Top 5 Significant Motifs - Global Matching (Highest to Lowest)
| Dataset #: 3 | Motif ID: 24 | Motif name: SP1 |
| Original motif Consensus sequence: CCCCKCCCCC | Reverse complement motif Consensus sequence: GGGGGYGGGG |
 |
 |
Best Matches for Top Significant Motif ID 24 (Highest to Lowest)
| Dataset #: | 2 |
| Motif ID: | 7 |
| Motif name: | Motif 7 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.00451594 |
Alignment:
CSKCCCCGCCCCSY
---CCCCKCCCCC-
| Original motif Consensus sequence: CSKCCCCGCCCCSY | Reverse complement motif Consensus sequence: MSGGGGCGGGGYSG |
 |
 |
| Dataset #: | 4 |
| Motif ID: | 36 |
| Motif name: | csGCCCCGCCCCsc |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.00965796 |
Alignment:
HVGCCCCGCCCCBB
---CCCCKCCCCC-
| Original motif Consensus sequence: HVGCCCCGCCCCBB | Reverse complement motif Consensus sequence: BBGGGGCGGGGCVD |
 |
 |
| Dataset #: | 4 |
| Motif ID: | 38 |
| Motif name: | cccGCCCCGCCCCsb |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0127292 |
Alignment:
BCCGCCCCGCCCCBB
----CCCCKCCCCC-
| Original motif Consensus sequence: BCCGCCCCGCCCCBB | Reverse complement motif Consensus sequence: BBGGGGCGGGGCGGB |
 |
 |
| Dataset #: | 5 |
| Motif ID: | 50 |
| Motif name: | TFF11 |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0174405 |
Alignment:
GGMGGRGGCGGVGC
---GGGGGYGGGG-
| Original motif Consensus sequence: GGMGGRGGCGGVGC | Reverse complement motif Consensus sequence: GCVCCGCCMCCYCC |
 |
 |
| Dataset #: | 5 |
| Motif ID: | 54 |
| Motif name: | TFM12 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0301548 |
Alignment:
CYYCBBCYYYTCCHCCTYYY
------CCCCKCCCCC----
| Original motif Consensus sequence: CYYCBBCYYYTCCHCCTYYY | Reverse complement motif Consensus sequence: KKKAGGDGGAKKMGBBGKMG |
 |
 |
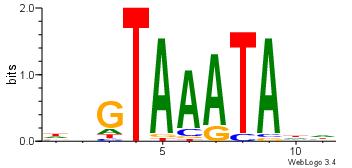
| Dataset #: 4 | Motif ID: 45 | Motif name: wbgTAAATAww |
| Original motif Consensus sequence: DBGTAAATAHD | Reverse complement motif Consensus sequence: DHTATTTACBD |
 |
 |
Best Matches for Top Significant Motif ID 45 (Highest to Lowest)
| Dataset #: | 5 |
| Motif ID: | 56 |
| Motif name: | TFM11 |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 15 |
| Number of overlap: | 11 |
| Similarity score: | 0.0192768 |
Alignment:
TWVHWWWYTTTYTTTTTHTTTVWBH
--------------DHTATTTACBD
| Original motif Consensus sequence: HDWVAAAHAAAAAMAAAMWWWHBWA | Reverse complement motif Consensus sequence: TWVHWWWYTTTYTTTTTHTTTVWBH |
 |
 |
| Dataset #: | 2 |
| Motif ID: | 9 |
| Motif name: | Motif 9 |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0250541 |
Alignment:
AATHATATWTHAAA
DHTATTTACBD---
| Original motif Consensus sequence: AATHATATWTHAAA | Reverse complement motif Consensus sequence: TTTDAWATATHATT |
 |
 |
| Dataset #: | 2 |
| Motif ID: | 6 |
| Motif name: | Motif 6 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0.027895 |
Alignment:
AATTYDGAARTAWW
---DBGTAAATAHD
| Original motif Consensus sequence: AATTYDGAARTAWW | Reverse complement motif Consensus sequence: WWTAKTTCDKAATT |
 |
 |
| Dataset #: | 5 |
| Motif ID: | 52 |
| Motif name: | TFM1 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0.0283217 |
Alignment:
ABAAAAAAWHAAAAARAW
DBGTAAATAHD-------
| Original motif Consensus sequence: WTKTTTTTHWTTTTTTBT | Reverse complement motif Consensus sequence: ABAAAAAAWHAAAAARAW |
 |
 |
| Dataset #: | 5 |
| Motif ID: | 53 |
| Motif name: | TFM3 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.0308097 |
Alignment:
TWAAWTTVTGAAAAAHWW
------DBGTAAATAHD-
| Original motif Consensus sequence: WWHTTTTTCABAAWTTWA | Reverse complement motif Consensus sequence: TWAAWTTVTGAAAAAHWW |
 |
 |
| Dataset #: 3 | Motif ID: 27 | Motif name: Klf4 |
| Original motif Consensus sequence: DGGGYGKGGC | Reverse complement motif Consensus sequence: GCCYCMCCCD |
 |
 |
Best Matches for Top Significant Motif ID 27 (Highest to Lowest)
| Dataset #: | 2 |
| Motif ID: | 7 |
| Motif name: | Motif 7 |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0152457 |
Alignment:
CSKCCCCGCCCCSY
--GCCYCMCCCD--
| Original motif Consensus sequence: CSKCCCCGCCCCSY | Reverse complement motif Consensus sequence: MSGGGGCGGGGYSG |
 |
 |
| Dataset #: | 4 |
| Motif ID: | 36 |
| Motif name: | csGCCCCGCCCCsc |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0160185 |
Alignment:
BBGGGGCGGGGCVD
--DGGGYGKGGC--
| Original motif Consensus sequence: HVGCCCCGCCCCBB | Reverse complement motif Consensus sequence: BBGGGGCGGGGCVD |
 |
 |
| Dataset #: | 4 |
| Motif ID: | 38 |
| Motif name: | cccGCCCCGCCCCsb |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 10 |
| Similarity score: | 0.0194893 |
Alignment:
BBGGGGCGGGGCGGB
--DGGGYGKGGC---
| Original motif Consensus sequence: BCCGCCCCGCCCCBB | Reverse complement motif Consensus sequence: BBGGGGCGGGGCGGB |
 |
 |
| Dataset #: | 5 |
| Motif ID: | 50 |
| Motif name: | TFF11 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0402906 |
Alignment:
GGMGGRGGCGGVGC
----DGGGYGKGGC
| Original motif Consensus sequence: GGMGGRGGCGGVGC | Reverse complement motif Consensus sequence: GCVCCGCCMCCYCC |
 |
 |
| Dataset #: | 5 |
| Motif ID: | 54 |
| Motif name: | TFM12 |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0556427 |
Alignment:
CYYCBBCYYYTCCHCCTYYY
----GCCYCMCCCD------
| Original motif Consensus sequence: CYYCBBCYYYTCCHCCTYYY | Reverse complement motif Consensus sequence: KKKAGGDGGAKKMGBBGKMG |
 |
 |
| Dataset #: 4 | Motif ID: 44 | Motif name: dhACATTCTkh |
| Original motif Consensus sequence: DHACATTCTGH | Reverse complement motif Consensus sequence: HCAGAATGTHD |
 |
 |
Best Matches for Top Significant Motif ID 44 (Highest to Lowest)
| Dataset #: | 5 |
| Motif ID: | 53 |
| Motif name: | TFM3 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.0217516 |
Alignment:
TWAAWTTVTGAAAAAHWW
DHACATTCTGH-------
| Original motif Consensus sequence: WWHTTTTTCABAAWTTWA | Reverse complement motif Consensus sequence: TWAAWTTVTGAAAAAHWW |
 |
 |
| Dataset #: | 2 |
| Motif ID: | 16 |
| Motif name: | Motif 16 |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0258838 |
Alignment:
TCAAAATKAWTTGT
HCAGAATGTHD---
| Original motif Consensus sequence: ACAAWTRATTTTGA | Reverse complement motif Consensus sequence: TCAAAATKAWTTGT |
 |
 |
| Dataset #: | 5 |
| Motif ID: | 56 |
| Motif name: | TFM11 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 11 |
| Similarity score: | 0.0307858 |
Alignment:
TWVHWWWYTTTYTTTTTHTTTVWBH
----DHACATTCTGH----------
| Original motif Consensus sequence: HDWVAAAHAAAAAMAAAMWWWHBWA | Reverse complement motif Consensus sequence: TWVHWWWYTTTYTTTTTHTTTVWBH |
 |
 |
| Dataset #: | 5 |
| Motif ID: | 55 |
| Motif name: | TFM13 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.0357323 |
Alignment:
ATKAAWTTTTRMAABAHHTW
-DHACATTCTGH--------
| Original motif Consensus sequence: ATKAAWTTTTRMAABAHHTW | Reverse complement motif Consensus sequence: WAHHTVTTYKAAAAWTTRAT |
 |
 |
| Dataset #: | 2 |
| Motif ID: | 6 |
| Motif name: | Motif 6 |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0400884 |
Alignment:
AATTYDGAARTAWW
HCAGAATGTHD---
| Original motif Consensus sequence: AATTYDGAARTAWW | Reverse complement motif Consensus sequence: WWTAKTTCDKAATT |
 |
 |
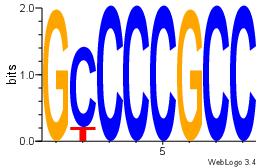
| Dataset #: 1 | Motif ID: 1 | Motif name: Motif 1 |
| Original motif Consensus sequence: GGCGGGGC | Reverse complement motif Consensus sequence: GCCCCGCC |
 |
 |
Best Matches for Top Significant Motif ID 1 (Highest to Lowest)
| Dataset #: | 4 |
| Motif ID: | 36 |
| Motif name: | csGCCCCGCCCCsc |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
BBGGGGCGGGGCVD
----GGCGGGGC--
| Original motif Consensus sequence: HVGCCCCGCCCCBB | Reverse complement motif Consensus sequence: BBGGGGCGGGGCVD |
 |
 |
| Dataset #: | 4 |
| Motif ID: | 38 |
| Motif name: | cccGCCCCGCCCCsb |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.00250079 |
Alignment:
BBGGGGCGGGGCGGB
----GGCGGGGC---
| Original motif Consensus sequence: BCCGCCCCGCCCCBB | Reverse complement motif Consensus sequence: BBGGGGCGGGGCGGB |
 |
 |
| Dataset #: | 2 |
| Motif ID: | 7 |
| Motif name: | Motif 7 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0149958 |
Alignment:
MSGGGGCGGGGYSG
----GGCGGGGC--
| Original motif Consensus sequence: CSKCCCCGCCCCSY | Reverse complement motif Consensus sequence: MSGGGGCGGGGYSG |
 |
 |
| Dataset #: | 5 |
| Motif ID: | 50 |
| Motif name: | TFF11 |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0348969 |
Alignment:
GCVCCGCCMCCYCC
GCCCCGCC------
| Original motif Consensus sequence: GGMGGRGGCGGVGC | Reverse complement motif Consensus sequence: GCVCCGCCMCCYCC |
 |
 |
| Dataset #: | 3 |
| Motif ID: | 27 |
| Motif name: | Klf4 |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 8 |
| Similarity score: | 0.0349983 |
Alignment:
DGGGYGKGGC
--GGCGGGGC
| Original motif Consensus sequence: DGGGYGKGGC | Reverse complement motif Consensus sequence: GCCYCMCCCD |
 |
 |
| Dataset #: | 3 |
| Motif ID: | 22 |
| Motif name: | Zfx |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0708224 |
Alignment:
VAGGCCBBGGCVBB
---GCCCCGCC---
| Original motif Consensus sequence: BBVGCCBVGGCCTV | Reverse complement motif Consensus sequence: VAGGCCBBGGCVBB |
 |
 |