |
| Motif ID: | UP00035 |
| Motif name: | Hic1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.047846 |
Alignment:
----DDBHGGTTGGCATVBD
GGAAGGAAGGAAGGAAGG--
| Original motif | Reverse complement motif |
| Consensus sequence: DBBATGCCAACCHVHH | Consensus sequence: DDBHGGTTGGCATVBD |
 |
 |
| Motif ID: | UP00090 |
| Motif name: | Elf3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 13 |
| Similarity score: | 0.539121 |
Alignment:
-----TDAMTTCCYYDTD
CCTTCCTTCCTTCCTTCC---
| Original motif | Reverse complement motif |
| Consensus sequence: DAHMMGGAARTDA | Consensus sequence: TDAMTTCCYYDTD |
 |
 |
| Motif ID: | UP00062 |
| Motif name: | Sox4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 13 |
| Similarity score: | 0.547912 |
Alignment:
HHBBMCTTTGTTHDDBH-----
----CCTTCCTTCCTTCCTTCC
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDAACAAAGRVBHH | Consensus sequence: HHBBMCTTTGTTHDDBH |
 |
 |
| Motif ID: | UP00021 |
| Motif name: | Zfp281_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 13 |
| Similarity score: | 0.548284 |
Alignment:
-----GKRGGGGGGGGGGBD
GGAAGGAAGGAAGGAAGG--
| Original motif | Reverse complement motif |
| Consensus sequence: DBCCCCCCCCCCMYC | Consensus sequence: GKRGGGGGGGGGGBD |
 |
 |
| Motif ID: | UP00030 |
| Motif name: | Sox11_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 13 |
| Similarity score: | 0.54837 |
Alignment:
-----HHBTMCTTTGTTMDDVH
CCTTCCTTCCTTCCTTCC-----
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDRAACAAAGRABHH | Consensus sequence: HHBTMCTTTGTTMDDVH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 12 |
| Similarity score: | 1.04069 |
Alignment:
HVGVSAGGAGGGTCYHHHHYVH------
------GGAAGGAAGGAAGGAAGG----
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |
| Motif ID: | UP00409 |
| Motif name: | Elf5 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 12 |
| Similarity score: | 1.04159 |
Alignment:
------BWAHSMGGAAGTWD
GGAAGGAAGGAAGGAAGG--
| Original motif | Reverse complement motif |
| Consensus sequence: BWAHSMGGAAGTWD | Consensus sequence: DWACTTCCRSDTWV |
 |
 |
| Motif ID: | UP00056 |
| Motif name: | Rfx4_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 12 |
| Similarity score: | 1.04349 |
Alignment:
------HHBCHTRGWTACVVH
GGAAGGAAGGAAGGAAGG------
| Original motif | Reverse complement motif |
| Consensus sequence: HHBCHTRGWTACVVH | Consensus sequence: HVVGTAWCKADGBHH |
 |
 |
| Dataset #: 2 | Motif ID: 33 | Motif name: HIF1AARNT |
| Original motif | Reverse complement motif |
| Consensus sequence: VBACGTGV | Consensus sequence: VCACGTBV |
 |
 |
Best Matches for Motif ID 33 (Highest to Lowest)
| Motif ID: | UP00097 |
| Motif name: | Mtf1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
HDYTTGCACACGDHBH
-------VCACGTBV-
| Original motif | Reverse complement motif |
| Consensus sequence: DBDHCGTGTGCAAMDH | Consensus sequence: HDYTTGCACACGDHBH |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.000574345 |
Alignment:
HHVBABCACGTGSTHD
-----VCACGTBV---
| Original motif | Reverse complement motif |
| Consensus sequence: DDASCACGTGBTBVDD | Consensus sequence: HHVBABCACGTGSTHD |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0.00642455 |
Alignment:
YDYBDHTMCACGTGGADDBMDGT
-------VBACGTGV--------
| Original motif | Reverse complement motif |
| Consensus sequence: YDYBDHTMCACGTGGADDBMDGT | Consensus sequence: ACHRBHDTCCACGTGYAHHBMHM |
 |
 |
| Motif ID: | UP00084 |
| Motif name: | Gmeb1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0131171 |
Alignment:
BVVDVRYACGTRYVBHB
-----VBACGTGV----
| Original motif | Reverse complement motif |
| Consensus sequence: BHBBKKACGTMMVDBVB | Consensus sequence: BVVDVRYACGTRYVBHB |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.014006 |
Alignment:
RDHDBVDTCACGTGASBHVHDH
-------VCACGTBV-------
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0184296 |
Alignment:
VBHHVCAGGTGCDVDHD
----VBACGTGV-----
| Original motif | Reverse complement motif |
| Consensus sequence: VBHHVCAGGTGCDVDHD | Consensus sequence: DHDBHGCACCTGBDDVB |
 |
 |
| Motif ID: | UP00068 |
| Motif name: | Eomes_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 8 |
| Similarity score: | 0.0188154 |
Alignment:
VVBVCKACACCTHVBV
------VCACGTBV--
| Original motif | Reverse complement motif |
| Consensus sequence: VBVDAGGTGTYGVBBV | Consensus sequence: VVBVCKACACCTHVBV |
 |
 |
| Motif ID: | UP00026 |
| Motif name: | Zscan4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0.0189568 |
Alignment:
DHVDTGTGCACAYAHDH
-VBACGTGV--------
| Original motif | Reverse complement motif |
| Consensus sequence: DHVDTGTGCACAYAHDH | Consensus sequence: HDHTMTGTGCACADVHD |
 |
 |
| Motif ID: | UP00042 |
| Motif name: | Gm397_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.0198987 |
Alignment:
HHRTATGTGCACATHHD
--VBACGTGV-------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDATGTGCACATAMDH | Consensus sequence: HHRTATGTGCACATHHD |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.021967 |
Alignment:
BBGHCACGCGACDD
---VCACGTBV---
| Original motif | Reverse complement motif |
| Consensus sequence: BBGHCACGCGACDD | Consensus sequence: HDGTCGCGTGDCVB |
 |
 |
| Dataset #: 2 | Motif ID: 34 | Motif name: HNF4A |
| Original motif | Reverse complement motif |
| Consensus sequence: RGGBCAAAGKYCA | Consensus sequence: TGMYCTTTGBCCK |
 |
 |
Best Matches for Motif ID 34 (Highest to Lowest)
| Motif ID: | UP00058 |
| Motif name: | Tcf3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 13 |
| Similarity score: | 0.0357776 |
Alignment:
HHHASATCAAAGVRHHH
---RGGBCAAAGKYCA-
| Original motif | Reverse complement motif |
| Consensus sequence: HHHASATCAAAGVRHHH | Consensus sequence: HHHKVCTTTGATSTHHH |
 |
 |
| Motif ID: | UP00054 |
| Motif name: | Tcf7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 13 |
| Similarity score: | 0.0357792 |
Alignment:
BHHASATCAAAGGAHHH
---RGGBCAAAGKYCA-
| Original motif | Reverse complement motif |
| Consensus sequence: BHHASATCAAAGGAHHH | Consensus sequence: HHHTCCTTTGATSTHHV |
 |
 |
| Motif ID: | UP00067 |
| Motif name: | Lef1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 13 |
| Similarity score: | 0.0359011 |
Alignment:
VDDAGATCAAAGGKDDD
---RGGBCAAAGKYCA-
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATCTDDV | Consensus sequence: VDDAGATCAAAGGKDDD |
 |
 |
| Motif ID: | UP00083 |
| Motif name: | Tcf7l2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 13 |
| Similarity score: | 0.0360786 |
Alignment:
BDDASATCAAAGGMDDD
---RGGBCAAAGKYCA-
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATSTDDV | Consensus sequence: BDDASATCAAAGGMDDD |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 13 |
| Similarity score: | 0.0405764 |
Alignment:
HVDDVCAGATGKYHBBV
-RGGBCAAAGKYCA---
| Original motif | Reverse complement motif |
| Consensus sequence: HVDDVCAGATGKYHBBV | Consensus sequence: VBBDMYCATCTGVHHBH |
 |
 |
| Motif ID: | UP00062 |
| Motif name: | Sox4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 13 |
| Similarity score: | 0.0409915 |
Alignment:
HHBBMCTTTGTTHDDBH
-TGMYCTTTGBCCK---
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDAACAAAGRVBHH | Consensus sequence: HHBBMCTTTGTTHDDBH |
 |
 |
| Motif ID: | UP00101 |
| Motif name: | Sox12_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 13 |
| Similarity score: | 0.0413218 |
Alignment:
HDTMCTTTGTSTVDBB
TGMYCTTTGBCCK---
| Original motif | Reverse complement motif |
| Consensus sequence: VVDBASACAAAGRADH | Consensus sequence: HDTMCTTTGTSTVDBB |
 |
 |
| Motif ID: | UP00030 |
| Motif name: | Sox11_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 13 |
| Similarity score: | 0.0421105 |
Alignment:
HHBTMCTTTGTTMDDVH
-TGMYCTTTGBCCK---
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDRAACAAAGRABHH | Consensus sequence: HHBTMCTTTGTTMDDVH |
 |
 |
| Motif ID: | UP00086 |
| Motif name: | Irf3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 13 |
| Similarity score: | 0.0442004 |
Alignment:
DDAGAADGGDSCDV
RGGBCAAAGKYCA-
| Original motif | Reverse complement motif |
| Consensus sequence: DDAGAADGGDSCDV | Consensus sequence: BHGSDCCDTTCTHH |
 |
 |
| Motif ID: | UP00082 |
| Motif name: | Zfp187_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 13 |
| Similarity score: | 0.0446884 |
Alignment:
VHMDGGACVAGGRCDH
---RGGBCAAAGKYCA
| Original motif | Reverse complement motif |
| Consensus sequence: DDGMCCTBGTCCHYHV | Consensus sequence: VHMDGGACVAGGRCDH |
 |
 |
| Dataset #: 2 | Motif ID: 35 | Motif name: INSM1 |
| Original motif | Reverse complement motif |
| Consensus sequence: TGYCAGGGGGCR | Consensus sequence: MGCCCCCTGMCA |
 |
 |
Best Matches for Motif ID 35 (Highest to Lowest)
| Motif ID: | UP00022 |
| Motif name: | Zfp740_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 12 |
| Similarity score: | 0.00085101 |
Alignment:
HVKDCCGGGGGGWADDD
--TGYCAGGGGGCR---
| Original motif | Reverse complement motif |
| Consensus sequence: DDDTWCCCCCCGGDRVH | Consensus sequence: HVKDCCGGGGGGWADDD |
 |
 |
| Motif ID: | UP00002 |
| Motif name: | Sp4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 12 |
| Similarity score: | 0.00406431 |
Alignment:
BVVHAGGGGGCGRDHHB
TGYCAGGGGGCR-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDHHMCGCCCCCTHVBB | Consensus sequence: BVVHAGGGGGCGRDHHB |
 |
 |
| Motif ID: | UP00047 |
| Motif name: | Zbtb7b_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 12 |
| Similarity score: | 0.00486866 |
Alignment:
VHWTTHGGGGGGVDD
--TGYCAGGGGGCR-
| Original motif | Reverse complement motif |
| Consensus sequence: DDVCCCCCCHAAWHB | Consensus sequence: VHWTTHGGGGGGVDD |
 |
 |
| Motif ID: | UP00079 |
| Motif name: | Esrra_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 12 |
| Similarity score: | 0.00862553 |
Alignment:
VKBBAGGGGTCAHDBBH
TGYCAGGGGGCR-----
| Original motif | Reverse complement motif |
| Consensus sequence: VKBBAGGGGTCAHDBBH | Consensus sequence: DBBHHTGACCCCTBBYV |
 |
 |
| Motif ID: | UP00024 |
| Motif name: | Glis2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 12 |
| Similarity score: | 0.00936306 |
Alignment:
HKBBGGGGGGTCVHHH
-TGYCAGGGGGCR---
| Original motif | Reverse complement motif |
| Consensus sequence: HHHVGACCCCCCVBRD | Consensus sequence: HKBBGGGGGGTCVHHH |
 |
 |
| Motif ID: | UP00066 |
| Motif name: | Hnf4a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 12 |
| Similarity score: | 0.0123592 |
Alignment:
BDBVMGGGGTCAABHDV
TGYCAGGGGGCR-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDBVMGGGGTCAABHDV | Consensus sequence: BHHVTTGACCCCYVVDB |
 |
 |
| Motif ID: | UP00007 |
| Motif name: | Egr1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 12 |
| Similarity score: | 0.0155167 |
Alignment:
HCCGCCCCCGCAHB
--MGCCCCCTGMCA
| Original motif | Reverse complement motif |
| Consensus sequence: HCCGCCCCCGCAHB | Consensus sequence: VHTGCGGGGGCGGH |
 |
 |
| Motif ID: | UP00022 |
| Motif name: | Zfp740_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 12 |
| Similarity score: | 0.0176638 |
Alignment:
HVBCCCCCCCCMHHHB
---MGCCCCCTGMCA-
| Original motif | Reverse complement motif |
| Consensus sequence: HVBCCCCCCCCMHHHB | Consensus sequence: BHHDYGGGGGGGGBVD |
 |
 |
| Motif ID: | UP00043 |
| Motif name: | Bcl6b_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.0182432 |
Alignment:
BHHHWGGGGCGGBBVH
TGYCAGGGGGCR----
| Original motif | Reverse complement motif |
| Consensus sequence: HBBBCCGCCCCWHHHV | Consensus sequence: BHHHWGGGGCGGBBVH |
 |
 |
| Motif ID: | UP00070 |
| Motif name: | Gcm1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 12 |
| Similarity score: | 0.0200587 |
Alignment:
BYMBKCCCCTATDVBHD
--MGCCCCCTGMCA---
| Original motif | Reverse complement motif |
| Consensus sequence: DDBVHATAGGGGRBRMB | Consensus sequence: BYMBKCCCCTATDVBHD |
 |
 |
| Dataset #: 2 | Motif ID: 36 | Motif name: Klf4 |
| Original motif | Reverse complement motif |
| Consensus sequence: DGGGYGKGGC | Consensus sequence: GCCYCMCCCD |
 |
 |
Best Matches for Motif ID 36 (Highest to Lowest)
| Motif ID: | UP00093 |
| Motif name: | Klf7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0 |
Alignment:
DHHDGGGCGRGGKHBH
---DGGGYGKGGC---
| Original motif | Reverse complement motif |
| Consensus sequence: HBDRCCMCGCCCHHHD | Consensus sequence: DHHDGGGCGRGGKHBH |
 |
 |
| Motif ID: | UP00021 |
| Motif name: | Zfp281_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0153308 |
Alignment:
DBCCCCCCCCCCMYC
---GCCYCMCCCD--
| Original motif | Reverse complement motif |
| Consensus sequence: DBCCCCCCCCCCMYC | Consensus sequence: GKRGGGGGGGGGGBD |
 |
 |
| Motif ID: | UP00002 |
| Motif name: | Sp4_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0186791 |
Alignment:
BDWAGGCGTGBCHHD
--DGGGYGKGGC---
| Original motif | Reverse complement motif |
| Consensus sequence: BDWAGGCGTGBCHHD | Consensus sequence: HHDGBCACGCCTWDB |
 |
 |
| Motif ID: | UP00022 |
| Motif name: | Zfp740_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0218399 |
Alignment:
HVBCCCCCCCCMHHHB
--GCCYCMCCCD----
| Original motif | Reverse complement motif |
| Consensus sequence: HVBCCCCCCCCMHHHB | Consensus sequence: BHHDYGGGGGGGGBVD |
 |
 |
| Motif ID: | UP00007 |
| Motif name: | Egr1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.0249586 |
Alignment:
VHTGCGGGGGCGGH
GCCYCMCCCD----
| Original motif | Reverse complement motif |
| Consensus sequence: HCCGCCCCCGCAHB | Consensus sequence: VHTGCGGGGGCGGH |
 |
 |
| Motif ID: | UP00002 |
| Motif name: | Sp4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0299159 |
Alignment:
BVVHAGGGGGCGRDHHB
------DGGGYGKGGC-
| Original motif | Reverse complement motif |
| Consensus sequence: BDHHMCGCCCCCTHVBB | Consensus sequence: BVVHAGGGGGCGRDHHB |
 |
 |
| Motif ID: | UP00000 |
| Motif name: | Smad3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0308595 |
Alignment:
BHBHDTGGCGGGGBDHD
----DGGGYGKGGC---
| Original motif | Reverse complement motif |
| Consensus sequence: DHHBCCCCGCCAHHBHB | Consensus sequence: BHBHDTGGCGGGGBDHD |
 |
 |
| Motif ID: | UP00047 |
| Motif name: | Zbtb7b_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.0345891 |
Alignment:
DDVCCCCCCHAAWHB
GCCYCMCCCD-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDVCCCCCCHAAWHB | Consensus sequence: VHWTTHGGGGGGVDD |
 |
 |
| Motif ID: | UP00043 |
| Motif name: | Bcl6b_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0346735 |
Alignment:
HBBBCCGCCCCWHHHV
--GCCYCMCCCD----
| Original motif | Reverse complement motif |
| Consensus sequence: HBBBCCGCCCCWHHHV | Consensus sequence: BHHHWGGGGCGGBBVH |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.0351496 |
Alignment:
MHHHWHYTMCCHCCCACVAABVH
------GCCYCMCCCD-------
| Original motif | Reverse complement motif |
| Consensus sequence: MHHHWHYTMCCHCCCACVAABVH | Consensus sequence: DVVTTVGTGGGHGGYAMHWHHHY |
 |
 |
| Dataset #: 2 | Motif ID: 37 | Motif name: MIZF |
| Original motif | Reverse complement motif |
| Consensus sequence: BAACGTCCGC | Consensus sequence: GCGGACGTTV |
 |
 |
Best Matches for Motif ID 37 (Highest to Lowest)
| Motif ID: | UP00419 |
| Motif name: | Spic |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0 |
Alignment:
DWACTTCCRCTTTB
BAACGTCCGC----
| Original motif | Reverse complement motif |
| Consensus sequence: VAAAGMGGAAGTWD | Consensus sequence: DWACTTCCRCTTTB |
 |
 |
| Motif ID: | UP00409 |
| Motif name: | Elf5 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.00379999 |
Alignment:
DWACTTCCRSDTWV
BAACGTCCGC----
| Original motif | Reverse complement motif |
| Consensus sequence: BWAHSMGGAAGTWD | Consensus sequence: DWACTTCCRSDTWV |
 |
 |
| Motif ID: | UP00413 |
| Motif name: | Elf4 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0124101 |
Alignment:
DBBHACTTCCGGKWTH
--BAACGTCCGC----
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHACTTCCGGKWTH | Consensus sequence: HAWYCCGGAAGTHBBD |
 |
 |
| Motif ID: | UP00404 |
| Motif name: | Elf2 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0128348 |
Alignment:
DBBHACTTCCGGDWDB
--BAACGTCCGC----
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHACTTCCGGDWDB | Consensus sequence: BDWDCCGGAAGTHBBD |
 |
 |
| Motif ID: | UP00085 |
| Motif name: | Sfpi1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0141516 |
Alignment:
HDACTTCCBCWTDV
BAACGTCCGC----
| Original motif | Reverse complement motif |
| Consensus sequence: BDAWGVGGAAGTDH | Consensus sequence: HDACTTCCBCWTDV |
 |
 |
| Motif ID: | UP00423 |
| Motif name: | Gm5454 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0144665 |
Alignment:
DVHACCGGAAGTHDDV
----GCGGACGTTV--
| Original motif | Reverse complement motif |
| Consensus sequence: VDHHACTTCCGGTHVH | Consensus sequence: DVHACCGGAAGTHDDV |
 |
 |
| Motif ID: | UP00412 |
| Motif name: | Etv5 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0155825 |
Alignment:
DHHACCGGAWGTTDDV
----GCGGACGTTV--
| Original motif | Reverse complement motif |
| Consensus sequence: VDHAACWTCCGGTHHH | Consensus sequence: DHHACCGGAWGTTDDV |
 |
 |
| Motif ID: | UP00015 |
| Motif name: | Ehf_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0162018 |
Alignment:
VBDABCCGGAAGTDD
-----GCGGACGTTV
| Original motif | Reverse complement motif |
| Consensus sequence: VBDABCCGGAAGTDD | Consensus sequence: DDACTTCCGGBTHBB |
 |
 |
| Motif ID: | UP00418 |
| Motif name: | Etv6 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0171234 |
Alignment:
DVHBTACTTCCGGHTHB
---BAACGTCCGC----
| Original motif | Reverse complement motif |
| Consensus sequence: DVHBTACTTCCGGHTHB | Consensus sequence: BHAHCCGGAAGTABDVD |
 |
 |
| Motif ID: | UP00421 |
| Motif name: | Etv1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0184147 |
Alignment:
MVHBACCGGAAGTVDVH
-----GCGGACGTTV--
| Original motif | Reverse complement motif |
| Consensus sequence: MVHBACCGGAAGTVDVH | Consensus sequence: HBDBACTTCCGGTBHVY |
 |
 |
| Dataset #: 2 | Motif ID: 38 | Motif name: Myc |
| Original motif | Reverse complement motif |
| Consensus sequence: VGCACGTGGH | Consensus sequence: DCCACGTGCV |
 |
 |
Best Matches for Motif ID 38 (Highest to Lowest)
| Motif ID: | UP00060 |
| Motif name: | Max_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0 |
Alignment:
DDASCACGTGBTBVDD
--VGCACGTGGH----
| Original motif | Reverse complement motif |
| Consensus sequence: DDASCACGTGBTBVDD | Consensus sequence: HHVBABCACGTGSTHD |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.00631885 |
Alignment:
ACHRBHDTCCACGTGYAHHBMHM
-------DCCACGTGCV------
| Original motif | Reverse complement motif |
| Consensus sequence: YDYBDHTMCACGTGGADDBMDGT | Consensus sequence: ACHRBHDTCCACGTGYAHHBMHM |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.00821668 |
Alignment:
DDHBHBSTCACGTGAHBBDHHM
------DCCACGTGCV------
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0140005 |
Alignment:
DHDBHGCACCTGBDDVB
----VGCACGTGGH---
| Original motif | Reverse complement motif |
| Consensus sequence: VBHHVCAGGTGCDVDHD | Consensus sequence: DHDBHGCACCTGBDDVB |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0308912 |
Alignment:
BBGHCACGCGACDD
--VGCACGTGGH--
| Original motif | Reverse complement motif |
| Consensus sequence: BBGHCACGCGACDD | Consensus sequence: HDGTCGCGTGDCVB |
 |
 |
| Motif ID: | UP00165 |
| Motif name: | Titf1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0311723 |
Alignment:
BVABCCACTTGAMHTT
---DCCACGTGCV---
| Original motif | Reverse complement motif |
| Consensus sequence: BVABCCACTTGAMHTT | Consensus sequence: AAHYTCAAGTGGBTBV |
 |
 |
| Motif ID: | UP00249 |
| Motif name: | Nkx2-5 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.031308 |
Alignment:
HDADCCACTTRAAWTT
---DCCACGTGCV---
| Original motif | Reverse complement motif |
| Consensus sequence: HDADCCACTTRAAWTT | Consensus sequence: AAWTTMAAGTGGHTDH |
 |
 |
| Motif ID: | UP00231 |
| Motif name: | Nkx2-2 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0334845 |
Alignment:
HDVRCCACTTGARADBD
---DCCACGTGCV----
| Original motif | Reverse complement motif |
| Consensus sequence: HDVRCCACTTGARADBD | Consensus sequence: DVDTKTCAAGTGGKBDH |
 |
 |
| Motif ID: | UP00147 |
| Motif name: | Nkx2-6 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0345548 |
Alignment:
DDADCCACTTAAVHTT
---DCCACGTGCV---
| Original motif | Reverse complement motif |
| Consensus sequence: DDADCCACTTAAVHTT | Consensus sequence: AAHVTTAAGTGGHTDD |
 |
 |
| Motif ID: | UP00527 |
| Motif name: | Foxn4_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0363818 |
Alignment:
VDWDARRGACGCGGWHGGAKHV
-----VGCACGTGGH-------
| Original motif | Reverse complement motif |
| Consensus sequence: VDWDARRGACGCGGWHGGAKHV | Consensus sequence: VHYTCCDWCCGCGTCMMTHWHV |
 |
 |
| Dataset #: 2 | Motif ID: 39 | Motif name: MYCMAX |
| Original motif | Reverse complement motif |
| Consensus sequence: RASCACGTGGT | Consensus sequence: ACCACGTGSTM |
 |
 |
Best Matches for Motif ID 39 (Highest to Lowest)
| Motif ID: | UP00060 |
| Motif name: | Max_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0 |
Alignment:
HHVBABCACGTGSTHD
---RASCACGTGGT--
| Original motif | Reverse complement motif |
| Consensus sequence: DDASCACGTGBTBVDD | Consensus sequence: HHVBABCACGTGSTHD |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.025477 |
Alignment:
ACHRBHDTCCACGTGYAHHBMHM
-------ACCACGTGSTM-----
| Original motif | Reverse complement motif |
| Consensus sequence: YDYBDHTMCACGTGGADDBMDGT | Consensus sequence: ACHRBHDTCCACGTGYAHHBMHM |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.033078 |
Alignment:
RDHDBVDTCACGTGASBHVHDH
-----RASCACGTGGT------
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0375583 |
Alignment:
VBHHVCAGGTGCDVDHD
---ACCACGTGSTM---
| Original motif | Reverse complement motif |
| Consensus sequence: VBHHVCAGGTGCDVDHD | Consensus sequence: DHDBHGCACCTGBDDVB |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0452419 |
Alignment:
HVDDVCAGATGKYHBBV
--RASCACGTGGT----
| Original motif | Reverse complement motif |
| Consensus sequence: HVDDVCAGATGKYHBBV | Consensus sequence: VBBDMYCATCTGVHHBH |
 |
 |
| Motif ID: | UP00036 |
| Motif name: | Myf6_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0471968 |
Alignment:
VVDBAACAGBTGBYHB
---RASCACGTGGT--
| Original motif | Reverse complement motif |
| Consensus sequence: VVDBAACAGBTGBYHB | Consensus sequence: BDKVCABCTGTTBDBV |
 |
 |
| Motif ID: | UP00099 |
| Motif name: | Ascl2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0504418 |
Alignment:
HHDVVGCAGCTGVBKVB
---RASCACGTGGT---
| Original motif | Reverse complement motif |
| Consensus sequence: BBYVVCAGCTGCBVHHD | Consensus sequence: HHDVVGCAGCTGVBKVB |
 |
 |
| Motif ID: | UP00084 |
| Motif name: | Gmeb1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0522039 |
Alignment:
HHMACKACGTMGBVCD
---ACCACGTGSTM--
| Original motif | Reverse complement motif |
| Consensus sequence: DGVBCRACGTYGTYHH | Consensus sequence: HHMACKACGTMGBVCD |
 |
 |
| Motif ID: | UP00165 |
| Motif name: | Titf1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0554094 |
Alignment:
BVABCCACTTGAMHTT
---ACCACGTGSTM--
| Original motif | Reverse complement motif |
| Consensus sequence: BVABCCACTTGAMHTT | Consensus sequence: AAHYTCAAGTGGBTBV |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.0557594 |
Alignment:
HDGTCGCGTGDCVB
-RASCACGTGGT--
| Original motif | Reverse complement motif |
| Consensus sequence: BBGHCACGCGACDD | Consensus sequence: HDGTCGCGTGDCVB |
 |
 |
| Dataset #: 2 | Motif ID: 40 | Motif name: Mycn |
| Original motif | Reverse complement motif |
| Consensus sequence: HSCACGTGGC | Consensus sequence: GCCACGTGSD |
 |
 |
Best Matches for Motif ID 40 (Highest to Lowest)
| Motif ID: | UP00060 |
| Motif name: | Max_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0 |
Alignment:
DDASCACGTGBTBVDD
--HSCACGTGGC----
| Original motif | Reverse complement motif |
| Consensus sequence: DDASCACGTGBTBVDD | Consensus sequence: HHVBABCACGTGSTHD |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.00783793 |
Alignment:
ACHRBHDTCCACGTGYAHHBMHM
-------GCCACGTGSD------
| Original motif | Reverse complement motif |
| Consensus sequence: YDYBDHTMCACGTGGADDBMDGT | Consensus sequence: ACHRBHDTCCACGTGYAHHBMHM |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0132858 |
Alignment:
DDHBHBSTCACGTGAHBBDHHM
------GCCACGTGSD------
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0227933 |
Alignment:
VBHHVCAGGTGCDVDHD
---GCCACGTGSD----
| Original motif | Reverse complement motif |
| Consensus sequence: VBHHVCAGGTGCDVDHD | Consensus sequence: DHDBHGCACCTGBDDVB |
 |
 |
| Motif ID: | UP00060 |
| Motif name: | Max_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0257011 |
Alignment:
BBGHCACGCGACDD
--HSCACGTGGC--
| Original motif | Reverse complement motif |
| Consensus sequence: BBGHCACGCGACDD | Consensus sequence: HDGTCGCGTGDCVB |
 |
 |
| Motif ID: | UP00165 |
| Motif name: | Titf1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0344833 |
Alignment:
BVABCCACTTGAMHTT
---GCCACGTGSD---
| Original motif | Reverse complement motif |
| Consensus sequence: BVABCCACTTGAMHTT | Consensus sequence: AAHYTCAAGTGGBTBV |
 |
 |
| Motif ID: | UP00231 |
| Motif name: | Nkx2-2 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.035228 |
Alignment:
HDVRCCACTTGARADBD
---GCCACGTGSD----
| Original motif | Reverse complement motif |
| Consensus sequence: HDVRCCACTTGARADBD | Consensus sequence: DVDTKTCAAGTGGKBDH |
 |
 |
| Motif ID: | UP00084 |
| Motif name: | Gmeb1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0358581 |
Alignment:
BVVDVRYACGTRYVBHB
----GCCACGTGSD---
| Original motif | Reverse complement motif |
| Consensus sequence: BHBBKKACGTMMVDBVB | Consensus sequence: BVVDVRYACGTRYVBHB |
 |
 |
| Motif ID: | UP00084 |
| Motif name: | Gmeb1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.03725 |
Alignment:
HHMACKACGTMGBVCD
---HSCACGTGGC---
| Original motif | Reverse complement motif |
| Consensus sequence: DGVBCRACGTYGTYHH | Consensus sequence: HHMACKACGTMGBVCD |
 |
 |
| Motif ID: | UP00526 |
| Motif name: | Foxn1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0383187 |
Alignment:
WHVBVVMACGCGCGTCDYHWDH
------HSCACGTGGC------
| Original motif | Reverse complement motif |
| Consensus sequence: WHVBVVMACGCGCGTCDYHWDH | Consensus sequence: HDWHMHGACGCGCGTRBVVBHW |
 |
 |
| Dataset #: 2 | Motif ID: 41 | Motif name: MZF1_5-13 |
| Original motif | Reverse complement motif |
| Consensus sequence: BKAGGGGDAD | Consensus sequence: BTHCCCCTYB |
 |
 |
Best Matches for Motif ID 41 (Highest to Lowest)
| Motif ID: | UP00022 |
| Motif name: | Zfp740_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0090535 |
Alignment:
DDDTWCCCCCCGGDRVH
---BTHCCCCTYB----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDTWCCCCCCGGDRVH | Consensus sequence: HVKDCCGGGGGGWADDD |
 |
 |
| Motif ID: | UP00007 |
| Motif name: | Egr1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.00910178 |
Alignment:
BBHBGAGTGGGAHHDB
---BKAGGGGDAD---
| Original motif | Reverse complement motif |
| Consensus sequence: BBHBGAGTGGGAHHDB | Consensus sequence: BHHDTCCCACTCBDBV |
 |
 |
| Motif ID: | UP00070 |
| Motif name: | Gcm1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0101177 |
Alignment:
DDBVHATAGGGGRBRMB
-----BKAGGGGDAD--
| Original motif | Reverse complement motif |
| Consensus sequence: DDBVHATAGGGGRBRMB | Consensus sequence: BYMBKCCCCTATDVBHD |
 |
 |
| Motif ID: | UP00021 |
| Motif name: | Zfp281_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0103225 |
Alignment:
GKRGGGGGGGGGGBD
BKAGGGGDAD-----
| Original motif | Reverse complement motif |
| Consensus sequence: DBCCCCCCCCCCMYC | Consensus sequence: GKRGGGGGGGGGGBD |
 |
 |
| Motif ID: | UP00099 |
| Motif name: | Ascl2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0126235 |
Alignment:
HVDVGGGHGGGGDBHB
-----BKAGGGGDAD-
| Original motif | Reverse complement motif |
| Consensus sequence: BHVDCCCCDCCCBDBH | Consensus sequence: HVDVGGGHGGGGDBHB |
 |
 |
| Motif ID: | UP00013 |
| Motif name: | Gabpa_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.013531 |
Alignment:
DBVVDRSGGAWKVHKG
-BKAGGGGDAD-----
| Original motif | Reverse complement motif |
| Consensus sequence: CYDBYWTCCSMHBVVH | Consensus sequence: DBVVDRSGGAWKVHKG |
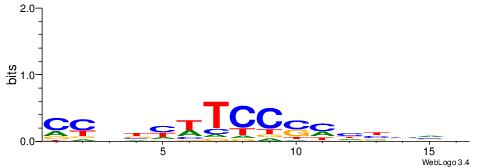 |
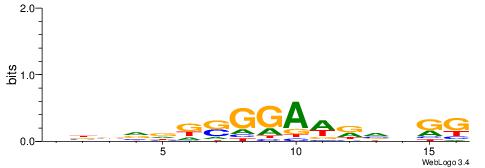 |
| Motif ID: | UP00022 |
| Motif name: | Zfp740_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0152996 |
Alignment:
BHHDYGGGGGGGGBVD
---BKAGGGGDAD---
| Original motif | Reverse complement motif |
| Consensus sequence: HVBCCCCCCCCMHHHB | Consensus sequence: BHHDYGGGGGGGGBVD |
 |
 |
| Motif ID: | UP00055 |
| Motif name: | Hbp1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.0176527 |
Alignment:
DDDYCCCATTGTVHABH
-BTHCCCCTYB------
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCCATTGTVHABH | Consensus sequence: HBTHVACAATGGGMDHD |
 |
 |
| Motif ID: | UP00002 |
| Motif name: | Sp4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0183345 |
Alignment:
BDHHMCGCCCCCTHVBB
-----BTHCCCCTYB--
| Original motif | Reverse complement motif |
| Consensus sequence: BDHHMCGCCCCCTHVBB | Consensus sequence: BVVHAGGGGGCGRDHHB |
 |
 |
| Motif ID: | UP00239 |
| Motif name: | Obox2 |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.019782 |
Alignment:
DBRGGGGGATTAAMTAH
BKAGGGGDAD-------
| Original motif | Reverse complement motif |
| Consensus sequence: DBRGGGGGATTAAMTAH | Consensus sequence: HTARTTAATCCCCCKBD |
 |
 |
| Dataset #: 2 | Motif ID: 42 | Motif name: NFKB1 |
| Original motif | Reverse complement motif |
| Consensus sequence: GGGGRTTCCCC | Consensus sequence: GGGGAAKCCCC |
 |
 |
Best Matches for Motif ID 42 (Highest to Lowest)
| Motif ID: | UP00021 |
| Motif name: | Zfp281_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0.0371794 |
Alignment:
VDDAHTGGGGGTHWYYD
------GGGGRTTCCCC
| Original motif | Reverse complement motif |
| Consensus sequence: DKKWDACCCCCAHTDDV | Consensus sequence: VDDAHTGGGGGTHWYYD |
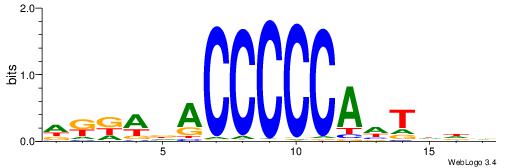 |
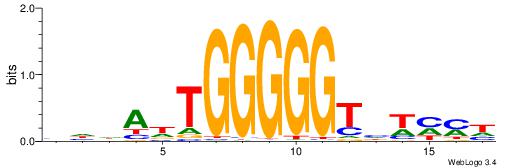 |
| Motif ID: | UP00088 |
| Motif name: | Plagl1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0407901 |
Alignment:
DBHGGGGGGTACCHHHD
----GGGGRTTCCCC--
| Original motif | Reverse complement motif |
| Consensus sequence: DBHGGGGGGTACCHHHD | Consensus sequence: DHDDGGTACCCCCCHBH |
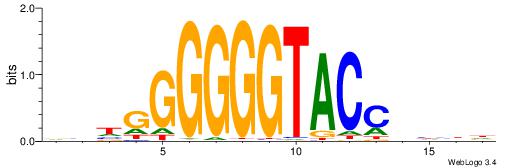 |
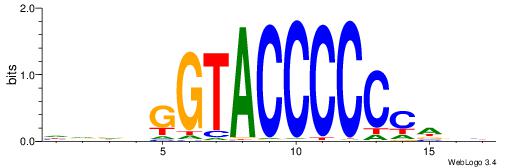 |
| Motif ID: | UP00088 |
| Motif name: | Plagl1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0459696 |
Alignment:
BHDGGGGSSCCCCBVV
--GGGGRTTCCCC---
| Original motif | Reverse complement motif |
| Consensus sequence: BHDGGGGSSCCCCBVV | Consensus sequence: VBVGGGGSSCCCCHHV |
 |
 |
| Motif ID: | UP00176 |
| Motif name: | Crx |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.0464973 |
Alignment:
DGGMTAATCCCCDWBK
-GGGGAAKCCCC----
| Original motif | Reverse complement motif |
| Consensus sequence: YBWDGGGGATTARCCD | Consensus sequence: DGGMTAATCCCCDWBK |
 |
 |
| Motif ID: | UP00265 |
| Motif name: | Pitx3 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.048533 |
Alignment:
GBHDGCTAATCCCCCB
---GGGGAAKCCCC--
| Original motif | Reverse complement motif |
| Consensus sequence: VGGGGGATTAGCDDBC | Consensus sequence: GBHDGCTAATCCCCCB |
 |
 |
| Motif ID: | UP00216 |
| Motif name: | Obox1 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0502644 |
Alignment:
DHDGTTAATCCCCKTDD
--GGGGAAKCCCC----
| Original motif | Reverse complement motif |
| Consensus sequence: DDARGGGGATTAACDHH | Consensus sequence: DHDGTTAATCCCCKTDD |
 |
 |
| Motif ID: | UP00239 |
| Motif name: | Obox2 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0515498 |
Alignment:
HTARTTAATCCCCCKBD
--GGGGAAKCCCC----
| Original motif | Reverse complement motif |
| Consensus sequence: DBRGGGGGATTAAMTAH | Consensus sequence: HTARTTAATCCCCCKBD |
 |
 |
| Motif ID: | UP00160 |
| Motif name: | Obox3 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0520605 |
Alignment:
HHWGTTAATCCCCCBBH
--GGGGAAKCCCC----
| Original motif | Reverse complement motif |
| Consensus sequence: HBVGGGGGATTAACWHH | Consensus sequence: HHWGTTAATCCCCCBBH |
 |
 |
| Motif ID: | UP00024 |
| Motif name: | Glis2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0.052761 |
Alignment:
HHHVGACCCCCCVBRD
-----GGGGRTTCCCC
| Original motif | Reverse complement motif |
| Consensus sequence: HHHVGACCCCCCVBRD | Consensus sequence: HKBBGGGGGGTCVHHH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0555704 |
Alignment:
CBDMCMGGGTGGTCCHVBVBAH
------GGGGAAKCCCC-----
| Original motif | Reverse complement motif |
| Consensus sequence: HTBVVVDGGACCACCCRGRDBG | Consensus sequence: CBDMCMGGGTGGTCCHVBVBAH |
 |
 |
| Dataset #: 2 | Motif ID: 43 | Motif name: NR1H2RXRA |
| Original motif | Reverse complement motif |
| Consensus sequence: AAAGGTCAAAGGTCAAC | Consensus sequence: GTTGACCTTTGACCTTT |
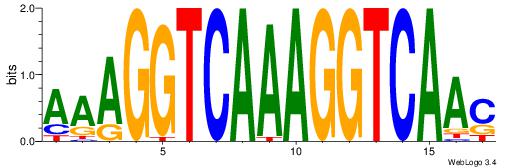 |
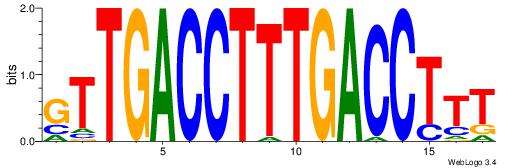 |
Best Matches for Motif ID 43 (Highest to Lowest)
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 17 |
| Similarity score: | 0.0538452 |
Alignment:
DVMHHDHKGACCCTCCTSVCBH
-----GTTGACCTTTGACCTTT
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |
| Motif ID: | UP00032 |
| Motif name: | Gata3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 17 |
| Similarity score: | 0.0545098 |
Alignment:
BDHDDADAGATAAGADHBDHVD
-----AAAGGTCAAAGGTCAAC
| Original motif | Reverse complement motif |
| Consensus sequence: BDHDDADAGATAAGADHBDHVD | Consensus sequence: HBHDVHDTCTTATCTHTDDHDV |
 |
 |
| Motif ID: | UP00067 |
| Motif name: | Lef1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 16 |
| Similarity score: | 0.521114 |
Alignment:
VDDAGATCAAAGGKDDD-
-AAAGGTCAAAGGTCAAC
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATCTDDV | Consensus sequence: VDDAGATCAAAGGKDDD |
 |
 |
| Motif ID: | UP00083 |
| Motif name: | Tcf7l2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 16 |
| Similarity score: | 0.527825 |
Alignment:
BDDASATCAAAGGMDDD-
-GTTGACCTTTGACCTTT
| Original motif | Reverse complement motif |
| Consensus sequence: DDDYCCTTTGATSTDDV | Consensus sequence: BDDASATCAAAGGMDDD |
 |
 |
| Motif ID: | UP00054 |
| Motif name: | Tcf7_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 16 |
| Similarity score: | 0.531154 |
Alignment:
-BHHASATCAAAGGAHHH
AAAGGTCAAAGGTCAAC-
| Original motif | Reverse complement motif |
| Consensus sequence: BHHASATCAAAGGAHHH | Consensus sequence: HHHTCCTTTGATSTHHV |
 |
 |
| Motif ID: | UP00058 |
| Motif name: | Tcf3_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 16 |
| Similarity score: | 0.531876 |
Alignment:
-HHHASATCAAAGVRHHH
AAAGGTCAAAGGTCAAC-
| Original motif | Reverse complement motif |
| Consensus sequence: HHHASATCAAAGVRHHH | Consensus sequence: HHHKVCTTTGATSTHHH |
 |
 |
| Motif ID: | UP00030 |
| Motif name: | Sox11_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 16 |
| Similarity score: | 0.544403 |
Alignment:
-HBDDRAACAAAGRABHH
AAAGGTCAAAGGTCAAC-
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDRAACAAAGRABHH | Consensus sequence: HHBTMCTTTGTTMDDVH |
 |
 |
| Motif ID: | UP00062 |
| Motif name: | Sox4_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 16 |
| Similarity score: | 0.544565 |
Alignment:
-HHBBMCTTTGTTHDDBH
GTTGACCTTTGACCTTT-
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDAACAAAGRVBHH | Consensus sequence: HHBBMCTTTGTTHDDBH |
 |
 |
| Motif ID: | UP00019 |
| Motif name: | Zbtb12_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 16 |
| Similarity score: | 0.550511 |
Alignment:
-BBGATCTAGAACBKHDB
GTTGACCTTTGACCTTT-
| Original motif | Reverse complement motif |
| Consensus sequence: BDHRBGTTCTAGATCVB | Consensus sequence: BBGATCTAGAACBKHDB |
 |
 |
| Motif ID: | UP00046 |
| Motif name: | Tcfe2a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 16 |
| Similarity score: | 0.552064 |
Alignment:
HVDDVCAGATGKYHBBV-
-AAAGGTCAAAGGTCAAC
| Original motif | Reverse complement motif |
| Consensus sequence: HVDDVCAGATGKYHBBV | Consensus sequence: VBBDMYCATCTGVHHBH |
 |
 |
| Dataset #: 2 | Motif ID: 44 | Motif name: NR4A2 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAGGTCAC | Consensus sequence: GTGACCTT |
 |
 |
Best Matches for Motif ID 44 (Highest to Lowest)
| Motif ID: | UP00048 |
| Motif name: | Rara |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
DHBHAAAGGTCACVHD
-----AAGGTCAC---
| Original motif | Reverse complement motif |
| Consensus sequence: DHBHAAAGGTCACVHD | Consensus sequence: HHVGTGACCTTTDVDD |
 |
 |
| Motif ID: | UP00009 |
| Motif name: | Nr2f2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0074527 |
Alignment:
DHBHAAAGGTCAHVHB
-----AAGGTCAC---
| Original motif | Reverse complement motif |
| Consensus sequence: DHBHAAAGGTCAHVHB | Consensus sequence: BHVDTGACCTTTDVDD |
 |
 |
| Motif ID: | UP00079 |
| Motif name: | Esrra_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.01542 |
Alignment:
DHDBCAAGGTCAHVBDH
-----AAGGTCAC----
| Original motif | Reverse complement motif |
| Consensus sequence: DHDBCAAGGTCAHVBDH | Consensus sequence: HHBVHTGACCTTGVDHD |
 |
 |
| Motif ID: | UP00009 |
| Motif name: | Nr2f2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0395644 |
Alignment:
VBVBCSGGGTCAVBBH
-----AAGGTCAC---
| Original motif | Reverse complement motif |
| Consensus sequence: VBVBCSGGGTCAVBBH | Consensus sequence: HVBVTGACCCSGBVBV |
 |
 |
| Motif ID: | UP00082 |
| Motif name: | Zfp187_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0407874 |
Alignment:
DDGMCCTBGTCCHYHV
GTGACCTT--------
| Original motif | Reverse complement motif |
| Consensus sequence: DDGMCCTBGTCCHYHV | Consensus sequence: VHMDGGACVAGGRCDH |
 |
 |
| Motif ID: | UP00053 |
| Motif name: | Rxra_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0422933 |
Alignment:
DBHBDTGACCCCDHVHB
----GTGACCTT-----
| Original motif | Reverse complement motif |
| Consensus sequence: DBHBDTGACCCCDHVHB | Consensus sequence: VHBHDGGGGTCAHBHBD |
 |
 |
| Motif ID: | UP00079 |
| Motif name: | Esrra_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0447444 |
Alignment:
VKBBAGGGGTCAHDBBH
-----AAGGTCAC----
| Original motif | Reverse complement motif |
| Consensus sequence: VKBBAGGGGTCAHDBBH | Consensus sequence: DBBHHTGACCCCTBBYV |
 |
 |
| Motif ID: | UP00066 |
| Motif name: | Hnf4a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.0481418 |
Alignment:
BBHHAAAGTCCAMTHH
-----AAGGTCAC---
| Original motif | Reverse complement motif |
| Consensus sequence: BBHHAAAGTCCAMTHH | Consensus sequence: HHAYTGGACTTTHDBV |
 |
 |
| Motif ID: | UP00197 |
| Motif name: | Hoxc9 |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.0497679 |
Alignment:
VDHHTTAATGACCBHV
-------GTGACCTT-
| Original motif | Reverse complement motif |
| Consensus sequence: VDVGGTCATTAAHHDB | Consensus sequence: VDHHTTAATGACCBHV |
 |
 |
| Motif ID: | UP00035 |
| Motif name: | Hic1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0500062 |
Alignment:
DBDDRTGCCCAWDHDV
----GTGACCTT----
| Original motif | Reverse complement motif |
| Consensus sequence: DBDDRTGCCCAWDHDV | Consensus sequence: VHHDWTGGGCAMDHBH |
 |
 |
| Dataset #: 2 | Motif ID: 45 | Motif name: Pax5 |
| Original motif | Reverse complement motif |
| Consensus sequence: DGVBCABTGDWGCGKRRCSR | Consensus sequence: MSGKKRCGCWDCABTGBBCD |
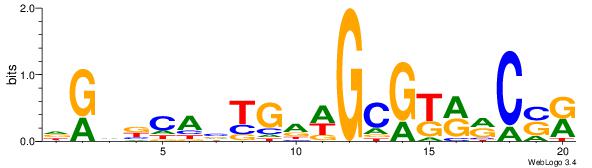 |
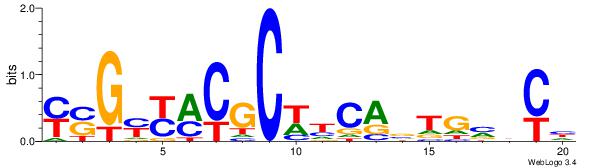 |
Best Matches for Motif ID 45 (Highest to Lowest)
| Motif ID: | UP00526 |
| Motif name: | Foxn1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0499537 |
Alignment:
DDRMHBACGCGYGCGTDGBBDB
-DGVBCABTGDWGCGKRRCSR-
| Original motif | Reverse complement motif |
| Consensus sequence: DDRMHBACGCGYGCGTDGBBDB | Consensus sequence: BDVBCDACGCKCGCGTBHRKHD |
 |
 |
| Motif ID: | UP00526 |
| Motif name: | Foxn1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0520196 |
Alignment:
HDWHMHGACGCGCGTRBVVBHW
DGVBCABTGDWGCGKRRCSR--
| Original motif | Reverse complement motif |
| Consensus sequence: WHVBVVMACGCGCGTCDYHWDH | Consensus sequence: HDWHMHGACGCGCGTRBVVBHW |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0542764 |
Alignment:
HDBABCGBKRGYGGCGMSBHAK
-DGVBCABTGDWGCGKRRCSR-
| Original motif | Reverse complement motif |
| Consensus sequence: RTHBSYCGCCMCMYVCGBTVDH | Consensus sequence: HDBABCGBKRGYGGCGMSBHAK |
 |
 |
| Motif ID: | UP00528 |
| Motif name: | Foxm1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0560636 |
Alignment:
DCATDBTGHGCATKMTTBRSWR
--MSGKKRCGCWDCABTGBBCD
| Original motif | Reverse complement motif |
| Consensus sequence: MWSMBAARRATGCDCABHATGD | Consensus sequence: DCATDBTGHGCATKMTTBRSWR |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0595671 |
Alignment:
VYWVVHCRCCACMCACGAHSBHH
DGVBCABTGDWGCGKRRCSR---
| Original motif | Reverse complement motif |
| Consensus sequence: VYWVVHCRCCACMCACGAHSBHH | Consensus sequence: HHBSHTCGTGRGTGGKGDBVWMV |
 |
 |
| Motif ID: | UP00528 |
| Motif name: | Foxm1_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0602559 |
Alignment:
HADTBVADGATGCATCMTGMVHB
DGVBCABTGDWGCGKRRCSR---
| Original motif | Reverse complement motif |
| Consensus sequence: HADTBVADGATGCATCMTGMVHB | Consensus sequence: VHVRCAYGATGCATCDTVVADTH |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 20 |
| Similarity score: | 0.0607506 |
Alignment:
HVBDAGCGBGGGTGGCRDBDDDB
DGVBCABTGDWGCGKRRCSR---
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Motif ID: | UP00098 |
| Motif name: | Rfx3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 20 |
| Similarity score: | 0.0611227 |
Alignment:
BDBBVBVGTAWCCAAGVBHBAGB
MSGKKRCGCWDCABTGBBCD---
| Original motif | Reverse complement motif |
| Consensus sequence: VCTBHBVCTTGGWTACVVVVVDB | Consensus sequence: BDBBVBVGTAWCCAAGVBHBAGB |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0611361 |
Alignment:
RDHDBVDTCACGTGASBHVHDH
--MSGKKRCGCWDCABTGBBCD
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.061403 |
Alignment:
GTVBDAATTGTVTGHGDDHKVB
DGVBCABTGDWGCGKRRCSR--
| Original motif | Reverse complement motif |
| Consensus sequence: GTVBDAATTGTVTGHGDDHKVB | Consensus sequence: VVYDDDCHCAVACAATTDBVAC |
 |
 |
| Dataset #: 2 | Motif ID: 46 | Motif name: RORA_1 |
| Original motif | Reverse complement motif |
| Consensus sequence: AWVDAGGTCA | Consensus sequence: TGACCTDVWT |
 |
 |
Best Matches for Motif ID 46 (Highest to Lowest)
| Motif ID: | UP00009 |
| Motif name: | Nr2f2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0154749 |
Alignment:
DHBHAAAGGTCAHVHB
--AWVDAGGTCA----
| Original motif | Reverse complement motif |
| Consensus sequence: DHBHAAAGGTCAHVHB | Consensus sequence: BHVDTGACCTTTDVDD |
 |
 |
| Motif ID: | UP00079 |
| Motif name: | Esrra_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0176661 |
Alignment:
HHBVHTGACCTTGVDHD
-----TGACCTDVWT--
| Original motif | Reverse complement motif |
| Consensus sequence: DHDBCAAGGTCAHVBDH | Consensus sequence: HHBVHTGACCTTGVDHD |
 |
 |
| Motif ID: | UP00048 |
| Motif name: | Rara |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0189599 |
Alignment:
DHBHAAAGGTCACVHD
--AWVDAGGTCA----
| Original motif | Reverse complement motif |
| Consensus sequence: DHBHAAAGGTCACVHD | Consensus sequence: HHVGTGACCTTTDVDD |
 |
 |
| Motif ID: | UP00009 |
| Motif name: | Nr2f2_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0331999 |
Alignment:
VBVBCSGGGTCAVBBH
--AWVDAGGTCA----
| Original motif | Reverse complement motif |
| Consensus sequence: VBVBCSGGGTCAVBBH | Consensus sequence: HVBVTGACCCSGBVBV |
 |
 |
| Motif ID: | UP00066 |
| Motif name: | Hnf4a_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0341771 |
Alignment:
BDBVMGGGGTCAABHDV
--AWVDAGGTCA-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDBVMGGGGTCAABHDV | Consensus sequence: BHHVTTGACCCCYVVDB |
 |
 |
| Motif ID: | UP00053 |
| Motif name: | Rxra_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0363566 |
Alignment:
DBHBDTGACCCCDHVHB
-----TGACCTDVWT--
| Original motif | Reverse complement motif |
| Consensus sequence: DBHBDTGACCCCDHVHB | Consensus sequence: VHBHDGGGGTCAHBHBD |
 |
 |
| Motif ID: | UP00079 |
| Motif name: | Esrra_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0391284 |
Alignment:
VKBBAGGGGTCAHDBBH
--AWVDAGGTCA-----
| Original motif | Reverse complement motif |
| Consensus sequence: VKBBAGGGGTCAHDBBH | Consensus sequence: DBBHHTGACCCCTBBYV |
 |
 |
| Motif ID: | UP00035 |
| Motif name: | Hic1_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.0444205 |
Alignment:
DBDDRTGCCCAWDHDV
-----TGACCTDVWT-
| Original motif | Reverse complement motif |
| Consensus sequence: DBDDRTGCCCAWDHDV | Consensus sequence: VHHDWTGGGCAMDHBH |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.0461434 |
Alignment:
DDHBHBSTCACGTGAHBBDHHM
AWVDAGGTCA------------
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0489571 |
Alignment:
BHCBCBCCGGGTGGTCYHVHDCH
-----------TGACCTDVWT--
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Dataset #: 2 | Motif ID: 47 | Motif name: RXRRAR_DR5 |
| Original motif | Reverse complement motif |
| Consensus sequence: RGKTCABVVRGAGGTCA | Consensus sequence: TGACCTCKVVBTGAYCK |
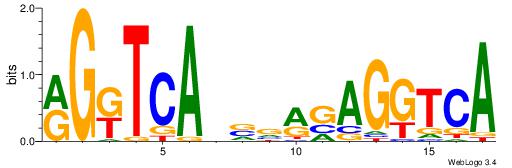 |
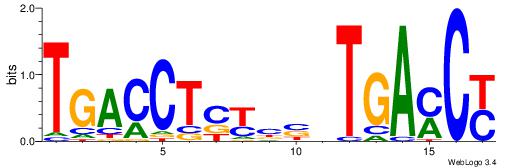 |
Best Matches for Motif ID 47 (Highest to Lowest)
| Motif ID: | UP00540 |
| Motif name: | Gli3_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 17 |
| Similarity score: | 0.0585087 |
Alignment:
DVTTVVCVYGGHGGYCHHDMYB
RGKTCABVVRGAGGTCA-----
| Original motif | Reverse complement motif |
| Consensus sequence: DVTTVVCVYGGHGGYCHHDMYB | Consensus sequence: VMYDHDGMCCHCCKBGVVAAVH |
 |
 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 17 |
| Similarity score: | 0.0709955 |
Alignment:
GTVBDAATTGTVTGHGDDHKVB
-----RGKTCABVVRGAGGTCA
| Original motif | Reverse complement motif |
| Consensus sequence: GTVBDAATTGTVTGHGDDHKVB | Consensus sequence: VVYDDDCHCAVACAATTDBVAC |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 17 |
| Similarity score: | 0.075055 |
Alignment:
DVVTTVGTGGGHGGYAMHWHHHY
RGKTCABVVRGAGGTCA------
| Original motif | Reverse complement motif |
| Consensus sequence: MHHHWHYTMCCHCCCACVAABVH | Consensus sequence: DVVTTVGTGGGHGGYAMHWHHHY |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 17 |
| Similarity score: | 0.0758824 |
Alignment:
BBBDVVRGACCACCCAVGABBAB
------RGKTCABVVRGAGGTCA
| Original motif | Reverse complement motif |
| Consensus sequence: BBBDVVRGACCACCCAVGABBAB | Consensus sequence: BTBVTCVTGGGTGGTCMVVDVBB |
 |
 |
| Motif ID: | UP00032 |
| Motif name: | Gata3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 17 |
| Similarity score: | 0.0759953 |
Alignment:
HBHDVBWGATHBTATCRVBHHH
TGACCTCKVVBTGAYCK-----
| Original motif | Reverse complement motif |
| Consensus sequence: HBHDVBWGATHBTATCRVBHHH | Consensus sequence: HHHBBMGATAVHATCWVVDHVH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 17 |
| Similarity score: | 0.0760211 |
Alignment:
DTVBDDRGACCACCCAGDWDGBH
------RGKTCABVVRGAGGTCA
| Original motif | Reverse complement motif |
| Consensus sequence: DTVBDDRGACCACCCAGDWDGBH | Consensus sequence: DBCDWHCTGGGTGGTCMDHBBAH |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 17 |
| Similarity score: | 0.0763398 |
Alignment:
BHCBCBCCGGGTGGTCYHVHDCH
TGACCTCKVVBTGAYCK------
| Original motif | Reverse complement motif |
| Consensus sequence: BHCBCBCCGGGTGGTCYHVHDCH | Consensus sequence: HGDHVHKGACCACCCGGBGBGDB |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 17 |
| Similarity score: | 0.0768658 |
Alignment:
HDBABCGBKRGYGGCGMSBHAK
RGKTCABVVRGAGGTCA-----
| Original motif | Reverse complement motif |
| Consensus sequence: RTHBSYCGCCMCMYVCGBTVDH | Consensus sequence: HDBABCGBKRGYGGCGMSBHAK |
 |
 |
| Motif ID: | UP00050 |
| Motif name: | Bhlhb2_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 17 |
| Similarity score: | 0.0784072 |
Alignment:
DDHBHBSTCACGTGAHBBDHHM
----RGKTCABVVRGAGGTCA-
| Original motif | Reverse complement motif |
| Consensus sequence: RDHDBVDTCACGTGASBHVHDH | Consensus sequence: DDHBHBSTCACGTGAHBBDHHM |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 17 |
| Similarity score: | 0.0794665 |
Alignment:
HVBDAGCGBGGGTGGCRDBDDDB
RGKTCABVVRGAGGTCA------
| Original motif | Reverse complement motif |
| Consensus sequence: HVBDAGCGBGGGTGGCRDBDDDB | Consensus sequence: BDDHVDKGCCACCCVCGCTDBVH |
 |
 |
| Dataset #: 2 | Motif ID: 48 | Motif name: Tcfcp2l1 |
| Original motif | Reverse complement motif |
| Consensus sequence: CCAGYYHVADCCRG | Consensus sequence: CKGGDTBDMMCTGG |
 |
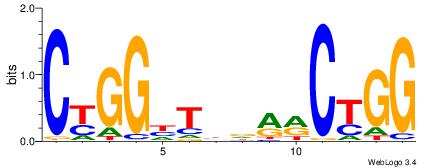 |
Best Matches for Motif ID 48 (Highest to Lowest)
| Motif ID: | UP00032 |
| Motif name: | Gata3_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0527249 |
Alignment:
HBHDVBWGATHBTATCRVBHHH
----CCAGYYHVADCCRG----
| Original motif | Reverse complement motif |
| Consensus sequence: HBHDVBWGATHBTATCRVBHHH | Consensus sequence: HHHBBMGATAVHATCWVVDHVH |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0551691 |
Alignment:
RTHBSYCGCCMCMYVCGBTVDH
--------CCAGYYHVADCCRG
| Original motif | Reverse complement motif |
| Consensus sequence: RTHBSYCGCCMCMYVCGBTVDH | Consensus sequence: HDBABCGBKRGYGGCGMSBHAK |
 |
 |
| Motif ID: | UP00005 |
| Motif name: | Tcfap2a_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0560983 |
Alignment:
HHRCCBBWGGDCDB
CCAGYYHVADCCRG
| Original motif | Reverse complement motif |
| Consensus sequence: HHRCCBBWGGDCDB | Consensus sequence: BDGHCCWBVGGKDH |
 |
 |
| Motif ID: | UP00013 |
| Motif name: | Gabpa_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0566467 |
Alignment:
CYDBYWTCCSMHBVVH
CCAGYYHVADCCRG--
| Original motif | Reverse complement motif |
| Consensus sequence: CYDBYWTCCSMHBVVH | Consensus sequence: DBVVDRSGGAWKVHKG |
 |
 |
| Motif ID: | UP00075 |
| Motif name: | Sox15_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0577636 |
Alignment:
BBVAATGDMATTHVA
-CCAGYYHVADCCRG
| Original motif | Reverse complement motif |
| Consensus sequence: BBVAATGDMATTHVA | Consensus sequence: TVDAATYDCATTVVV |
 |
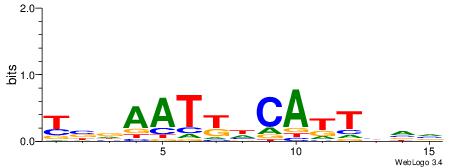 |
| Motif ID: | UP00096 |
| Motif name: | Sox13_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0587893 |
Alignment:
DDDHTGGGTGGGHAHHD
---CKGGDTBDMMCTGG
| Original motif | Reverse complement motif |
| Consensus sequence: DDDHTGGGTGGGHAHHD | Consensus sequence: DHHTHCCCACCCAHDDH |
 |
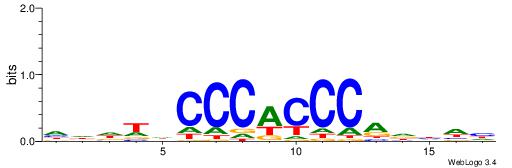 |
| Motif ID: | UP00034 |
| Motif name: | Sox7_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0590238 |
Alignment:
VVYDDDCHCAVACAATTDBVAC
--CCAGYYHVADCCRG------
| Original motif | Reverse complement motif |
| Consensus sequence: GTVBDAATTGTVTGHGDDHKVB | Consensus sequence: VVYDDDCHCAVACAATTDBVAC |
 |
 |
| Motif ID: | UP00540 |
| Motif name: | Gli3_v016060_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 14 |
| Similarity score: | 0.0596904 |
Alignment:
VYWVVHCRCCACMCACGAHSBHH
--------CCAGYYHVADCCRG-
| Original motif | Reverse complement motif |
| Consensus sequence: VYWVVHCRCCACMCACGAHSBHH | Consensus sequence: HHBSHTCGTGRGTGGKGDBVWMV |
 |
 |
| Motif ID: | UP00538 |
| Motif name: | Gli1_v015681_primary |
| Species: | Mus musculus |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0597695 |
Alignment:
BBBDVVRGACCACCCAVGABBAB
---------CCAGYYHVADCCRG
| Original motif | Reverse complement motif |
| Consensus sequence: BBBDVVRGACCACCCAVGABBAB | Consensus sequence: BTBVTCVTGGGTGGTCMVVDVBB |
 |
 |
| Motif ID: | UP00539 |
| Motif name: | Gli2_v015681_secondary |
| Species: | Mus musculus |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0598357 |
Alignment:
HVGVSAGGAGGGTCYHHHHYVH
-------CKGGDTBDMMCTGG-
| Original motif | Reverse complement motif |
| Consensus sequence: HVGVSAGGAGGGTCYHHHHYVH | Consensus sequence: DVMHHDHKGACCCTCCTSVCBH |
 |
 |