Best Matches in Database for Each Motif (Highest to Lowest)
| Dataset #: 1 | Motif ID: 1 | Motif name: Motif 1 |
| Original motif | Reverse complement motif |
| Consensus sequence: GGCGGGGC | Consensus sequence: GCCCCGCC |
 |
 |
Best Matches for Motif ID 1 (Highest to Lowest)
| Motif ID: | MA0290.1 |
| Motif name: | DAL81 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 11 |
| Number of overlap: | 8 |
| Similarity score: | 0.0497454 |
Alignment:
AAAAGCCGCGGGCGGGATT
----------GGCGGGGC-
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAGCCGCGGGCGGGATT | Consensus sequence: AATCCCGCCCGCGGCTTTT |
 |
 |
| Motif ID: | MA0283.1 |
| Motif name: | CHA4 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0552504 |
Alignment:
GGCGGAGW
GGCGGGGC
| Original motif | Reverse complement motif |
| Consensus sequence: GGCGGAGW | Consensus sequence: WCTCCGCC |
 |
 |
| Motif ID: | MA0344.1 |
| Motif name: | NHP10 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0635389 |
Alignment:
TCCCCGGC
GCCCCGCC
| Original motif | Reverse complement motif |
| Consensus sequence: GCCGGGGA | Consensus sequence: TCCCCGGC |
 |
 |
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0654186 |
Alignment:
VCCCCDCHHHDDBD
GCCCCGCC------
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Motif ID: | MA0431.1 |
| Motif name: | TDA9 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 8 |
| Similarity score: | 0.0668666 |
Alignment:
VCCCCDCWH
GCCCCGCC-
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCWH | Consensus sequence: DWGHGGGGB |
 |
 |
| Dataset #: 2 | Motif ID: 2 | Motif name: Motif 2 |
| Original motif | Reverse complement motif |
| Consensus sequence: RGRAGARRGARRAR | Consensus sequence: MTMMTCMMTCTKCK |
 |
 |
Best Matches for Motif ID 2 (Highest to Lowest)
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0351315 |
Alignment:
HHVHDHATMGAAGAAAHDWDW
-----RGRAGARRGARRAR--
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0308.1 |
| Motif name: | GSM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0437372 |
Alignment:
HDDVVTCCGGAGDTWDDDHVH
MTMMTCMMTCTKCK-------
| Original motif | Reverse complement motif |
| Consensus sequence: HBHDDDWADCTCCGGAVBDDH | Consensus sequence: HDDVVTCCGGAGDTWDDDHVH |
 |
 |
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0450939 |
Alignment:
DWVDVVGCACGTGCTDVBDB
-MTMMTCMMTCTKCK-----
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0451102 |
Alignment:
HHTHHHWATATATWWWRDDV
MTMMTCMMTCTKCK------
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Motif ID: | MA0413.1 |
| Motif name: | USV1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0452242 |
Alignment:
DDDTTMCCCCTGAAHHDBDV
MTMMTCMMTCTKCK------
| Original motif | Reverse complement motif |
| Consensus sequence: DDDTTMCCCCTGAAHHDBDV | Consensus sequence: VDBDHHTTCAGGGGRAADDD |
 |
 |
| Dataset #: 2 | Motif ID: 3 | Motif name: Motif 3 |
| Original motif | Reverse complement motif |
| Consensus sequence: AWAAAWTWAAASWA | Consensus sequence: TWSTTTWAWTTTWT |
 |
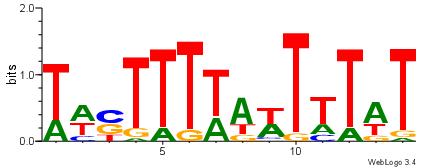 |
Best Matches for Motif ID 3 (Highest to Lowest)
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0383354 |
Alignment:
BHDKWWWATATATWHHHAHH
----AWAAAWTWAAASWA--
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0440227 |
Alignment:
BDACCTWTAATTAWAHBWVVD
----AWAAAWTWAAASWA---
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0465457 |
Alignment:
VHDHHYWHTATATAADDDHDH
-------AWAAAWTWAAASWA
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0479921 |
Alignment:
HHVHDHATMGAAGAAAHDWDW
------AWAAAWTWAAASWA-
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0507283 |
Alignment:
BHHDVTTTGTTTACAWWWHV
------TWSTTTWAWTTTWT
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Dataset #: 2 | Motif ID: 4 | Motif name: Motif 4 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAWTTRCWT | Consensus sequence: AWGKAAWTTTT |
 |
 |
Best Matches for Motif ID 4 (Highest to Lowest)
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.00483292 |
Alignment:
GBYHAAAWTTTTTCACTBHDD
-AWGKAAWTTTT---------
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0107712 |
Alignment:
DHDDDRAAAAWTTTTYYHBDD
------AAAAWTTRCWT----
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 11 |
| Similarity score: | 0.0282258 |
Alignment:
WDWDHTTTCTTCYATHHHVHH
AWGKAAWTTTT----------
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0336.1 |
| Motif name: | MGA1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 11 |
| Similarity score: | 0.0394154 |
Alignment:
BHDHDDKTGTTCTATVDBVHD
AWGKAAWTTTT----------
| Original motif | Reverse complement motif |
| Consensus sequence: DDVVHBATAGAACAYDHHDHV | Consensus sequence: BHDHDDKTGTTCTATVDBVHD |
 |
 |
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 11 |
| Similarity score: | 0.0409802 |
Alignment:
BHDKWWWATATATWHHHAHH
AWGKAAWTTTT---------
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Dataset #: 2 | Motif ID: 5 | Motif name: Motif 5 |
| Original motif | Reverse complement motif |
| Consensus sequence: AWTAAATAYAATTT | Consensus sequence: AAATTKTATTTAWT |
 |
 |
Best Matches for Motif ID 5 (Highest to Lowest)
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0787324 |
Alignment:
HHBHMKAAAAWTTTTKDHDHD
-------AAATTKTATTTAWT
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0386.1 |
| Motif name: | SPT15 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0787361 |
Alignment:
DHVSDATATATATWDCTHDHV
----AWTAAATAYAATTT---
| Original motif | Reverse complement motif |
| Consensus sequence: VHHHAGDWATATATATDSVHD | Consensus sequence: DHVSDATATATATWDCTHDHV |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0815545 |
Alignment:
HHHDDDTTATATAHWKDHHHB
-AAATTKTATTTAWT------
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0932265 |
Alignment:
BHHDVTTTGTTTACAWWWHV
-AAATTKTATTTAWT-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0950567 |
Alignment:
DVVWVHTWTAATTAWAGGTDV
------AWTAAATAYAATTT-
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |
| Dataset #: 2 | Motif ID: 6 | Motif name: Motif 6 |
| Original motif | Reverse complement motif |
| Consensus sequence: AATTYDGAARTAWW | Consensus sequence: WWTAKTTCDKAATT |
 |
 |
Best Matches for Motif ID 6 (Highest to Lowest)
| Motif ID: | MA0400.1 |
| Motif name: | SUT2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0673673 |
Alignment:
DVDHHTTCGGAGTTKHHHDH
WWTAKTTCDKAATT------
| Original motif | Reverse complement motif |
| Consensus sequence: HDDHHRAACTCCGAAHHDBD | Consensus sequence: DVDHHTTCGGAGTTKHHHDH |
 |
 |
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0693729 |
Alignment:
HHVHDHATMGAAGAAAHDWDW
---AATTYDGAARTAWW----
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0415.1 |
| Motif name: | YAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0741794 |
Alignment:
HBDDDMTTACGTAAKVHHVH
--WWTAKTTCDKAATT----
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDMTTACGTAAKVHHVH | Consensus sequence: HVDHVYTTACGTAARHDDVH |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0778093 |
Alignment:
BDWWWTGTAAACAAAVDDDV
-AATTYDGAARTAWW-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0793696 |
Alignment:
GBYHAAAWTTTTTCACTBHDD
------AATTYDGAARTAWW-
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Dataset #: 2 | Motif ID: 7 | Motif name: Motif 7 |
| Original motif | Reverse complement motif |
| Consensus sequence: CSKCCCCGCCCCSY | Consensus sequence: MSGGGGCGGGGYSG |
 |
 |
Best Matches for Motif ID 7 (Highest to Lowest)
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0478852 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
---MSGGGGCGGGGYSG---
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0351.1 |
| Motif name: | DOT6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0530075 |
Alignment:
DBBBDBWSCTCATCGCVHBBD
-------CSKCCCCGCCCCSY
| Original motif | Reverse complement motif |
| Consensus sequence: DBBBDBWSCTCATCGCVHBBD | Consensus sequence: DBBHBGCGATGAGSWBHVBVD |
 |
 |
| Motif ID: | MA0350.1 |
| Motif name: | TOD6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0615081 |
Alignment:
DBBHDHASCTCATCGCVHHHD
-------CSKCCCCGCCCCSY
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHDHASCTCATCGCVHHHD | Consensus sequence: DHHHVGCGATGAGSTDDDBBD |
 |
 |
| Motif ID: | MA0269.1 |
| Motif name: | AFT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0669459 |
Alignment:
VBWDBSGGGTGCARTAHBDVV
-----MSGGGGCGGGGYSG--
| Original motif | Reverse complement motif |
| Consensus sequence: BBDBHTAKTGCACCCSVDWBV | Consensus sequence: VBWDBSGGGTGCARTAHBDVV |
 |
 |
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0706512 |
Alignment:
DBDDHHHGDGGGGB
MSGGGGCGGGGYSG
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Dataset #: 2 | Motif ID: 8 | Motif name: Motif 8 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAATRWTAAAATCA | Consensus sequence: TGATTTTAWKATTT |
 |
 |
Best Matches for Motif ID 8 (Highest to Lowest)
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0751523 |
Alignment:
HHHBAGTGAAAAAWTTTDKVC
---AAATRWTAAAATCA----
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0435.1 |
| Motif name: | YPR015C |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.075789 |
Alignment:
TBDHDACGTAAATCMTDDHH
-AAATRWTAAAATCA-----
| Original motif | Reverse complement motif |
| Consensus sequence: TBDHDACGTAAATCMTDDHH | Consensus sequence: HDDDARGATTTACGTHHDBA |
 |
 |
| Motif ID: | MA0369.1 |
| Motif name: | RLM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 13 |
| Similarity score: | 0.564575 |
Alignment:
VKMYCTATWWWTAGAAHB-
-TGATTTTAWKATTT----
| Original motif | Reverse complement motif |
| Consensus sequence: VDTTCTAWWWATAGMYYV | Consensus sequence: VKMYCTATWWWTAGAAHB |
 |
 |
| Motif ID: | MA0407.1 |
| Motif name: | THI2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 13 |
| Similarity score: | 0.570221 |
Alignment:
-GGMAACYSWAAGARC
-AAATRWTAAAATCA-
| Original motif | Reverse complement motif |
| Consensus sequence: GGMAACYSWAAGARC | Consensus sequence: GKTCTTWSMGTTYCC |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 13 |
| Similarity score: | 0.575864 |
Alignment:
BDWWWTGTAAACAAAVDDDV-
-AAATRWTAAAATCA------
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Dataset #: 2 | Motif ID: 9 | Motif name: Motif 9 |
| Original motif | Reverse complement motif |
| Consensus sequence: AATHATATWTHAAA | Consensus sequence: TTTDAWATATHATT |
 |
 |
Best Matches for Motif ID 9 (Highest to Lowest)
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0354375 |
Alignment:
HHHDDDTTATATAHWKDHHHB
----TTTDAWATATHATT---
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.041212 |
Alignment:
HHTHHHWATATATWWWRDDV
---AATHATATWTHAAA---
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0513433 |
Alignment:
DVVWVHTWTAATTAWAGGTDV
-----TTTDAWATATHATT--
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |
| Motif ID: | MA0386.1 |
| Motif name: | SPT15 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0562045 |
Alignment:
VHHHAGDWATATATATDSVHD
------TTTDAWATATHATT-
| Original motif | Reverse complement motif |
| Consensus sequence: VHHHAGDWATATATATDSVHD | Consensus sequence: DHVSDATATATATWDCTHDHV |
 |
 |
| Motif ID: | MA0369.1 |
| Motif name: | RLM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0619139 |
Alignment:
VDTTCTAWWWATAGMYYV
--TTTDAWATATHATT--
| Original motif | Reverse complement motif |
| Consensus sequence: VDTTCTAWWWATAGMYYV | Consensus sequence: VKMYCTATWWWTAGAAHB |
 |
 |
| Dataset #: 2 | Motif ID: 10 | Motif name: Motif 10 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTTCATAAWT | Consensus sequence: AWTTATGAAA |
 |
 |
Best Matches for Motif ID 10 (Highest to Lowest)
| Motif ID: | MA0415.1 |
| Motif name: | YAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0643778 |
Alignment:
HVDHVYTTACGTAARHDDVH
----AWTTATGAAA------
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDMTTACGTAAKVHHVH | Consensus sequence: HVDHVYTTACGTAARHDDVH |
 |
 |
| Motif ID: | MA0387.1 |
| Motif name: | SPT2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.0680925 |
Alignment:
KTYBTTAAHW
TTTCATAAWT
| Original motif | Reverse complement motif |
| Consensus sequence: WHTTAAVKAR | Consensus sequence: KTYBTTAAHW |
 |
 |
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.0721515 |
Alignment:
HHHBAGTGAAAAAWTTTDKVC
-AWTTATGAAA----------
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0369.1 |
| Motif name: | RLM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0782947 |
Alignment:
VKMYCTATWWWTAGAAHB
------AWTTATGAAA--
| Original motif | Reverse complement motif |
| Consensus sequence: VDTTCTAWWWATAGMYYV | Consensus sequence: VKMYCTATWWWTAGAAHB |
 |
 |
| Motif ID: | MA0418.1 |
| Motif name: | YAP6 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0840819 |
Alignment:
VGKYHGMTTAYGTAAKHRDV
-----AWTTATGAAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: VGKYHGMTTAYGTAAKHRDV | Consensus sequence: VDMDYTTACKTAAYCHKRCV |
 |
 |
| Dataset #: 2 | Motif ID: 11 | Motif name: Motif 11 |
| Original motif | Reverse complement motif |
| Consensus sequence: ATTTWATGAAA | Consensus sequence: TTTCATWAAAT |
 |
 |
Best Matches for Motif ID 11 (Highest to Lowest)
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0.0759198 |
Alignment:
HHHBAGTGAAAAAWTTTDKVC
ATTTWATGAAA----------
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0369.1 |
| Motif name: | RLM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.0779159 |
Alignment:
VKMYCTATWWWTAGAAHB
------ATTTWATGAAA-
| Original motif | Reverse complement motif |
| Consensus sequence: VDTTCTAWWWATAGMYYV | Consensus sequence: VKMYCTATWWWTAGAAHB |
 |
 |
| Motif ID: | MA0415.1 |
| Motif name: | YAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0864254 |
Alignment:
HVDHVYTTACGTAARHDDVH
---ATTTWATGAAA------
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDMTTACGTAAKVHHVH | Consensus sequence: HVDHVYTTACGTAARHDDVH |
 |
 |
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.087477 |
Alignment:
BDACCTWTAATTAWAHBWVVD
-----TTTCATWAAAT-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |
| Motif ID: | MA0407.1 |
| Motif name: | THI2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0885853 |
Alignment:
GGMAACYSWAAGARC
----ATTTWATGAAA
| Original motif | Reverse complement motif |
| Consensus sequence: GGMAACYSWAAGARC | Consensus sequence: GKTCTTWSMGTTYCC |
 |
 |
| Dataset #: 2 | Motif ID: 12 | Motif name: Motif 12 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAAACAAA | Consensus sequence: TTTGTTTT |
 |
 |
Best Matches for Motif ID 12 (Highest to Lowest)
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.00812138 |
Alignment:
BHHDVTTTGTTTACAWWWHV
-----TTTGTTTT-------
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Motif ID: | MA0371.1 |
| Motif name: | ROX1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0247396 |
Alignment:
YCHATTGTTCTC
---TTTGTTTT-
| Original motif | Reverse complement motif |
| Consensus sequence: YCHATTGTTCTC | Consensus sequence: GAGAACAATDGK |
 |
 |
| Motif ID: | MA0277.1 |
| Motif name: | AZF1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0259235 |
Alignment:
AAAAAGAAA
-AAAACAAA
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAAGAAA | Consensus sequence: TTTCTTTTT |
 |
 |
| Motif ID: | MA0343.1 |
| Motif name: | NDT80 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.0382796 |
Alignment:
HHBHHKGMCACAAAAHCSVDV
------AAAACAAA-------
| Original motif | Reverse complement motif |
| Consensus sequence: HHBHHKGMCACAAAAHCSVDV | Consensus sequence: BDVSGHTTTTGTGRCYDDVHH |
 |
 |
| Motif ID: | MA0929.1 |
| Motif name: | NCU00019 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0487695 |
Alignment:
DBDGTAAAYAHD
----AAAACAAA
| Original motif | Reverse complement motif |
| Consensus sequence: DBDGTAAAYAHD | Consensus sequence: BHTKTTTACDDB |
 |
 |
| Dataset #: 2 | Motif ID: 13 | Motif name: Motif 13 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAAGATTT | Consensus sequence: AAATCTTT |
 |
 |
Best Matches for Motif ID 13 (Highest to Lowest)
| Motif ID: | MA0435.1 |
| Motif name: | YPR015C |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 8 |
| Similarity score: | 0.0140145 |
Alignment:
HDDDARGATTTACGTHHDBA
---AAAGATTT---------
| Original motif | Reverse complement motif |
| Consensus sequence: TBDHDACGTAAATCMTDDHH | Consensus sequence: HDDDARGATTTACGTHHDBA |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0232736 |
Alignment:
DHDDDRAAAAWTTTTYYHBDD
------AAAGATTT-------
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.030962 |
Alignment:
GBYHAAAWTTTTTCACTBHDD
-----AAATCTTT--------
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0398.1 |
| Motif name: | SUM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0460574 |
Alignment:
AAAAATWWD
AAAGATTT-
| Original motif | Reverse complement motif |
| Consensus sequence: DWWATTTTT | Consensus sequence: AAAAATWWD |
 |
 |
| Motif ID: | MA0302.1 |
| Motif name: | GAT4 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0477168 |
Alignment:
BBBAGATCTHB
---AAATCTTT
| Original motif | Reverse complement motif |
| Consensus sequence: VHAGATCTVVH | Consensus sequence: BBBAGATCTHB |
 |
 |
| Dataset #: 2 | Motif ID: 14 | Motif name: Motif 14 |
| Original motif | Reverse complement motif |
| Consensus sequence: AWAAATAA | Consensus sequence: TTATTTWT |
 |
 |
Best Matches for Motif ID 14 (Highest to Lowest)
| Motif ID: | MA0369.1 |
| Motif name: | RLM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0179286 |
Alignment:
VKMYCTATWWWTAGAAHB
----TTATTTWT------
| Original motif | Reverse complement motif |
| Consensus sequence: VDTTCTAWWWATAGMYYV | Consensus sequence: VKMYCTATWWWTAGAAHB |
 |
 |
| Motif ID: | MA0398.1 |
| Motif name: | SUM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 8 |
| Similarity score: | 0.0179741 |
Alignment:
DWWATTTTT
-TTATTTWT
| Original motif | Reverse complement motif |
| Consensus sequence: DWWATTTTT | Consensus sequence: AAAAATWWD |
 |
 |
| Motif ID: | MA0433.1 |
| Motif name: | YOX1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0225467 |
Alignment:
TTAATTAA
AWAAATAA
| Original motif | Reverse complement motif |
| Consensus sequence: TTAATTAA | Consensus sequence: TTAATTAA |
 |
 |
| Motif ID: | MA0317.1 |
| Motif name: | HCM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0233838 |
Alignment:
ATAAACAA
AWAAATAA
| Original motif | Reverse complement motif |
| Consensus sequence: ATAAACAA | Consensus sequence: TTGTTTAT |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0238432 |
Alignment:
BDWWWTGTAAACAAAVDDDV
------AWAAATAA------
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Dataset #: 2 | Motif ID: 15 | Motif name: Motif 15 |
| Original motif | Reverse complement motif |
| Consensus sequence: ATMACAATAAAA | Consensus sequence: TTTTATTGTYAT |
 |
 |
Best Matches for Motif ID 15 (Highest to Lowest)
| Motif ID: | MA0314.1 |
| Motif name: | HAP3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 12 |
| Similarity score: | 0.106236 |
Alignment:
TMKVMMCCAATSAGA
---ATMACAATAAAA
| Original motif | Reverse complement motif |
| Consensus sequence: TCTSATTGGYYVRRA | Consensus sequence: TMKVMMCCAATSAGA |
 |
 |
| Motif ID: | MA0440.1 |
| Motif name: | ZAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 12 |
| Similarity score: | 0.109512 |
Alignment:
ACCYTMAAGGTBATG
--TTTTATTGTYAT-
| Original motif | Reverse complement motif |
| Consensus sequence: ACCYTMAAGGTBATG | Consensus sequence: CATBACCTTYAMGGT |
 |
 |
| Motif ID: | MA0369.1 |
| Motif name: | RLM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.116071 |
Alignment:
VKMYCTATWWWTAGAAHB
----ATMACAATAAAA--
| Original motif | Reverse complement motif |
| Consensus sequence: VDTTCTAWWWATAGMYYV | Consensus sequence: VKMYCTATWWWTAGAAHB |
 |
 |
| Motif ID: | MA0343.1 |
| Motif name: | NDT80 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 12 |
| Similarity score: | 0.116213 |
Alignment:
BDVSGHTTTTGTGRCYDDVHH
------TTTTATTGTYAT---
| Original motif | Reverse complement motif |
| Consensus sequence: HHBHHKGMCACAAAAHCSVDV | Consensus sequence: BDVSGHTTTTGTGRCYDDVHH |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.122106 |
Alignment:
BHHDVTTTGTTTACAWWWHV
----TTTTATTGTYAT----
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Dataset #: 2 | Motif ID: 16 | Motif name: Motif 16 |
| Original motif | Reverse complement motif |
| Consensus sequence: ACAAWTRATTTTGA | Consensus sequence: TCAAAATKAWTTGT |
 |
 |
Best Matches for Motif ID 16 (Highest to Lowest)
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0889454 |
Alignment:
HHHBAGTGAAAAAWTTTDKVC
------TCAAAATKAWTTGT-
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0980629 |
Alignment:
HHBHMKAAAAWTTTTKDHDHD
----TCAAAATKAWTTGT---
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0343.1 |
| Motif name: | NDT80 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 12 |
| Similarity score: | 1.0933 |
Alignment:
--BDVSGHTTTTGTGRCYDDVHH
ACAAWTRATTTTGA---------
| Original motif | Reverse complement motif |
| Consensus sequence: HHBHHKGMCACAAAAHCSVDV | Consensus sequence: BDVSGHTTTTGTGRCYDDVHH |
 |
 |
| Motif ID: | MA0440.1 |
| Motif name: | ZAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 1.59678 |
Alignment:
---CATBACCTTYAMGGT
-TCAAAATKAWTTGT---
| Original motif | Reverse complement motif |
| Consensus sequence: ACCYTMAAGGTBATG | Consensus sequence: CATBACCTTYAMGGT |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 2.07585 |
Alignment:
----BDWWWTGTAAACAAAVDDDV
------ACAAWTRATTTTGA----
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Dataset #: 2 | Motif ID: 17 | Motif name: Motif 17 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAATGAAT | Consensus sequence: ATTCATTTTT |
 |
 |
Best Matches for Motif ID 17 (Highest to Lowest)
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 10 |
| Similarity score: | 0.0496392 |
Alignment:
HHHBAGTGAAAAAWTTTDKVC
--------AAAAATGAAT---
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0303.1 |
| Motif name: | GCN4 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 10 |
| Similarity score: | 0.0605292 |
Alignment:
DVHHDHMTGASTCAKHHHHBB
---------ATTCATTTTT--
| Original motif | Reverse complement motif |
| Consensus sequence: BVHDDDRTGASTCAYHHHHVD | Consensus sequence: DVHHDHMTGASTCAKHHHHBB |
 |
 |
| Motif ID: | MA0343.1 |
| Motif name: | NDT80 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.0620677 |
Alignment:
HHBHHKGMCACAAAAHCSVDV
-----------AAAAATGAAT
| Original motif | Reverse complement motif |
| Consensus sequence: HHBHHKGMCACAAAAHCSVDV | Consensus sequence: BDVSGHTTTTGTGRCYDDVHH |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0653327 |
Alignment:
HHBHMKAAAAWTTTTKDHDHD
------AAAAATGAAT-----
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0369.1 |
| Motif name: | RLM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0663763 |
Alignment:
VKMYCTATWWWTAGAAHB
--ATTCATTTTT------
| Original motif | Reverse complement motif |
| Consensus sequence: VDTTCTAWWWATAGMYYV | Consensus sequence: VKMYCTATWWWTAGAAHB |
 |
 |
| Dataset #: 2 | Motif ID: 18 | Motif name: Motif 18 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTAGWTWATAA | Consensus sequence: TTATWAWCTAA |
 |
 |
Best Matches for Motif ID 18 (Highest to Lowest)
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0529918 |
Alignment:
BDWWWTGTAAACAAAVDDDV
----TTATWAWCTAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Motif ID: | MA0415.1 |
| Motif name: | YAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0535956 |
Alignment:
HBDDDMTTACGTAAKVHHVH
---TTATWAWCTAA------
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDMTTACGTAAKVHHVH | Consensus sequence: HVDHVYTTACGTAARHDDVH |
 |
 |
| Motif ID: | MA0418.1 |
| Motif name: | YAP6 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0641812 |
Alignment:
VGKYHGMTTAYGTAAKHRDV
----TTATWAWCTAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: VGKYHGMTTAYGTAAKHRDV | Consensus sequence: VDMDYTTACKTAAYCHKRCV |
 |
 |
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.0671782 |
Alignment:
BHDKWWWATATATWHHHAHH
-------TTATWAWCTAA--
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.0678808 |
Alignment:
HHHDDDTTATATAHWKDHHHB
---TTATWAWCTAA-------
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Dataset #: 2 | Motif ID: 19 | Motif name: Motif 19 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTCWTAGATTAWA | Consensus sequence: TWTAATCTAWGAA |
 |
 |
Best Matches for Motif ID 19 (Highest to Lowest)
| Motif ID: | MA0407.1 |
| Motif name: | THI2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 13 |
| Similarity score: | 0.0781943 |
Alignment:
GKTCTTWSMGTTYCC
-TTCWTAGATTAWA-
| Original motif | Reverse complement motif |
| Consensus sequence: GGMAACYSWAAGARC | Consensus sequence: GKTCTTWSMGTTYCC |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 13 |
| Similarity score: | 0.0790099 |
Alignment:
BDWWWTGTAAACAAAVDDDV
-----TWTAATCTAWGAA--
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Motif ID: | MA0369.1 |
| Motif name: | RLM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 13 |
| Similarity score: | 0.0859661 |
Alignment:
VDTTCTAWWWATAGMYYV
--TTCWTAGATTAWA---
| Original motif | Reverse complement motif |
| Consensus sequence: VDTTCTAWWWATAGMYYV | Consensus sequence: VKMYCTATWWWTAGAAHB |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 13 |
| Similarity score: | 0.0862971 |
Alignment:
VHDHHYWHTATATAADDDHDH
----TTCWTAGATTAWA----
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0400.1 |
| Motif name: | SUT2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 13 |
| Similarity score: | 0.0872006 |
Alignment:
DVDHHTTCGGAGTTKHHHDH
-----TTCWTAGATTAWA--
| Original motif | Reverse complement motif |
| Consensus sequence: HDDHHRAACTCCGAAHHDBD | Consensus sequence: DVDHHTTCGGAGTTKHHHDH |
 |
 |
| Dataset #: 2 | Motif ID: 20 | Motif name: Motif 20 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTMAAAGATTT | Consensus sequence: AAATCTTTYAA |
 |
 |
Best Matches for Motif ID 20 (Highest to Lowest)
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.0477023 |
Alignment:
GBYHAAAWTTTTTCACTBHDD
-----AAATCTTTYAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0435.1 |
| Motif name: | YPR015C |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0.0501411 |
Alignment:
TBDHDACGTAAATCMTDDHH
TTMAAAGATTT---------
| Original motif | Reverse complement motif |
| Consensus sequence: TBDHDACGTAAATCMTDDHH | Consensus sequence: HDDDARGATTTACGTHHDBA |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0548685 |
Alignment:
DHDDDRAAAAWTTTTYYHBDD
-------AAATCTTTYAA---
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0407.1 |
| Motif name: | THI2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0682282 |
Alignment:
GKTCTTWSMGTTYCC
-TTMAAAGATTT---
| Original motif | Reverse complement motif |
| Consensus sequence: GGMAACYSWAAGARC | Consensus sequence: GKTCTTWSMGTTYCC |
 |
 |
| Motif ID: | MA0440.1 |
| Motif name: | ZAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0708025 |
Alignment:
ACCYTMAAGGTBATG
---TTMAAAGATTT-
| Original motif | Reverse complement motif |
| Consensus sequence: ACCYTMAAGGTBATG | Consensus sequence: CATBACCTTYAMGGT |
 |
 |
| Dataset #: 2 | Motif ID: 21 | Motif name: Motif 21 |
| Original motif | Reverse complement motif |
| Consensus sequence: ATAAAA | Consensus sequence: TTTTAT |
 |
 |
Best Matches for Motif ID 21 (Highest to Lowest)
| Motif ID: | MA0343.1 |
| Motif name: | NDT80 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.0600039 |
Alignment:
BDVSGHTTTTGTGRCYDDVHH
------TTTTAT---------
| Original motif | Reverse complement motif |
| Consensus sequence: HHBHHKGMCACAAAAHCSVDV | Consensus sequence: BDVSGHTTTTGTGRCYDDVHH |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 6 |
| Similarity score: | 0.0715567 |
Alignment:
HHBHMKAAAAWTTTTKDHDHD
----ATAAAA-----------
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0369.1 |
| Motif name: | RLM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 6 |
| Similarity score: | 0.0744048 |
Alignment:
VDTTCTAWWWATAGMYYV
--TTTTAT----------
| Original motif | Reverse complement motif |
| Consensus sequence: VDTTCTAWWWATAGMYYV | Consensus sequence: VKMYCTATWWWTAGAAHB |
 |
 |
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0.0757497 |
Alignment:
HHHBAGTGAAAAAWTTTDKVC
--------ATAAAA-------
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0309.1 |
| Motif name: | GZF3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 6 |
| Similarity score: | 0.0766832 |
Alignment:
YGATAASV
--ATAAAA
| Original motif | Reverse complement motif |
| Consensus sequence: YGATAASV | Consensus sequence: BSTTATCM |
 |
 |
| Dataset #: 3 | Motif ID: 22 | Motif name: Zfx |
| Original motif | Reverse complement motif |
| Consensus sequence: BBVGCCBVGGCCTV | Consensus sequence: VAGGCCBBGGCVBB |
 |
 |
Best Matches for Motif ID 22 (Highest to Lowest)
| Motif ID: | MA0273.1 |
| Motif name: | ARO80 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0605125 |
Alignment:
GVBVVHKCTCGGBAABDHHDH
--BBVGCCBVGGCCTV-----
| Original motif | Reverse complement motif |
| Consensus sequence: GVBVVHKCTCGGBAABDHHDH | Consensus sequence: HHHHDBTTBCCGAGRHBBVBC |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0610807 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
-----VAGGCCBBGGCVBB-
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0347.1 |
| Motif name: | NRG1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0633057 |
Alignment:
DVCBDWGGACCCTRHDVBHD
------VAGGCCBBGGCVBB
| Original motif | Reverse complement motif |
| Consensus sequence: HHVVDHMAGGGTCCWDBGVD | Consensus sequence: DVCBDWGGACCCTRHDVBHD |
 |
 |
| Motif ID: | MA0278.1 |
| Motif name: | BAS1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0666116 |
Alignment:
BBBWHKTGACTCBGGCKBWGB
-----VAGGCCBBGGCVBB--
| Original motif | Reverse complement motif |
| Consensus sequence: BCWBRGCCVGAGTCARDWBVV | Consensus sequence: BBBWHKTGACTCBGGCKBWGB |
 |
 |
| Motif ID: | MA0351.1 |
| Motif name: | DOT6 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0714887 |
Alignment:
DBBHBGCGATGAGSWBHVBVD
-------VAGGCCBBGGCVBB
| Original motif | Reverse complement motif |
| Consensus sequence: DBBBDBWSCTCATCGCVHBBD | Consensus sequence: DBBHBGCGATGAGSWBHVBVD |
 |
 |
| Dataset #: 3 | Motif ID: 23 | Motif name: Egr1 |
| Original motif | Reverse complement motif |
| Consensus sequence: HGCGTGGGCGK | Consensus sequence: YCGCCCACGCH |
 |
 |
Best Matches for Motif ID 23 (Highest to Lowest)
| Motif ID: | MA0290.1 |
| Motif name: | DAL81 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0310356 |
Alignment:
AATCCCGCCCGCGGCTTTT
----YCGCCCACGCH----
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAGCCGCGGGCGGGATT | Consensus sequence: AATCCCGCCCGCGGCTTTT |
 |
 |
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0552565 |
Alignment:
VDVBDAGCACGTGCBBDVWH
----YCGCCCACGCH-----
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0562067 |
Alignment:
VCCCCDCHHHDDBD
---YCGCCCACGCH
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Motif ID: | MA0323.1 |
| Motif name: | IXR1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.056625 |
Alignment:
AARCCGGRAGCGGKG
--HGCGTGGGCGK--
| Original motif | Reverse complement motif |
| Consensus sequence: AARCCGGRAGCGGKG | Consensus sequence: CYCCGCTKCCGGMTT |
 |
 |
| Motif ID: | MA0400.1 |
| Motif name: | SUT2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.059616 |
Alignment:
DVDHHTTCGGAGTTKHHHDH
--HGCGTGGGCGK-------
| Original motif | Reverse complement motif |
| Consensus sequence: HDDHHRAACTCCGAAHHDBD | Consensus sequence: DVDHHTTCGGAGTTKHHHDH |
 |
 |
| Dataset #: 3 | Motif ID: 24 | Motif name: SP1 |
| Original motif | Reverse complement motif |
| Consensus sequence: CCCCKCCCCC | Consensus sequence: GGGGGYGGGG |
 |
 |
Best Matches for Motif ID 24 (Highest to Lowest)
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.0399399 |
Alignment:
VCCCCDCHHHDDBD
-CCCCKCCCCC---
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Motif ID: | MA0323.1 |
| Motif name: | IXR1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0418489 |
Alignment:
AARCCGGRAGCGGKG
-----GGGGGYGGGG
| Original motif | Reverse complement motif |
| Consensus sequence: AARCCGGRAGCGGKG | Consensus sequence: CYCCGCTKCCGGMTT |
 |
 |
| Motif ID: | MA0290.1 |
| Motif name: | DAL81 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.059697 |
Alignment:
AATCCCGCCCGCGGCTTTT
--CCCCKCCCCC-------
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAGCCGCGGGCGGGATT | Consensus sequence: AATCCCGCCCGCGGCTTTT |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0702654 |
Alignment:
HBDHVKKKCGGCGCCBHBVH
---GGGGGYGGGG-------
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0347.1 |
| Motif name: | NRG1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 10 |
| Similarity score: | 0.0720885 |
Alignment:
HHVVDHMAGGGTCCWDBGVD
-GGGGGYGGGG---------
| Original motif | Reverse complement motif |
| Consensus sequence: HHVVDHMAGGGTCCWDBGVD | Consensus sequence: DVCBDWGGACCCTRHDVBHD |
 |
 |
| Dataset #: 3 | Motif ID: 25 | Motif name: TFAP2A |
| Original motif | Reverse complement motif |
| Consensus sequence: GCCBBVRGS | Consensus sequence: SCMVBBGGC |
 |
 |
Best Matches for Motif ID 25 (Highest to Lowest)
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 12 |
| Number of overlap: | 9 |
| Similarity score: | 0.0523356 |
Alignment:
HBDHVKKKCGGCGCCBHBVH
SCMVBBGGC-----------
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0269.1 |
| Motif name: | AFT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 13 |
| Number of overlap: | 9 |
| Similarity score: | 0.0531916 |
Alignment:
VBWDBSGGGTGCARTAHBDVV
SCMVBBGGC------------
| Original motif | Reverse complement motif |
| Consensus sequence: BBDBHTAKTGCACCCSVDWBV | Consensus sequence: VBWDBSGGGTGCARTAHBDVV |
 |
 |
| Motif ID: | MA0347.1 |
| Motif name: | NRG1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 9 |
| Similarity score: | 0.0537997 |
Alignment:
DVCBDWGGACCCTRHDVBHD
-SCMVBBGGC----------
| Original motif | Reverse complement motif |
| Consensus sequence: HHVVDHMAGGGTCCWDBGVD | Consensus sequence: DVCBDWGGACCCTRHDVBHD |
 |
 |
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 9 |
| Similarity score: | 0.0555572 |
Alignment:
VCCCCDCHHHDDBD
-----GCCBBVRGS
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Motif ID: | MA0343.1 |
| Motif name: | NDT80 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 13 |
| Number of overlap: | 9 |
| Similarity score: | 0.0585669 |
Alignment:
HHBHHKGMCACAAAAHCSVDV
SCMVBBGGC------------
| Original motif | Reverse complement motif |
| Consensus sequence: HHBHHKGMCACAAAAHCSVDV | Consensus sequence: BDVSGHTTTTGTGRCYDDVHH |
 |
 |
| Dataset #: 3 | Motif ID: 26 | Motif name: MIZF |
| Original motif | Reverse complement motif |
| Consensus sequence: BAACGTCCGC | Consensus sequence: GCGGACGTTV |
 |
 |
Best Matches for Motif ID 26 (Highest to Lowest)
| Motif ID: | MA0400.1 |
| Motif name: | SUT2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0463449 |
Alignment:
DVDHHTTCGGAGTTKHHHDH
------GCGGACGTTV----
| Original motif | Reverse complement motif |
| Consensus sequence: HDDHHRAACTCCGAAHHDBD | Consensus sequence: DVDHHTTCGGAGTTKHHHDH |
 |
 |
| Motif ID: | MA0439.1 |
| Motif name: | YRR1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.0501225 |
Alignment:
VTTATHTCCGY
-BAACGTCCGC
| Original motif | Reverse complement motif |
| Consensus sequence: VTTATHTCCGY | Consensus sequence: KCGGADATAAV |
 |
 |
| Motif ID: | MA0308.1 |
| Motif name: | GSM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.052838 |
Alignment:
HDDVVTCCGGAGDTWDDDHVH
------GCGGACGTTV-----
| Original motif | Reverse complement motif |
| Consensus sequence: HBHDDDWADCTCCGGAVBDDH | Consensus sequence: HDDVVTCCGGAGDTWDDDHVH |
 |
 |
| Motif ID: | MA0422.1 |
| Motif name: | URC2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.0613126 |
Alignment:
HCGGAVWTAH
BAACGTCCGC
| Original motif | Reverse complement motif |
| Consensus sequence: HCGGAVWTAH | Consensus sequence: HTAWVTCCGD |
 |
 |
| Motif ID: | MA0367.1 |
| Motif name: | RGT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.0625248 |
Alignment:
HCGGAWDAWDD
GCGGACGTTV-
| Original motif | Reverse complement motif |
| Consensus sequence: DDWTDWTCCGD | Consensus sequence: HCGGAWDAWDD |
 |
 |
| Dataset #: 3 | Motif ID: 27 | Motif name: Klf4 |
| Original motif | Reverse complement motif |
| Consensus sequence: DGGGYGKGGC | Consensus sequence: GCCYCMCCCD |
 |
 |
Best Matches for Motif ID 27 (Highest to Lowest)
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0377858 |
Alignment:
VCCCCDCHHHDDBD
GCCYCMCCCD----
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.0479742 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
---DGGGYGKGGC-------
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0290.1 |
| Motif name: | DAL81 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 10 |
| Similarity score: | 0.0512896 |
Alignment:
AAAAGCCGCGGGCGGGATT
--------DGGGYGKGGC-
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAGCCGCGGGCGGGATT | Consensus sequence: AATCCCGCCCGCGGCTTTT |
 |
 |
| Motif ID: | MA0413.1 |
| Motif name: | USV1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 10 |
| Similarity score: | 0.0518653 |
Alignment:
DDDTTMCCCCTGAAHHDBDV
-GCCYCMCCCD---------
| Original motif | Reverse complement motif |
| Consensus sequence: DDDTTMCCCCTGAAHHDBDV | Consensus sequence: VDBDHHTTCAGGGGRAADDD |
 |
 |
| Motif ID: | MA0269.1 |
| Motif name: | AFT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0608491 |
Alignment:
VBWDBSGGGTGCARTAHBDVV
-----DGGGYGKGGC------
| Original motif | Reverse complement motif |
| Consensus sequence: BBDBHTAKTGCACCCSVDWBV | Consensus sequence: VBWDBSGGGTGCARTAHBDVV |
 |
 |
| Dataset #: 3 | Motif ID: 28 | Motif name: E2F1 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTTSGCGC | Consensus sequence: GCGCSAAA |
 |
 |
Best Matches for Motif ID 28 (Highest to Lowest)
| Motif ID: | MA0401.1 |
| Motif name: | SWI4 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.047828 |
Alignment:
ACGCGAAA
GCGCSAAA
| Original motif | Reverse complement motif |
| Consensus sequence: ACGCGAAA | Consensus sequence: TTTCGCGT |
 |
 |
| Motif ID: | MA0412.1 |
| Motif name: | UME6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0641639 |
Alignment:
AWWTAGCCGCCGA
--TTTSGCGC---
| Original motif | Reverse complement motif |
| Consensus sequence: TCGGCGGCTAWWT | Consensus sequence: AWWTAGCCGCCGA |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0656133 |
Alignment:
HBDHVKKKCGGCGCCBHBVH
------TTTSGCGC------
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0384.1 |
| Motif name: | SNT2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 8 |
| Similarity score: | 0.069239 |
Alignment:
MGGCGCTAMCA
--GCGCSAAA-
| Original motif | Reverse complement motif |
| Consensus sequence: TGRTAGCGCCR | Consensus sequence: MGGCGCTAMCA |
 |
 |
| Motif ID: | MA0396.1 |
| Motif name: | STP3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0748236 |
Alignment:
DGCGCTABH
-GCGCSAAA
| Original motif | Reverse complement motif |
| Consensus sequence: DBTAGCGCD | Consensus sequence: DGCGCTABH |
 |
 |
| Dataset #: 3 | Motif ID: 29 | Motif name: HIF1AARNT |
| Original motif | Reverse complement motif |
| Consensus sequence: VBACGTGV | Consensus sequence: VCACGTBV |
 |
 |
Best Matches for Motif ID 29 (Highest to Lowest)
| Motif ID: | MA0357.1 |
| Motif name: | PHO4 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.00818914 |
Alignment:
SCACGTGS
VBACGTGV
| Original motif | Reverse complement motif |
| Consensus sequence: SCACGTGS | Consensus sequence: SCACGTGS |
 |
 |
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0129728 |
Alignment:
VDVBDAGCACGTGCBBDVWH
------VCACGTBV------
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Motif ID: | MA0281.1 |
| Motif name: | CBF1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0134372 |
Alignment:
GCACGTGA
VBACGTGV
| Original motif | Reverse complement motif |
| Consensus sequence: GCACGTGA | Consensus sequence: TCACGTGC |
 |
 |
| Motif ID: | MA0321.1 |
| Motif name: | INO2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0403872 |
Alignment:
GCATGTGAA
VCACGTBV-
| Original motif | Reverse complement motif |
| Consensus sequence: GCATGTGAA | Consensus sequence: TTCACATGC |
 |
 |
| Motif ID: | MA0286.1 |
| Motif name: | CST6 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 8 |
| Similarity score: | 0.0442236 |
Alignment:
DHACGTCAK
VCACGTBV-
| Original motif | Reverse complement motif |
| Consensus sequence: RTGACGTHD | Consensus sequence: DHACGTCAK |
 |
 |
| Dataset #: 3 | Motif ID: 30 | Motif name: PLAG1 |
| Original motif | Reverse complement motif |
| Consensus sequence: GGGGCCCAAGGGGG | Consensus sequence: CCCCCTTGGGCCCC |
 |
 |
Best Matches for Motif ID 30 (Highest to Lowest)
| Motif ID: | MA0269.1 |
| Motif name: | AFT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.100886 |
Alignment:
VBWDBSGGGTGCARTAHBDVV
CCCCCTTGGGCCCC-------
| Original motif | Reverse complement motif |
| Consensus sequence: BBDBHTAKTGCACCCSVDWBV | Consensus sequence: VBWDBSGGGTGCARTAHBDVV |
 |
 |
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.102193 |
Alignment:
DBDDHHHGDGGGGB
GGGGCCCAAGGGGG
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.103695 |
Alignment:
HBDHVKKKCGGCGCCBHBVH
-CCCCCTTGGGCCCC-----
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0347.1 |
| Motif name: | NRG1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 13 |
| Similarity score: | 0.590746 |
Alignment:
HHVVDHMAGGGTCCWDBGVD-
-------GGGGCCCAAGGGGG
| Original motif | Reverse complement motif |
| Consensus sequence: HHVVDHMAGGGTCCWDBGVD | Consensus sequence: DVCBDWGGACCCTRHDVBHD |
 |
 |
| Motif ID: | MA0323.1 |
| Motif name: | IXR1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 13 |
| Similarity score: | 0.606398 |
Alignment:
-AARCCGGRAGCGGKG
GGGGCCCAAGGGGG--
| Original motif | Reverse complement motif |
| Consensus sequence: AARCCGGRAGCGGKG | Consensus sequence: CYCCGCTKCCGGMTT |
 |
 |
| Dataset #: 3 | Motif ID: 31 | Motif name: Pax5 |
| Original motif | Reverse complement motif |
| Consensus sequence: DGVBCABTGDWGCGKRRCSR | Consensus sequence: MSGKKRCGCWDCABTGBBCD |
 |
 |
Best Matches for Motif ID 31 (Highest to Lowest)
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0474871 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
DGVBCABTGDWGCGKRRCSR
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0351.1 |
| Motif name: | DOT6 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0524078 |
Alignment:
DBBHBGCGATGAGSWBHVBVD
MSGKKRCGCWDCABTGBBCD-
| Original motif | Reverse complement motif |
| Consensus sequence: DBBBDBWSCTCATCGCVHBBD | Consensus sequence: DBBHBGCGATGAGSWBHVBVD |
 |
 |
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0533034 |
Alignment:
WDWDHTTTCTTCYATHHHVHH
MSGKKRCGCWDCABTGBBCD-
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0303.1 |
| Motif name: | GCN4 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0547496 |
Alignment:
DVHHDHMTGASTCAKHHHHBB
-MSGKKRCGCWDCABTGBBCD
| Original motif | Reverse complement motif |
| Consensus sequence: BVHDDDRTGASTCAYHHHHVD | Consensus sequence: DVHHDHMTGASTCAKHHHHBB |
 |
 |
| Motif ID: | MA0273.1 |
| Motif name: | ARO80 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0582152 |
Alignment:
GVBVVHKCTCGGBAABDHHDH
-MSGKKRCGCWDCABTGBBCD
| Original motif | Reverse complement motif |
| Consensus sequence: GVBVVHKCTCGGBAABDHHDH | Consensus sequence: HHHHDBTTBCCGAGRHBBVBC |
 |
 |
| Dataset #: 3 | Motif ID: 32 | Motif name: ArntAhr |
| Original motif | Reverse complement motif |
| Consensus sequence: YGCGTG | Consensus sequence: CACGCM |
 |
 |
Best Matches for Motif ID 32 (Highest to Lowest)
| Motif ID: | MA0310.1 |
| Motif name: | HAC1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 6 |
| Similarity score: | 0.0161785 |
Alignment:
GACACGTV
--CACGCM
| Original motif | Reverse complement motif |
| Consensus sequence: GACACGTV | Consensus sequence: VACGTGTC |
 |
 |
| Motif ID: | MA0270.1 |
| Motif name: | AFT2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 6 |
| Similarity score: | 0.017883 |
Alignment:
BGGGTGTB
YGCGTG--
| Original motif | Reverse complement motif |
| Consensus sequence: BACACCCB | Consensus sequence: BGGGTGTB |
 |
 |
| Motif ID: | MA0295.1 |
| Motif name: | FHL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 6 |
| Similarity score: | 0.0196507 |
Alignment:
GACGCAVA
--YGCGTG
| Original motif | Reverse complement motif |
| Consensus sequence: GACGCAVA | Consensus sequence: TBTGCGTC |
 |
 |
| Motif ID: | MA0281.1 |
| Motif name: | CBF1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 6 |
| Similarity score: | 0.0277434 |
Alignment:
GCACGTGA
-YGCGTG-
| Original motif | Reverse complement motif |
| Consensus sequence: GCACGTGA | Consensus sequence: TCACGTGC |
 |
 |
| Motif ID: | MA0269.1 |
| Motif name: | AFT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 6 |
| Similarity score: | 0.0290394 |
Alignment:
VBWDBSGGGTGCARTAHBDVV
-----YGCGTG----------
| Original motif | Reverse complement motif |
| Consensus sequence: BBDBHTAKTGCACCCSVDWBV | Consensus sequence: VBWDBSGGGTGCARTAHBDVV |
 |
 |
| Dataset #: 3 | Motif ID: 33 | Motif name: Mycn |
| Original motif | Reverse complement motif |
| Consensus sequence: HSCACGTGGC | Consensus sequence: GCCACGTGSD |
 |
 |
Best Matches for Motif ID 33 (Highest to Lowest)
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0219866 |
Alignment:
VDVBDAGCACGTGCBBDVWH
-----HSCACGTGGC-----
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Motif ID: | MA0290.1 |
| Motif name: | DAL81 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0657718 |
Alignment:
AATCCCGCCCGCGGCTTTT
-----HSCACGTGGC----
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAGCCGCGGGCGGGATT | Consensus sequence: AATCCCGCCCGCGGCTTTT |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0693996 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
------GCCACGTGSD----
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0350.1 |
| Motif name: | TOD6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0710797 |
Alignment:
DBBHDHASCTCATCGCVHHHD
------HSCACGTGGC-----
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHDHASCTCATCGCVHHHD | Consensus sequence: DHHHVGCGATGAGSTDDDBBD |
 |
 |
| Motif ID: | MA0351.1 |
| Motif name: | DOT6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0713738 |
Alignment:
DBBBDBWSCTCATCGCVHBBD
------HSCACGTGGC-----
| Original motif | Reverse complement motif |
| Consensus sequence: DBBBDBWSCTCATCGCVHBBD | Consensus sequence: DBBHBGCGATGAGSWBHVBVD |
 |
 |
| Dataset #: 3 | Motif ID: 34 | Motif name: Myc |
| Original motif | Reverse complement motif |
| Consensus sequence: VGCACGTGGH | Consensus sequence: DCCACGTGCV |
 |
 |
Best Matches for Motif ID 34 (Highest to Lowest)
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0185747 |
Alignment:
DWVDVVGCACGTGCTDVBDB
-----VGCACGTGGH-----
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Motif ID: | MA0290.1 |
| Motif name: | DAL81 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0632131 |
Alignment:
AATCCCGCCCGCGGCTTTT
-----VGCACGTGGH----
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAGCCGCGGGCGGGATT | Consensus sequence: AATCCCGCCCGCGGCTTTT |
 |
 |
| Motif ID: | MA0269.1 |
| Motif name: | AFT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0677837 |
Alignment:
BBDBHTAKTGCACCCSVDWBV
--------VGCACGTGGH---
| Original motif | Reverse complement motif |
| Consensus sequence: BBDBHTAKTGCACCCSVDWBV | Consensus sequence: VBWDBSGGGTGCARTAHBDVV |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0717569 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
------DCCACGTGCV----
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0413.1 |
| Motif name: | USV1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0738781 |
Alignment:
DDDTTMCCCCTGAAHHDBDV
----DCCACGTGCV------
| Original motif | Reverse complement motif |
| Consensus sequence: DDDTTMCCCCTGAAHHDBDV | Consensus sequence: VDBDHHTTCAGGGGRAADDD |
 |
 |
| Dataset #: 3 | Motif ID: 35 | Motif name: GABPA |
| Original motif | Reverse complement motif |
| Consensus sequence: CCGGAAGTGVV | Consensus sequence: VVCACTTCCGG |
 |
 |
Best Matches for Motif ID 35 (Highest to Lowest)
| Motif ID: | MA0413.1 |
| Motif name: | USV1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.0573253 |
Alignment:
VDBDHHTTCAGGGGRAADDD
-VVCACTTCCGG--------
| Original motif | Reverse complement motif |
| Consensus sequence: DDDTTMCCCCTGAAHHDBDV | Consensus sequence: VDBDHHTTCAGGGGRAADDD |
 |
 |
| Motif ID: | MA0308.1 |
| Motif name: | GSM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0620377 |
Alignment:
HBHDDDWADCTCCGGAVBDDH
----VVCACTTCCGG------
| Original motif | Reverse complement motif |
| Consensus sequence: HBHDDDWADCTCCGGAVBDDH | Consensus sequence: HDDVVTCCGGAGDTWDDDHVH |
 |
 |
| Motif ID: | MA0415.1 |
| Motif name: | YAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.0640651 |
Alignment:
HVDHVYTTACGTAARHDDVH
-VVCACTTCCGG--------
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDMTTACGTAAKVHHVH | Consensus sequence: HVDHVYTTACGTAARHDDVH |
 |
 |
| Motif ID: | MA0323.1 |
| Motif name: | IXR1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0672647 |
Alignment:
AARCCGGRAGCGGKG
---CCGGAAGTGVV-
| Original motif | Reverse complement motif |
| Consensus sequence: AARCCGGRAGCGGKG | Consensus sequence: CYCCGCTKCCGGMTT |
 |
 |
| Motif ID: | MA0367.1 |
| Motif name: | RGT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0.0701765 |
Alignment:
DDWTDWTCCGD
CCGGAAGTGVV
| Original motif | Reverse complement motif |
| Consensus sequence: DDWTDWTCCGD | Consensus sequence: HCGGAWDAWDD |
 |
 |
| Dataset #: 4 | Motif ID: 36 | Motif name: csGCCCCGCCCCsc |
| Original motif | Reverse complement motif |
| Consensus sequence: HVGCCCCGCCCCBB | Consensus sequence: BBGGGGCGGGGCVD |
 |
 |
Best Matches for Motif ID 36 (Highest to Lowest)
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0443426 |
Alignment:
HBDHVKKKCGGCGCCBHBVH
--BBGGGGCGGGGCVD----
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0351.1 |
| Motif name: | DOT6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0711355 |
Alignment:
DBBBDBWSCTCATCGCVHBBD
-------HVGCCCCGCCCCBB
| Original motif | Reverse complement motif |
| Consensus sequence: DBBBDBWSCTCATCGCVHBBD | Consensus sequence: DBBHBGCGATGAGSWBHVBVD |
 |
 |
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0750988 |
Alignment:
VDVBDAGCACGTGCBBDVWH
---BBGGGGCGGGGCVD---
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Motif ID: | MA0269.1 |
| Motif name: | AFT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0753304 |
Alignment:
VBWDBSGGGTGCARTAHBDVV
-----BBGGGGCGGGGCVD--
| Original motif | Reverse complement motif |
| Consensus sequence: BBDBHTAKTGCACCCSVDWBV | Consensus sequence: VBWDBSGGGTGCARTAHBDVV |
 |
 |
| Motif ID: | MA0350.1 |
| Motif name: | TOD6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0758122 |
Alignment:
DBBHDHASCTCATCGCVHHHD
-------HVGCCCCGCCCCBB
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHDHASCTCATCGCVHHHD | Consensus sequence: DHHHVGCGATGAGSTDDDBBD |
 |
 |
| Dataset #: 4 | Motif ID: 37 | Motif name: tkAAATAATAtw |
| Original motif | Reverse complement motif |
| Consensus sequence: HDAAATAATAHW | Consensus sequence: WHTATTATTTDH |
 |
 |
Best Matches for Motif ID 37 (Highest to Lowest)
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.0286836 |
Alignment:
DVVWVHTWTAATTAWAGGTDV
----WHTATTATTTDH-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 12 |
| Similarity score: | 0.0302485 |
Alignment:
HHTHHHWATATATWWWRDDV
---WHTATTATTTDH-----
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 12 |
| Similarity score: | 0.0314807 |
Alignment:
HHBHMKAAAAWTTTTKDHDHD
------WHTATTATTTDH---
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 12 |
| Similarity score: | 0.0324864 |
Alignment:
VHDHHYWHTATATAADDDHDH
-------HDAAATAATAHW--
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0336.1 |
| Motif name: | MGA1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 12 |
| Similarity score: | 0.0399294 |
Alignment:
DDVVHBATAGAACAYDHHDHV
----HDAAATAATAHW-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDVVHBATAGAACAYDHHDHV | Consensus sequence: BHDHDDKTGTTCTATVDBVHD |
 |
 |
| Dataset #: 4 | Motif ID: 38 | Motif name: cccGCCCCGCCCCsb |
| Original motif | Reverse complement motif |
| Consensus sequence: BCCGCCCCGCCCCBB | Consensus sequence: BBGGGGCGGGGCGGB |
 |
 |
Best Matches for Motif ID 38 (Highest to Lowest)
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 15 |
| Similarity score: | 0.0578957 |
Alignment:
HBDHVKKKCGGCGCCBHBVH
--BBGGGGCGGGGCGGB---
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0351.1 |
| Motif name: | DOT6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.0771995 |
Alignment:
DBBBDBWSCTCATCGCVHBBD
------BCCGCCCCGCCCCBB
| Original motif | Reverse complement motif |
| Consensus sequence: DBBBDBWSCTCATCGCVHBBD | Consensus sequence: DBBHBGCGATGAGSWBHVBVD |
 |
 |
| Motif ID: | MA0269.1 |
| Motif name: | AFT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 15 |
| Similarity score: | 0.0809779 |
Alignment:
VBWDBSGGGTGCARTAHBDVV
-----BBGGGGCGGGGCGGB-
| Original motif | Reverse complement motif |
| Consensus sequence: BBDBHTAKTGCACCCSVDWBV | Consensus sequence: VBWDBSGGGTGCARTAHBDVV |
 |
 |
| Motif ID: | MA0350.1 |
| Motif name: | TOD6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 15 |
| Similarity score: | 0.081315 |
Alignment:
DBBHDHASCTCATCGCVHHHD
------BCCGCCCCGCCCCBB
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHDHASCTCATCGCVHHHD | Consensus sequence: DHHHVGCGATGAGSTDDDBBD |
 |
 |
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 15 |
| Similarity score: | 0.0831289 |
Alignment:
VDVBDAGCACGTGCBBDVWH
---BBGGGGCGGGGCGGB--
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Dataset #: 4 | Motif ID: 39 | Motif name: kCAGCCAATmr |
| Original motif | Reverse complement motif |
| Consensus sequence: DCAGCCAATVR | Consensus sequence: MBATTGGCTGH |
 |
 |
Best Matches for Motif ID 39 (Highest to Lowest)
| Motif ID: | MA0314.1 |
| Motif name: | HAP3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0335999 |
Alignment:
TCTSATTGGYYVRRA
--MBATTGGCTGH--
| Original motif | Reverse complement motif |
| Consensus sequence: TCTSATTGGYYVRRA | Consensus sequence: TMKVMMCCAATSAGA |
 |
 |
| Motif ID: | MA0278.1 |
| Motif name: | BAS1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.0653972 |
Alignment:
BBBWHKTGACTCBGGCKBWGB
--MBATTGGCTGH--------
| Original motif | Reverse complement motif |
| Consensus sequence: BCWBRGCCVGAGTCARDWBVV | Consensus sequence: BBBWHKTGACTCBGGCKBWGB |
 |
 |
| Motif ID: | MA0350.1 |
| Motif name: | TOD6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0758689 |
Alignment:
DBBHDHASCTCATCGCVHHHD
----DCAGCCAATVR------
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHDHASCTCATCGCVHHHD | Consensus sequence: DHHHVGCGATGAGSTDDDBBD |
 |
 |
| Motif ID: | MA0351.1 |
| Motif name: | DOT6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0769469 |
Alignment:
DBBBDBWSCTCATCGCVHBBD
----DCAGCCAATVR------
| Original motif | Reverse complement motif |
| Consensus sequence: DBBBDBWSCTCATCGCVHBBD | Consensus sequence: DBBHBGCGATGAGSWBHVBVD |
 |
 |
| Motif ID: | MA0435.1 |
| Motif name: | YPR015C |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.0778865 |
Alignment:
HDDDARGATTTACGTHHDBA
-----MBATTGGCTGH----
| Original motif | Reverse complement motif |
| Consensus sequence: TBDHDACGTAAATCMTDDHH | Consensus sequence: HDDDARGATTTACGTHHDBA |
 |
 |
| Dataset #: 4 | Motif ID: 40 | Motif name: kcACCTGCAgc |
| Original motif | Reverse complement motif |
| Consensus sequence: BCACCTGCABC | Consensus sequence: GBTGCAGGTGB |
 |
 |
Best Matches for Motif ID 40 (Highest to Lowest)
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0263419 |
Alignment:
DWVDVVGCACGTGCTDVBDB
---GBTGCAGGTGB------
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Motif ID: | MA0413.1 |
| Motif name: | USV1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0285119 |
Alignment:
VDBDHHTTCAGGGGRAADDD
----GBTGCAGGTGB-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDTTMCCCCTGAAHHDBDV | Consensus sequence: VDBDHHTTCAGGGGRAADDD |
 |
 |
| Motif ID: | MA0440.1 |
| Motif name: | ZAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0342832 |
Alignment:
CATBACCTTYAMGGT
--BCACCTGCABC--
| Original motif | Reverse complement motif |
| Consensus sequence: ACCYTMAAGGTBATG | Consensus sequence: CATBACCTTYAMGGT |
 |
 |
| Motif ID: | MA0269.1 |
| Motif name: | AFT1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0417278 |
Alignment:
BBDBHTAKTGCACCCSVDWBV
------GBTGCAGGTGB----
| Original motif | Reverse complement motif |
| Consensus sequence: BBDBHTAKTGCACCCSVDWBV | Consensus sequence: VBWDBSGGGTGCARTAHBDVV |
 |
 |
| Motif ID: | MA0400.1 |
| Motif name: | SUT2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.0531282 |
Alignment:
DVDHHTTCGGAGTTKHHHDH
-----GBTGCAGGTGB----
| Original motif | Reverse complement motif |
| Consensus sequence: HDDHHRAACTCCGAAHHDBD | Consensus sequence: DVDHHTTCGGAGTTKHHHDH |
 |
 |
| Dataset #: 4 | Motif ID: 41 | Motif name: wwAAATAATAtw |
| Original motif | Reverse complement motif |
| Consensus sequence: HDAAATAATADD | Consensus sequence: DDTATTATTTDH |
 |
 |
Best Matches for Motif ID 41 (Highest to Lowest)
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 12 |
| Similarity score: | 0.0160013 |
Alignment:
VHDHHYWHTATATAADDDHDH
-------HDAAATAATADD--
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.0173741 |
Alignment:
BDACCTWTAATTAWAHBWVVD
-----HDAAATAATADD----
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 12 |
| Similarity score: | 0.0175581 |
Alignment:
HHTHHHWATATATWWWRDDV
---DDTATTATTTDH-----
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 12 |
| Similarity score: | 0.0272758 |
Alignment:
HHBHMKAAAAWTTTTKDHDHD
------DDTATTATTTDH---
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0336.1 |
| Motif name: | MGA1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 12 |
| Similarity score: | 0.032139 |
Alignment:
DDVVHBATAGAACAYDHHDHV
----HDAAATAATADD-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDVVHBATAGAACAYDHHDHV | Consensus sequence: BHDHDDKTGTTCTATVDBVHD |
 |
 |
| Dataset #: 4 | Motif ID: 42 | Motif name: sSGTCACGTGACSs |
| Original motif | Reverse complement motif |
| Consensus sequence: SGGTCACGTGACCS | Consensus sequence: SGGTCACGTGACCS |
 |
 |
Best Matches for Motif ID 42 (Highest to Lowest)
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0699483 |
Alignment:
VDVBDAGCACGTGCBBDVWH
---SGGTCACGTGACCS---
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Motif ID: | MA0418.1 |
| Motif name: | YAP6 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0812819 |
Alignment:
VDMDYTTACKTAAYCHKRCV
--SGGTCACGTGACCS----
| Original motif | Reverse complement motif |
| Consensus sequence: VGKYHGMTTAYGTAAKHRDV | Consensus sequence: VDMDYTTACKTAAYCHKRCV |
 |
 |
| Motif ID: | MA0415.1 |
| Motif name: | YAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.088253 |
Alignment:
HVDHVYTTACGTAARHDDVH
---SGGTCACGTGACCS---
| Original motif | Reverse complement motif |
| Consensus sequence: HBDDDMTTACGTAAKVHHVH | Consensus sequence: HVDHVYTTACGTAARHDDVH |
 |
 |
| Motif ID: | MA0343.1 |
| Motif name: | NDT80 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.090505 |
Alignment:
BDVSGHTTTTGTGRCYDDVHH
---SGGTCACGTGACCS----
| Original motif | Reverse complement motif |
| Consensus sequence: HHBHHKGMCACAAAAHCSVDV | Consensus sequence: BDVSGHTTTTGTGRCYDDVHH |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.103029 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
----SGGTCACGTGACCS--
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Dataset #: 4 | Motif ID: 43 | Motif name: wsTACwGTAsw |
| Original motif | Reverse complement motif |
| Consensus sequence: HVTACWGTABD | Consensus sequence: DBTACWGTAVH |
 |
 |
Best Matches for Motif ID 43 (Highest to Lowest)
| Motif ID: | MA0299.1 |
| Motif name: | GAL4 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.04133 |
Alignment:
GAKBACTVTBBHCCG
-DBTACWGTAVH---
| Original motif | Reverse complement motif |
| Consensus sequence: CGGDBVAVAGTVYTC | Consensus sequence: GAKBACTVTBBHCCG |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0.0426543 |
Alignment:
BHHDVTTTGTTTACAWWWHV
---------DBTACWGTAVH
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.045564 |
Alignment:
HHVHDHATMGAAGAAAHDWDW
---HVTACWGTABD-------
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0336.1 |
| Motif name: | MGA1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.0471974 |
Alignment:
BHDHDDKTGTTCTATVDBVHD
-------HVTACWGTABD---
| Original motif | Reverse complement motif |
| Consensus sequence: DDVVHBATAGAACAYDHHDHV | Consensus sequence: BHDHDDKTGTTCTATVDBVHD |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 11 |
| Similarity score: | 0.0507352 |
Alignment:
HHHDDDTTATATAHWKDHHHB
---------DBTACWGTAVH-
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Dataset #: 4 | Motif ID: 44 | Motif name: dhACATTCTkh |
| Original motif | Reverse complement motif |
| Consensus sequence: DHACATTCTGH | Consensus sequence: HCAGAATGTHD |
 |
 |
Best Matches for Motif ID 44 (Highest to Lowest)
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0436478 |
Alignment:
HHVHDHATMGAAGAAAHDWDW
------HCAGAATGTHD----
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0336.1 |
| Motif name: | MGA1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0439501 |
Alignment:
DDVVHBATAGAACAYDHHDHV
------HCAGAATGTHD----
| Original motif | Reverse complement motif |
| Consensus sequence: DDVVHBATAGAACAYDHHDHV | Consensus sequence: BHDHDDKTGTTCTATVDBVHD |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.0443222 |
Alignment:
DHDDDRAAAAWTTTTYYHBDD
-----DHACATTCTGH-----
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Motif ID: | MA0440.1 |
| Motif name: | ZAP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0482357 |
Alignment:
ACCYTMAAGGTBATG
--HCAGAATGTHD--
| Original motif | Reverse complement motif |
| Consensus sequence: ACCYTMAAGGTBATG | Consensus sequence: CATBACCTTYAMGGT |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0504146 |
Alignment:
VHDHHYWHTATATAADDDHDH
----HCAGAATGTHD------
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Dataset #: 4 | Motif ID: 45 | Motif name: wbgTAAATAww |
| Original motif | Reverse complement motif |
| Consensus sequence: DBGTAAATAHD | Consensus sequence: DHTATTTACBD |
 |
 |
Best Matches for Motif ID 45 (Highest to Lowest)
| Motif ID: | MA0929.1 |
| Motif name: | NCU00019 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0 |
Alignment:
DBDGTAAAYAHD
-DBGTAAATAHD
| Original motif | Reverse complement motif |
| Consensus sequence: DBDGTAAAYAHD | Consensus sequence: BHTKTTTACDDB |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0222559 |
Alignment:
HHHDDDTTATATAHWKDHHHB
----DBGTAAATAHD------
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0296.1 |
| Motif name: | FKH1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.024745 |
Alignment:
BDWWWTGTAAACAAAVDDDV
----DBGTAAATAHD-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDWWWTGTAAACAAAVDDDV | Consensus sequence: BHHDVTTTGTTTACAWWWHV |
 |
 |
| Motif ID: | MA0386.1 |
| Motif name: | SPT15 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0327275 |
Alignment:
VHHHAGDWATATATATDSVHD
------DBGTAAATAHD----
| Original motif | Reverse complement motif |
| Consensus sequence: VHHHAGDWATATATATDSVHD | Consensus sequence: DHVSDATATATATWDCTHDHV |
 |
 |
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0338073 |
Alignment:
BHDKWWWATATATWHHHAHH
------DHTATTTACBD---
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Dataset #: 5 | Motif ID: 46 | Motif name: TFW1 |
| Original motif | Reverse complement motif |
| Consensus sequence: GTCGCG | Consensus sequence: CGCGAC |
 |
 |
Best Matches for Motif ID 46 (Highest to Lowest)
| Motif ID: | MA0290.1 |
| Motif name: | DAL81 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.0317132 |
Alignment:
AAAAGCCGCGGGCGGGATT
----GTCGCG---------
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAGCCGCGGGCGGGATT | Consensus sequence: AATCCCGCCCGCGGCTTTT |
 |
 |
| Motif ID: | MA0374.1 |
| Motif name: | RSC3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 6 |
| Similarity score: | 0.035812 |
Alignment:
VVGCGCG
-GTCGCG
| Original motif | Reverse complement motif |
| Consensus sequence: CGCGCVV | Consensus sequence: VVGCGCG |
 |
 |
| Motif ID: | MA0375.1 |
| Motif name: | RSC30 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 6 |
| Similarity score: | 0.0365344 |
Alignment:
CGCGCGSV
--CGCGAC
| Original motif | Reverse complement motif |
| Consensus sequence: VSCGCGCG | Consensus sequence: CGCGCGSV |
 |
 |
| Motif ID: | MA0329.1 |
| Motif name: | MBP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 6 |
| Similarity score: | 0.0377355 |
Alignment:
ACGCGHH
-CGCGAC
| Original motif | Reverse complement motif |
| Consensus sequence: ACGCGHH | Consensus sequence: BHCGCGT |
 |
 |
| Motif ID: | MA0330.1 |
| Motif name: | MBP1::SWI6 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 6 |
| Similarity score: | 0.0502588 |
Alignment:
ACGCGTY
-CGCGAC
| Original motif | Reverse complement motif |
| Consensus sequence: ACGCGTY | Consensus sequence: MACGCGT |
 |
 |
| Dataset #: 5 | Motif ID: 47 | Motif name: TFW2 |
| Original motif | Reverse complement motif |
| Consensus sequence: SCGCGCGG | Consensus sequence: CCGCGCGS |
 |
 |
Best Matches for Motif ID 47 (Highest to Lowest)
| Motif ID: | MA0290.1 |
| Motif name: | DAL81 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0420825 |
Alignment:
AATCCCGCCCGCGGCTTTT
----CCGCGCGS-------
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAGCCGCGGGCGGGATT | Consensus sequence: AATCCCGCCCGCGGCTTTT |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 8 |
| Similarity score: | 0.0502501 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
---------CCGCGCGS---
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0273.1 |
| Motif name: | ARO80 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0651707 |
Alignment:
GVBVVHKCTCGGBAABDHHDH
----SCGCGCGG---------
| Original motif | Reverse complement motif |
| Consensus sequence: GVBVVHKCTCGGBAABDHHDH | Consensus sequence: HHHHDBTTBCCGAGRHBBVBC |
 |
 |
| Motif ID: | MA0357.1 |
| Motif name: | PHO4 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 8 |
| Similarity score: | 0.0659131 |
Alignment:
SCACGTGS
SCGCGCGG
| Original motif | Reverse complement motif |
| Consensus sequence: SCACGTGS | Consensus sequence: SCACGTGS |
 |
 |
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.0667998 |
Alignment:
VCCCCDCHHHDDBD
-CCGCGCGS-----
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Dataset #: 5 | Motif ID: 48 | Motif name: TFW3 |
| Original motif | Reverse complement motif |
| Consensus sequence: CGGCYBCGCG | Consensus sequence: CGCGBMGCCG |
 |
 |
Best Matches for Motif ID 48 (Highest to Lowest)
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0494805 |
Alignment:
HBDHVKKKCGGCGCCBHBVH
--------CGGCYBCGCG--
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0299.1 |
| Motif name: | GAL4 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.0660923 |
Alignment:
GAKBACTVTBBHCCG
-----CGCGBMGCCG
| Original motif | Reverse complement motif |
| Consensus sequence: CGGDBVAVAGTVYTC | Consensus sequence: GAKBACTVTBBHCCG |
 |
 |
| Motif ID: | MA0290.1 |
| Motif name: | DAL81 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0731045 |
Alignment:
AATCCCGCCCGCGGCTTTT
-----CGGCYBCGCG----
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAGCCGCGGGCGGGATT | Consensus sequence: AATCCCGCCCGCGGCTTTT |
 |
 |
| Motif ID: | MA0323.1 |
| Motif name: | IXR1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0735542 |
Alignment:
CYCCGCTKCCGGMTT
--CGGCYBCGCG---
| Original motif | Reverse complement motif |
| Consensus sequence: AARCCGGRAGCGGKG | Consensus sequence: CYCCGCTKCCGGMTT |
 |
 |
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0830377 |
Alignment:
VCCCCDCHHHDDBD
---CGGCYBCGCG-
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Dataset #: 5 | Motif ID: 49 | Motif name: TFF1 |
| Original motif | Reverse complement motif |
| Consensus sequence: CGGVGCCGCVGC | Consensus sequence: GCVGCGGCBCCG |
 |
 |
Best Matches for Motif ID 49 (Highest to Lowest)
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.0423489 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
----CGGVGCCGCVGC----
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0347.1 |
| Motif name: | NRG1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 12 |
| Similarity score: | 0.0692583 |
Alignment:
DVCBDWGGACCCTRHDVBHD
-GCVGCGGCBCCG-------
| Original motif | Reverse complement motif |
| Consensus sequence: HHVVDHMAGGGTCCWDBGVD | Consensus sequence: DVCBDWGGACCCTRHDVBHD |
 |
 |
| Motif ID: | MA0308.1 |
| Motif name: | GSM1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 12 |
| Similarity score: | 0.0733694 |
Alignment:
HDDVVTCCGGAGDTWDDDHVH
-CGGVGCCGCVGC--------
| Original motif | Reverse complement motif |
| Consensus sequence: HBHDDDWADCTCCGGAVBDDH | Consensus sequence: HDDVVTCCGGAGDTWDDDHVH |
 |
 |
| Motif ID: | MA0299.1 |
| Motif name: | GAL4 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 12 |
| Similarity score: | 0.0735331 |
Alignment:
GAKBACTVTBBHCCG
---GCVGCGGCBCCG
| Original motif | Reverse complement motif |
| Consensus sequence: CGGDBVAVAGTVYTC | Consensus sequence: GAKBACTVTBBHCCG |
 |
 |
| Motif ID: | MA0273.1 |
| Motif name: | ARO80 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 12 |
| Similarity score: | 0.074724 |
Alignment:
HHHHDBTTBCCGAGRHBBVBC
---------CGGVGCCGCVGC
| Original motif | Reverse complement motif |
| Consensus sequence: GVBVVHKCTCGGBAABDHHDH | Consensus sequence: HHHHDBTTBCCGAGRHBBVBC |
 |
 |
| Dataset #: 5 | Motif ID: 50 | Motif name: TFF11 |
| Original motif | Reverse complement motif |
| Consensus sequence: GGMGGRGGCGGVGC | Consensus sequence: GCVCCGCCMCCYCC |
 |
 |
Best Matches for Motif ID 50 (Highest to Lowest)
| Motif ID: | MA0425.1 |
| Motif name: | YGR067C |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0442772 |
Alignment:
VCCCCDCHHHDDBD
GCVCCGCCMCCYCC
| Original motif | Reverse complement motif |
| Consensus sequence: VCCCCDCHHHDDBD | Consensus sequence: DBDDHHHGDGGGGB |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0513557 |
Alignment:
HBDHVKKKCGGCGCCBHBVH
GGMGGRGGCGGVGC------
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0376.1 |
| Motif name: | RTG3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.065682 |
Alignment:
VDVBDAGCACGTGCBBDVWH
GGMGGRGGCGGVGC------
| Original motif | Reverse complement motif |
| Consensus sequence: VDVBDAGCACGTGCBBDVWH | Consensus sequence: DWVDVVGCACGTGCTDVBDB |
 |
 |
| Motif ID: | MA0278.1 |
| Motif name: | BAS1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0674206 |
Alignment:
BBBWHKTGACTCBGGCKBWGB
GCVCCGCCMCCYCC-------
| Original motif | Reverse complement motif |
| Consensus sequence: BCWBRGCCVGAGTCARDWBVV | Consensus sequence: BBBWHKTGACTCBGGCKBWGB |
 |
 |
| Motif ID: | MA0350.1 |
| Motif name: | TOD6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0684329 |
Alignment:
DHHHVGCGATGAGSTDDDBBD
GGMGGRGGCGGVGC-------
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHDHASCTCATCGCVHHHD | Consensus sequence: DHHHVGCGATGAGSTDDDBBD |
 |
 |
| Dataset #: 5 | Motif ID: 51 | Motif name: TFM2 |
| Original motif | Reverse complement motif |
| Consensus sequence: RGRGGAGRRGGHGGDG | Consensus sequence: CHCCBCCKMCTCCKCM |
 |
 |
Best Matches for Motif ID 51 (Highest to Lowest)
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 16 |
| Similarity score: | 0.0415781 |
Alignment:
HBDHVKKKCGGCGCCBHBVH
--CHCCBCCKMCTCCKCM--
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0303.1 |
| Motif name: | GCN4 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 16 |
| Similarity score: | 0.0417084 |
Alignment:
DVHHDHMTGASTCAKHHHHBB
-CHCCBCCKMCTCCKCM----
| Original motif | Reverse complement motif |
| Consensus sequence: BVHDDDRTGASTCAYHHHHVD | Consensus sequence: DVHHDHMTGASTCAKHHHHBB |
 |
 |
| Motif ID: | MA0351.1 |
| Motif name: | DOT6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 16 |
| Similarity score: | 0.0417741 |
Alignment:
DBBHBGCGATGAGSWBHVBVD
---RGRGGAGRRGGHGGDG--
| Original motif | Reverse complement motif |
| Consensus sequence: DBBBDBWSCTCATCGCVHBBD | Consensus sequence: DBBHBGCGATGAGSWBHVBVD |
 |
 |
| Motif ID: | MA0350.1 |
| Motif name: | TOD6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 16 |
| Similarity score: | 0.0433269 |
Alignment:
DHHHVGCGATGAGSTDDDBBD
---RGRGGAGRRGGHGGDG--
| Original motif | Reverse complement motif |
| Consensus sequence: DBBHDHASCTCATCGCVHHHD | Consensus sequence: DHHHVGCGATGAGSTDDDBBD |
 |
 |
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 16 |
| Similarity score: | 0.0500315 |
Alignment:
HHVHDHATMGAAGAAAHDWDW
---RGRGGAGRRGGHGGDG--
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Dataset #: 5 | Motif ID: 52 | Motif name: TFM1 |
| Original motif | Reverse complement motif |
| Consensus sequence: WTKTTTTTHWTTTTTTBT | Consensus sequence: ABAAAAAAWHAAAAARAW |
 |
 |
Best Matches for Motif ID 52 (Highest to Lowest)
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 18 |
| Similarity score: | 0.0314002 |
Alignment:
WDWDHTTTCTTCYATHHHVHH
WTKTTTTTHWTTTTTTBT---
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0404368 |
Alignment:
BHDKWWWATATATWHHHAHH
--ABAAAAAAWHAAAAARAW
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 18 |
| Similarity score: | 0.0405522 |
Alignment:
HHHDDDTTATATAHWKDHHHB
WTKTTTTTHWTTTTTTBT---
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 18 |
| Similarity score: | 0.0419477 |
Alignment:
BDACCTWTAATTAWAHBWVVD
WTKTTTTTHWTTTTTTBT---
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |
| Motif ID: | MA0378.1 |
| Motif name: | SFP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 18 |
| Similarity score: | 0.0436589 |
Alignment:
DHDDDRAAAAWTTTTYYHBDD
WTKTTTTTHWTTTTTTBT---
| Original motif | Reverse complement motif |
| Consensus sequence: DHDDDRAAAAWTTTTYYHBDD | Consensus sequence: HHBHMKAAAAWTTTTKDHDHD |
 |
 |
| Dataset #: 5 | Motif ID: 53 | Motif name: TFM3 |
| Original motif | Reverse complement motif |
| Consensus sequence: WWHTTTTTCABAAWTTWA | Consensus sequence: TWAAWTTVTGAAAAAHWW |
 |
 |
Best Matches for Motif ID 53 (Highest to Lowest)
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0343553 |
Alignment:
HHTHHHWATATATWWWRDDV
--TWAAWTTVTGAAAAAHWW
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0370702 |
Alignment:
VHDHHYWHTATATAADDDHDH
--TWAAWTTVTGAAAAAHWW-
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 18 |
| Similarity score: | 0.0412997 |
Alignment:
HHVHDHATMGAAGAAAHDWDW
WWHTTTTTCABAAWTTWA---
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 18 |
| Similarity score: | 0.0471804 |
Alignment:
BDACCTWTAATTAWAHBWVVD
---WWHTTTTTCABAAWTTWA
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |
| Motif ID: | MA0418.1 |
| Motif name: | YAP6 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0519178 |
Alignment:
VDMDYTTACKTAAYCHKRCV
WWHTTTTTCABAAWTTWA--
| Original motif | Reverse complement motif |
| Consensus sequence: VGKYHGMTTAYGTAAKHRDV | Consensus sequence: VDMDYTTACKTAAYCHKRCV |
 |
 |
| Dataset #: 5 | Motif ID: 54 | Motif name: TFM12 |
| Original motif | Reverse complement motif |
| Consensus sequence: CYYCBBCYYYTCCHCCTYYY | Consensus sequence: KKKAGGDGGAKKMGBBGKMG |
 |
 |
Best Matches for Motif ID 54 (Highest to Lowest)
| Motif ID: | MA0278.1 |
| Motif name: | BAS1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0588509 |
Alignment:
BBBWHKTGACTCBGGCKBWGB
KKKAGGDGGAKKMGBBGKMG-
| Original motif | Reverse complement motif |
| Consensus sequence: BCWBRGCCVGAGTCARDWBVV | Consensus sequence: BBBWHKTGACTCBGGCKBWGB |
 |
 |
| Motif ID: | MA0303.1 |
| Motif name: | GCN4 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.060319 |
Alignment:
DVHHDHMTGASTCAKHHHHBB
KKKAGGDGGAKKMGBBGKMG-
| Original motif | Reverse complement motif |
| Consensus sequence: BVHDDDRTGASTCAYHHHHVD | Consensus sequence: DVHHDHMTGASTCAKHHHHBB |
 |
 |
| Motif ID: | MA0395.1 |
| Motif name: | STP2 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0604525 |
Alignment:
HVVHBGGCGCCGYRYVHHBH
CYYCBBCYYYTCCHCCTYYY
| Original motif | Reverse complement motif |
| Consensus sequence: HVVHBGGCGCCGYRYVHHBH | Consensus sequence: HBDHVKKKCGGCGCCBHBVH |
 |
 |
| Motif ID: | MA0273.1 |
| Motif name: | ARO80 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0624127 |
Alignment:
HHHHDBTTBCCGAGRHBBVBC
KKKAGGDGGAKKMGBBGKMG-
| Original motif | Reverse complement motif |
| Consensus sequence: GVBVVHKCTCGGBAABDHHDH | Consensus sequence: HHHHDBTTBCCGAGRHBBVBC |
 |
 |
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0630527 |
Alignment:
HHVHDHATMGAAGAAAHDWDW
CYYCBBCYYYTCCHCCTYYY-
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Dataset #: 5 | Motif ID: 55 | Motif name: TFM13 |
| Original motif | Reverse complement motif |
| Consensus sequence: ATKAAWTTTTRMAABAHHTW | Consensus sequence: WAHHTVTTYKAAAAWTTRAT |
 |
 |
Best Matches for Motif ID 55 (Highest to Lowest)
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0402024 |
Alignment:
VHDHHYWHTATATAADDDHDH
-WAHHTVTTYKAAAAWTTRAT
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0437557 |
Alignment:
HHVHDHATMGAAGAAAHDWDW
WAHHTVTTYKAAAAWTTRAT-
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0465576 |
Alignment:
BDACCTWTAATTAWAHBWVVD
-WAHHTVTTYKAAAAWTTRAT
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0479924 |
Alignment:
GBYHAAAWTTTTTCACTBHDD
-ATKAAWTTTTRMAABAHHTW
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0346.1 |
| Motif name: | NHP6B |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0485472 |
Alignment:
HHTHHHWATATATWWWRDDV
ATKAAWTTTTRMAABAHHTW
| Original motif | Reverse complement motif |
| Consensus sequence: HHTHHHWATATATWWWRDDV | Consensus sequence: BHDKWWWATATATWHHHAHH |
 |
 |
| Dataset #: 5 | Motif ID: 56 | Motif name: TFM11 |
| Original motif | Reverse complement motif |
| Consensus sequence: HDWVAAAHAAAAAMAAAMWWWHBWA | Consensus sequence: TWVHWWWYTTTYTTTTTHTTTVWBH |
 |
 |
Best Matches for Motif ID 56 (Highest to Lowest)
| Motif ID: | MA0377.1 |
| Motif name: | SFL1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 21 |
| Similarity score: | 0.0547631 |
Alignment:
----HHVHDHATMGAAGAAAHDWDW
HDWVAAAHAAAAAMAAAMWWWHBWA-
| Original motif | Reverse complement motif |
| Consensus sequence: HHVHDHATMGAAGAAAHDWDW | Consensus sequence: WDWDHTTTCTTCYATHHHVHH |
 |
 |
| Motif ID: | MA0345.1 |
| Motif name: | NHP6A |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 21 |
| Similarity score: | 0.0599871 |
Alignment:
HHHDDDTTATATAHWKDHHHB----
-TWVHWWWYTTTYTTTTTHTTTVWBH
| Original motif | Reverse complement motif |
| Consensus sequence: VHDHHYWHTATATAADDDHDH | Consensus sequence: HHHDDDTTATATAHWKDHHHB |
 |
 |
| Motif ID: | MA0336.1 |
| Motif name: | MGA1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 21 |
| Similarity score: | 0.0605282 |
Alignment:
DDVVHBATAGAACAYDHHDHV----
----HDWVAAAHAAAAAMAAAMWWWHBWA
| Original motif | Reverse complement motif |
| Consensus sequence: DDVVHBATAGAACAYDHHDHV | Consensus sequence: BHDHDDKTGTTCTATVDBVHD |
 |
 |
| Motif ID: | MA0390.1 |
| Motif name: | STB3 |
| Taxon: | Fungi |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 21 |
| Similarity score: | 0.069776 |
Alignment:
GBYHAAAWTTTTTCACTBHDD----
TWVHWWWYTTTYTTTTTHTTTVWBH
| Original motif | Reverse complement motif |
| Consensus sequence: GBYHAAAWTTTTTCACTBHDD | Consensus sequence: HHHBAGTGAAAAAWTTTDKVC |
 |
 |
| Motif ID: | MA0383.1 |
| Motif name: | SMP1 |
| Taxon: | Fungi |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 21 |
| Similarity score: | 0.0708423 |
Alignment:
BDACCTWTAATTAWAHBWVVD----
HDWVAAAHAAAAAMAAAMWWWHBWA
| Original motif | Reverse complement motif |
| Consensus sequence: BDACCTWTAATTAWAHBWVVD | Consensus sequence: DVVWVHTWTAATTAWAGGTDV |
 |
 |