Best Matches in Database for Each Motif (Highest to Lowest)
| Dataset #: 1 | Motif ID: 1 | Motif name: Motif 1 |
| Original motif | Reverse complement motif |
| Consensus sequence: GGCGGGGC | Consensus sequence: GCCCCGCC |
 |
 |
Best Matches for Motif ID 1 (Highest to Lowest)
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
THWYHHDCCCCGCCCHHHYADA
------GCCCCGCC--------
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.0340362 |
Alignment:
HHHDHHDTGCGGGGGHHAHHHY
-------GGCGGGGC-------
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 8 |
| Similarity score: | 0.0368821 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
----GCCCCGCC----------
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0478168 |
Alignment:
HHHHDMATCCCYCGCATKVBTV
--------GCCCCGCC------
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0498717 |
Alignment:
RHVMHHDGMGGGGGAGHWHTHTT
------GGCGGGGC---------
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Dataset #: 2 | Motif ID: 2 | Motif name: Motif 2 |
| Original motif | Reverse complement motif |
| Consensus sequence: RGRAGARRGARRAR | Consensus sequence: MTMMTCMMTCTKCK |
 |
 |
Best Matches for Motif ID 2 (Highest to Lowest)
| Motif ID: | UP00611 |
| Motif name: | PAX6 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0234155 |
Alignment:
DHBHGCVTGTGGAVVGCVCBBVH
---------MTMMTCMMTCTKCK
| Original motif | Reverse complement motif |
| Consensus sequence: HVBBGBGCVVTCCACAVGCHBHD | Consensus sequence: DHBHGCVTGTGGAVVGCVCBBVH |
 |
 |
| Motif ID: | UP00624 |
| Motif name: | VENTX_R143C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0262931 |
Alignment:
SHAHHHVATATMBTAYVRGBWWG
MTMMTCMMTCTKCK---------
| Original motif | Reverse complement motif |
| Consensus sequence: CWWVCMVMTABYATATVHDDTHS | Consensus sequence: SHAHHHVATATMBTAYVRGBWWG |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_H322Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0268132 |
Alignment:
DDDDDHDGGGTGGGHHHDDDVVK
-------RGRAGARRGARRAR--
| Original motif | Reverse complement motif |
| Consensus sequence: DDDDDHDGGGTGGGHHHDDDVVK | Consensus sequence: YVVDHDHHHCCCACCCHHDDDDD |
 |
 |
| Motif ID: | UP00594 |
| Motif name: | HESX1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.03053 |
Alignment:
DTHGBBBWHDVHBHDBVDVBHB
----MTMMTCMMTCTKCK----
| Original motif | Reverse complement motif |
| Consensus sequence: BHBVHVBDHVHVDHWVBVCHAD | Consensus sequence: DTHGBBBWHDVHBHDBVDVBHB |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_S265Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 14 |
| Similarity score: | 0.0308188 |
Alignment:
VBHDDHARDGGAGACMHDVDVVA
--------RGRAGARRGARRAR-
| Original motif | Reverse complement motif |
| Consensus sequence: VBHDDHARDGGAGACMHDVDVVA | Consensus sequence: TBVDVDDRGTCTCCHKTHDHHBV |
 |
 |
| Dataset #: 2 | Motif ID: 3 | Motif name: Motif 3 |
| Original motif | Reverse complement motif |
| Consensus sequence: AWAAAWTWAAASWA | Consensus sequence: TWSTTTWAWTTTWT |
 |
 |
Best Matches for Motif ID 3 (Highest to Lowest)
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.00761781 |
Alignment:
TWTTHVWATAHAWATAAWHTHKG
-------AWAAAWTWAAASWA--
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Motif ID: | UP00599 |
| Motif name: | KLF1_R328H_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 14 |
| Similarity score: | 0.00934745 |
Alignment:
THMTWDTAATADATTATBHTTM
TWSTTTWAWTTTWT--------
| Original motif | Reverse complement motif |
| Consensus sequence: THMTWDTAATADATTATBHTTM | Consensus sequence: RAAHBATAATDTATTADWAYHA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.00935632 |
Alignment:
TDTTHVWATAHADATAAWHTHKG
-------AWAAAWTWAAASWA--
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0117725 |
Alignment:
TWTTHVWATAVADATAAWHTHKG
-------AWAAAWTWAAASWA--
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Motif ID: | UP00606 |
| Motif name: | NR1H4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0125288 |
Alignment:
WWHVDAWDDATTWATAVTWMAAT
-----AWAAAWTWAAASWA----
| Original motif | Reverse complement motif |
| Consensus sequence: WWHVDAWDDATTWATAVTWMAAT | Consensus sequence: ATTYWABTATWAATDDWTDBHWW |
 |
 |
| Dataset #: 2 | Motif ID: 4 | Motif name: Motif 4 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAWTTRCWT | Consensus sequence: AWGKAAWTTTT |
 |
 |
Best Matches for Motif ID 4 (Highest to Lowest)
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 11 |
| Similarity score: | 0.0171027 |
Alignment:
HHAVVDCCMATAAAATDBDDVY
-----------AAAAWTTRCWT
| Original motif | Reverse complement motif |
| Consensus sequence: HHAVVDCCMATAAAATDBDDVY | Consensus sequence: KBDHVDATTTTATYGGDVVTHH |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.0174438 |
Alignment:
CRHAHWTTATWTHTATWBHAAWA
-AAAAWTTRCWT-----------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394W_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.0198047 |
Alignment:
TWTTHVWATAHAWATAAWHTHKG
-----------AWGKAAWTTTT-
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CYHAHWTTATWTDTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.0206321 |
Alignment:
TWTTHVWATAVADATAAWHTHKG
-----------AWGKAAWTTTT-
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Motif ID: | UP00523 |
| Motif name: | FOXR1_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.0206685 |
Alignment:
WBHGBRGCAAACTTHAKVHBHHD
--------AAAAWTTRCWT----
| Original motif | Reverse complement motif |
| Consensus sequence: WBHGBRGCAAACTTHAKVHBHHD | Consensus sequence: DHHBHVRTDAAGTTTGCKBCHVW |
 |
 |
| Dataset #: 2 | Motif ID: 5 | Motif name: Motif 5 |
| Original motif | Reverse complement motif |
| Consensus sequence: AWTAAATAYAATTT | Consensus sequence: AAATTKTATTTAWT |
 |
 |
Best Matches for Motif ID 5 (Highest to Lowest)
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0123886 |
Alignment:
TWTTHVWATAHAWATAAWHTHKG
------AWTAAATAYAATTT---
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.013075 |
Alignment:
TDTTHVWATAHADATAAWHTHKG
------AWTAAATAYAATTT---
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394W_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.013818 |
Alignment:
TWTTHVWATAHAWATAAWHTHKG
------AWTAAATAYAATTT---
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CYHAHWTTATWTDTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0165843 |
Alignment:
TWTTHVWATAVADATAAWHTHKG
------AWTAAATAYAATTT---
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Motif ID: | UP00619 |
| Motif name: | PROP1_R99Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0167999 |
Alignment:
AWGDWTAATYTAATTADTHDHV
----AAATTKTATTTAWT----
| Original motif | Reverse complement motif |
| Consensus sequence: AWGDWTAATYTAATTADTHDHV | Consensus sequence: VHDHADTAATTAMATTAWDCWT |
 |
 |
| Dataset #: 2 | Motif ID: 6 | Motif name: Motif 6 |
| Original motif | Reverse complement motif |
| Consensus sequence: AATTYDGAARTAWW | Consensus sequence: WWTAKTTCDKAATT |
 |
 |
Best Matches for Motif ID 6 (Highest to Lowest)
| Motif ID: | UP00624 |
| Motif name: | VENTX_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0255961 |
Alignment:
TWTWDTHGBTAATTABCDHHBDW
WWTAKTTCDKAATT---------
| Original motif | Reverse complement motif |
| Consensus sequence: WDVDHDGBTAATTABCHADWAWA | Consensus sequence: TWTWDTHGBTAATTABCDHHBDW |
 |
 |
| Motif ID: | UP00596 |
| Motif name: | HOXC4_R158L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 14 |
| Similarity score: | 0.0332804 |
Alignment:
HTWDTDHGHTAATTARCDDHHHW
WWTAKTTCDKAATT---------
| Original motif | Reverse complement motif |
| Consensus sequence: WHHHDDGKTAATTAHCHHADWAH | Consensus sequence: HTWDTDHGHTAATTARCDDHHHW |
 |
 |
| Motif ID: | UP00624 |
| Motif name: | VENTX_E101K_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 14 |
| Similarity score: | 0.0354322 |
Alignment:
WHWWDHYBCTAATTAVVBWHVHA
WWTAKTTCDKAATT---------
| Original motif | Reverse complement motif |
| Consensus sequence: TDVDWBVVTAATTAGBMHDWWHW | Consensus sequence: WHWWDHYBCTAATTAVVBWHVHA |
 |
 |
| Motif ID: | UP00625 |
| Motif name: | VSX1_Q175H_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 14 |
| Similarity score: | 0.0359429 |
Alignment:
HHWDTHRGVTAATTAGCCHRHAW
WWTAKTTCDKAATT---------
| Original motif | Reverse complement motif |
| Consensus sequence: WTHKHGGCTAATTAVCMHADWHH | Consensus sequence: HHWDTHRGVTAATTAGCCHRHAW |
 |
 |
| Motif ID: | UP00588 |
| Motif name: | ESX1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 14 |
| Similarity score: | 0.0398477 |
Alignment:
THDDTHDGCTAATTAKCVVRHDA
WWTAKTTCDKAATT---------
| Original motif | Reverse complement motif |
| Consensus sequence: THHKVBGYTAATTAGCHHADDHA | Consensus sequence: THDDTHDGCTAATTAKCVVRHDA |
 |
 |
| Dataset #: 2 | Motif ID: 7 | Motif name: Motif 7 |
| Original motif | Reverse complement motif |
| Consensus sequence: CSKCCCCGCCCCSY | Consensus sequence: MSGGGGCGGGGYSG |
 |
 |
Best Matches for Motif ID 7 (Highest to Lowest)
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0 |
Alignment:
THWYHHDCCCCGCCCHHHYADA
----CSKCCCCGCCCCSY----
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0205365 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
--CSKCCCCGCCCCSY------
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0376028 |
Alignment:
KHHHTHHCCCCCGCAHDHDHHH
-----CSKCCCCGCCCCSY---
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.043376 |
Alignment:
RHVMHHDGMGGGGGAGHWHTHTT
-----MSGGGGCGGGGYSG----
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_H322Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0476064 |
Alignment:
DDDDDHDGGGTGGGHHHDDDVVK
----MSGGGGCGGGGYSG-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDDDHDGGGTGGGHHHDDDVVK | Consensus sequence: YVVDHDHHHCCCACCCHHDDDDD |
 |
 |
| Dataset #: 2 | Motif ID: 8 | Motif name: Motif 8 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAATRWTAAAATCA | Consensus sequence: TGATTTTAWKATTT |
 |
 |
Best Matches for Motif ID 8 (Highest to Lowest)
| Motif ID: | UP00619 |
| Motif name: | PROP1_R99Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0194922 |
Alignment:
AWGDWTAATYTAATTADTHDHV
-----TGATTTTAWKATTT---
| Original motif | Reverse complement motif |
| Consensus sequence: AWGDWTAATYTAATTADTHDHV | Consensus sequence: VHDHADTAATTAMATTAWDCWT |
 |
 |
| Motif ID: | UP00619 |
| Motif name: | PROP1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0209962 |
Alignment:
HAHTBTAATYWAATTADBTDVG
-----TGATTTTAWKATTT---
| Original motif | Reverse complement motif |
| Consensus sequence: HAHTBTAATYWAATTADBTDVG | Consensus sequence: CBHAVDTAATTWMATTABADTH |
 |
 |
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0258194 |
Alignment:
DHVDHDTTTTACGAGVWHHDDG
---TGATTTTAWKATTT-----
| Original motif | Reverse complement motif |
| Consensus sequence: CDDHHWBCTCGTAAAADHDVHD | Consensus sequence: DHVDHDTTTTACGAGVWHHDDG |
 |
 |
| Motif ID: | UP00607 |
| Motif name: | NR2E3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0263894 |
Alignment:
DDADHHHTGATCTTTGBHVDVD
-------TGATTTTAWKATTT-
| Original motif | Reverse complement motif |
| Consensus sequence: DVDBHBCAAAGATCAHDHDTDD | Consensus sequence: DDADHHHTGATCTTTGBHVDVD |
 |
 |
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.031562 |
Alignment:
HHAVVDCCMATAAAATDBDDVY
----AAATRWTAAAATCA----
| Original motif | Reverse complement motif |
| Consensus sequence: HHAVVDCCMATAAAATDBDDVY | Consensus sequence: KBDHVDATTTTATYGGDVVTHH |
 |
 |
| Dataset #: 2 | Motif ID: 9 | Motif name: Motif 9 |
| Original motif | Reverse complement motif |
| Consensus sequence: AATHATATWTHAAA | Consensus sequence: TTTDAWATATHATT |
 |
 |
Best Matches for Motif ID 9 (Highest to Lowest)
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0084574 |
Alignment:
CYHAHWTTATDTDTATWBHAADA
--------AATHATATWTHAAA-
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 14 |
| Similarity score: | 0.0100981 |
Alignment:
CYHAHWTTATDTVTATWBHAAWA
--------AATHATATWTHAAA-
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394W_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0118845 |
Alignment:
CYHAHWTTATWTDTATWBHAAWA
--------AATHATATWTHAAA-
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CYHAHWTTATWTDTATWBHAAWA |
 |
 |
| Motif ID: | UP00606 |
| Motif name: | NR1H4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.012666 |
Alignment:
WWHVDAWDDATTWATAVTWMAAT
---------AATHATATWTHAAA
| Original motif | Reverse complement motif |
| Consensus sequence: WWHVDAWDDATTWATAVTWMAAT | Consensus sequence: ATTYWABTATWAATDDWTDBHWW |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0128993 |
Alignment:
CRHAHWTTATWTHTATWBHAAWA
--------AATHATATWTHAAA-
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Dataset #: 2 | Motif ID: 10 | Motif name: Motif 10 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTTCATAAWT | Consensus sequence: AWTTATGAAA |
 |
 |
Best Matches for Motif ID 10 (Highest to Lowest)
| Motif ID: | UP00619 |
| Motif name: | PROP1_R99Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 10 |
| Similarity score: | 0 |
Alignment:
VHDHADTAATTAMATTAWDCWT
----AWTTATGAAA--------
| Original motif | Reverse complement motif |
| Consensus sequence: AWGDWTAATYTAATTADTHDHV | Consensus sequence: VHDHADTAATTAMATTAWDCWT |
 |
 |
| Motif ID: | UP00399 |
| Motif name: | Oct-1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 10 |
| Similarity score: | 0.00166104 |
Alignment:
THHDVDWTWTGCAWAWBBVVDH
----AWTTATGAAA--------
| Original motif | Reverse complement motif |
| Consensus sequence: THHDVDWTWTGCAWAWBBVVDH | Consensus sequence: HDVBBVWTWTGCAWAWDBDHHA |
 |
 |
| Motif ID: | UP00617 |
| Motif name: | POU4F3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0024968 |
Alignment:
THVHVBTTATTAATAWHBWDHB
----AWTTATGAAA--------
| Original motif | Reverse complement motif |
| Consensus sequence: THVHVBTTATTAATAWHBWDHB | Consensus sequence: BHDWVHWTATTAATAAVBHVHA |
 |
 |
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.00671131 |
Alignment:
CDDHHWBCTCGTAAAADHDVHD
------TTTCATAAWT------
| Original motif | Reverse complement motif |
| Consensus sequence: CDDHHWBCTCGTAAAADHDVHD | Consensus sequence: DHVDHDTTTTACGAGVWHHDDG |
 |
 |
| Motif ID: | UP00588 |
| Motif name: | ESX1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.00948327 |
Alignment:
THDDTHDGCTAATTAKCVVRHDA
----TTTCATAAWT---------
| Original motif | Reverse complement motif |
| Consensus sequence: THHKVBGYTAATTAGCHHADDHA | Consensus sequence: THDDTHDGCTAATTAKCVVRHDA |
 |
 |
| Dataset #: 2 | Motif ID: 11 | Motif name: Motif 11 |
| Original motif | Reverse complement motif |
| Consensus sequence: ATTTWATGAAA | Consensus sequence: TTTCATWAAAT |
 |
 |
Best Matches for Motif ID 11 (Highest to Lowest)
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0 |
Alignment:
CDDHHWBCTCGTAAAADHDVHD
------TTTCATWAAAT-----
| Original motif | Reverse complement motif |
| Consensus sequence: CDDHHWBCTCGTAAAADHDVHD | Consensus sequence: DHVDHDTTTTACGAGVWHHDDG |
 |
 |
| Motif ID: | UP00619 |
| Motif name: | PROP1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0031995 |
Alignment:
HAHTBTAATYWAATTADBTDVG
---TTTCATWAAAT--------
| Original motif | Reverse complement motif |
| Consensus sequence: HAHTBTAATYWAATTADBTDVG | Consensus sequence: CBHAVDTAATTWMATTABADTH |
 |
 |
| Motif ID: | UP00619 |
| Motif name: | PROP1_R99Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.0047045 |
Alignment:
VHDHADTAATTAMATTAWDCWT
----TTTCATWAAAT-------
| Original motif | Reverse complement motif |
| Consensus sequence: AWGDWTAATYTAATTADTHDHV | Consensus sequence: VHDHADTAATTAMATTAWDCWT |
 |
 |
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.00591402 |
Alignment:
HHAVVDCCMATAAAATDBDDVY
-----TTTCATWAAAT------
| Original motif | Reverse complement motif |
| Consensus sequence: HHAVVDCCMATAAAATDBDDVY | Consensus sequence: KBDHVDATTTTATYGGDVVTHH |
 |
 |
| Motif ID: | UP00626 |
| Motif name: | VSX2_R200Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.00705472 |
Alignment:
THVMDHVTTAATTAABDRTHDD
------TTTCATWAAAT-----
| Original motif | Reverse complement motif |
| Consensus sequence: THVMDHVTTAATTAABDRTHDD | Consensus sequence: HDHAMDVTTAATTAABDDRBHA |
 |
 |
| Dataset #: 2 | Motif ID: 12 | Motif name: Motif 12 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAAACAAA | Consensus sequence: TTTGTTTT |
 |
 |
Best Matches for Motif ID 12 (Highest to Lowest)
| Motif ID: | UP00401 |
| Motif name: | Sox4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
HBVMCWTTGTTDDD
-----TTTGTTTT-
| Original motif | Reverse complement motif |
| Consensus sequence: DDDAACAAWGRVVH | Consensus sequence: HBVMCWTTGTTDDD |
 |
 |
| Motif ID: | UP00589 |
| Motif name: | FOXC1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 11 |
| Number of overlap: | 8 |
| Similarity score: | 0.00359317 |
Alignment:
BHVVDDWATRTAAACAAWSDBDD
----------AAAACAAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: BHVVDDWATRTAAACAAWSDBDD | Consensus sequence: DHBHSWTTGTTTAMATWDDVBDB |
 |
 |
| Motif ID: | UP00523 |
| Motif name: | FOXR1_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 8 |
| Similarity score: | 0.0103105 |
Alignment:
DVVDBKHTTGTTTGDBDBHVGVV
------TTTGTTTT---------
| Original motif | Reverse complement motif |
| Consensus sequence: VVCVHBDVDCAAACAAHYVDBVD | Consensus sequence: DVVDBKHTTGTTTGDBDBHVGVV |
 |
 |
| Motif ID: | UP00523 |
| Motif name: | FOXR1_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0310557 |
Alignment:
DHHBHVRTDAAGTTTGCKBCHVW
------------TTTGTTTT---
| Original motif | Reverse complement motif |
| Consensus sequence: WBHGBRGCAAACTTHAKVHBHHD | Consensus sequence: DHHBHVRTDAAGTTTGCKBCHVW |
 |
 |
| Motif ID: | UP00607 |
| Motif name: | NR2E3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 8 |
| Similarity score: | 0.0433693 |
Alignment:
DVDBHBCAAAGATCAHDHDTDD
---------AAAACAAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: DVDBHBCAAAGATCAHDHDTDD | Consensus sequence: DDADHHHTGATCTTTGBHVDVD |
 |
 |
| Dataset #: 2 | Motif ID: 13 | Motif name: Motif 13 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAAGATTT | Consensus sequence: AAATCTTT |
 |
 |
Best Matches for Motif ID 13 (Highest to Lowest)
| Motif ID: | UP00592 |
| Motif name: | GFI1B_A204T_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
TDBHVVGVMCAGATTTVMAHHAD
--------AAAGATTT-------
| Original motif | Reverse complement motif |
| Consensus sequence: DTDHTRBAAATCTGRVCBVHBDA | Consensus sequence: TDBHVVGVMCAGATTTVMAHHAD |
 |
 |
| Motif ID: | UP00607 |
| Motif name: | NR2E3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.0198394 |
Alignment:
DVDBHBCAAAGATCAHDHDTDD
-------AAAGATTT-------
| Original motif | Reverse complement motif |
| Consensus sequence: DVDBHBCAAAGATCAHDHDTDD | Consensus sequence: DDADHHHTGATCTTTGBHVDVD |
 |
 |
| Motif ID: | UP00523 |
| Motif name: | FOXR1_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0.033006 |
Alignment:
WBHGBRGCAAACTTHAKVHBHHD
-------AAATCTTT--------
| Original motif | Reverse complement motif |
| Consensus sequence: WBHGBRGCAAACTTHAKVHBHHD | Consensus sequence: DHHBHVRTDAAGTTTGCKBCHVW |
 |
 |
| Motif ID: | UP00592 |
| Motif name: | GFI1B_A204T_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0.0331884 |
Alignment:
HAHDDBGCCGTGATTTHDVHDDY
--------AAAGATTT-------
| Original motif | Reverse complement motif |
| Consensus sequence: KDDHVDDAAATCACGGCVDDHTH | Consensus sequence: HAHDDBGCCGTGATTTHDVHDDY |
 |
 |
| Motif ID: | UP00609 |
| Motif name: | PAX3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0367542 |
Alignment:
HBBHDVTAATCGATTADBHDDB
--------AAAGATTT------
| Original motif | Reverse complement motif |
| Consensus sequence: BDDHBDTAATCGATTAVHDBVD | Consensus sequence: HBBHDVTAATCGATTADBHDDB |
 |
 |
| Dataset #: 2 | Motif ID: 14 | Motif name: Motif 14 |
| Original motif | Reverse complement motif |
| Consensus sequence: AWAAATAA | Consensus sequence: TTATTTWT |
 |
 |
Best Matches for Motif ID 14 (Highest to Lowest)
| Motif ID: | UP00617 |
| Motif name: | POU4F3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
BHDWVHWTATTAATAAVBHVHA
--------AWAAATAA------
| Original motif | Reverse complement motif |
| Consensus sequence: THVHVBTTATTAATAWHBWDHB | Consensus sequence: BHDWVHWTATTAATAAVBHVHA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 8 |
| Similarity score: | 0.000449511 |
Alignment:
CYHAHWTTATDTDTATWBHAADA
------TTATTTWT---------
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.000763656 |
Alignment:
CRHAHWTTATWTHTATWBHAAWA
------TTATTTWT---------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Motif ID: | UP00589 |
| Motif name: | FOXC1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.00312951 |
Alignment:
BHVVDDWATRTAAACAAWSDBDD
---------AWAAATAA------
| Original motif | Reverse complement motif |
| Consensus sequence: BHVVDDWATRTAAACAAWSDBDD | Consensus sequence: DHBHSWTTGTTTAMATWDDVBDB |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 8 |
| Similarity score: | 0.0065948 |
Alignment:
TWTTHVWATAVADATAAWHTHKG
---------AWAAATAA------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Dataset #: 2 | Motif ID: 15 | Motif name: Motif 15 |
| Original motif | Reverse complement motif |
| Consensus sequence: ATMACAATAAAA | Consensus sequence: TTTTATTGTYAT |
 |
 |
Best Matches for Motif ID 15 (Highest to Lowest)
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 12 |
| Similarity score: | 0 |
Alignment:
KBDHVDATTTTATYGGDVVTHH
-------TTTTATTGTYAT---
| Original motif | Reverse complement motif |
| Consensus sequence: HHAVVDCCMATAAAATDBDDVY | Consensus sequence: KBDHVDATTTTATYGGDVVTHH |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 12 |
| Similarity score: | 0.00155138 |
Alignment:
CRHAHWTTATWTHTATWBHAAWA
----TTTTATTGTYAT-------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.00507302 |
Alignment:
TDTTHVWATAHADATAAWHTHKG
-------ATMACAATAAAA----
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.00621945 |
Alignment:
CYHAHWTTATDTVTATWBHAAWA
----TTTTATTGTYAT-------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394W_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.010042 |
Alignment:
CYHAHWTTATWTDTATWBHAAWA
----TTTTATTGTYAT-------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CYHAHWTTATWTDTATWBHAAWA |
 |
 |
| Dataset #: 2 | Motif ID: 16 | Motif name: Motif 16 |
| Original motif | Reverse complement motif |
| Consensus sequence: ACAAWTRATTTTGA | Consensus sequence: TCAAAATKAWTTGT |
 |
 |
Best Matches for Motif ID 16 (Highest to Lowest)
| Motif ID: | UP00523 |
| Motif name: | FOXR1_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0239714 |
Alignment:
DHHBHVRTDAAGTTTGCKBCHVW
---ACAAWTRATTTTGA------
| Original motif | Reverse complement motif |
| Consensus sequence: WBHGBRGCAAACTTHAKVHBHHD | Consensus sequence: DHHBHVRTDAAGTTTGCKBCHVW |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0257984 |
Alignment:
CRHAHWTTATWTHTATWBHAAWA
-ACAAWTRATTTTGA--------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Motif ID: | UP00619 |
| Motif name: | PROP1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0270326 |
Alignment:
HAHTBTAATYWAATTADBTDVG
---TCAAAATKAWTTGT-----
| Original motif | Reverse complement motif |
| Consensus sequence: HAHTBTAATYWAATTADBTDVG | Consensus sequence: CBHAVDTAATTWMATTABADTH |
 |
 |
| Motif ID: | UP00626 |
| Motif name: | VSX2_R200Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0282702 |
Alignment:
HDHAMDVTTAATTAABDDRBHA
---ACAAWTRATTTTGA-----
| Original motif | Reverse complement motif |
| Consensus sequence: THVMDHVTTAATTAABDRTHDD | Consensus sequence: HDHAMDVTTAATTAABDDRBHA |
 |
 |
| Motif ID: | UP00607 |
| Motif name: | NR2E3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0288801 |
Alignment:
DDADHHHTGATCTTTGBHVDVD
--ACAAWTRATTTTGA------
| Original motif | Reverse complement motif |
| Consensus sequence: DVDBHBCAAAGATCAHDHDTDD | Consensus sequence: DDADHHHTGATCTTTGBHVDVD |
 |
 |
| Dataset #: 2 | Motif ID: 17 | Motif name: Motif 17 |
| Original motif | Reverse complement motif |
| Consensus sequence: AAAAATGAAT | Consensus sequence: ATTCATTTTT |
 |
 |
Best Matches for Motif ID 17 (Highest to Lowest)
| Motif ID: | UP00607 |
| Motif name: | NR2E3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.00963039 |
Alignment:
DVDBHBCAAAGATCAHDHDTDD
-------AAAAATGAAT-----
| Original motif | Reverse complement motif |
| Consensus sequence: DVDBHBCAAAGATCAHDHDTDD | Consensus sequence: DDADHHHTGATCTTTGBHVDVD |
 |
 |
| Motif ID: | UP00617 |
| Motif name: | POU4F3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 10 |
| Similarity score: | 0.0124223 |
Alignment:
THVHVBTTATTAATAWHBWDHB
--------ATTCATTTTT----
| Original motif | Reverse complement motif |
| Consensus sequence: THVHVBTTATTAATAWHBWDHB | Consensus sequence: BHDWVHWTATTAATAAVBHVHA |
 |
 |
| Motif ID: | UP00626 |
| Motif name: | VSX2_R200Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 10 |
| Similarity score: | 0.0175439 |
Alignment:
HDHAMDVTTAATTAABDDRBHA
--AAAAATGAAT----------
| Original motif | Reverse complement motif |
| Consensus sequence: THVMDHVTTAATTAABDRTHDD | Consensus sequence: HDHAMDVTTAATTAABDDRBHA |
 |
 |
| Motif ID: | UP00603 |
| Motif name: | MSX2_R172H_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.0180636 |
Alignment:
HDHGCVDTAATTGGTVHMVHHT
-----ATTCATTTTT-------
| Original motif | Reverse complement motif |
| Consensus sequence: AHHVYDVACCAATTADVGCHDH | Consensus sequence: HDHGCVDTAATTGGTVHMVHHT |
 |
 |
| Motif ID: | UP00401 |
| Motif name: | Sox4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.0214899 |
Alignment:
DDDAACAAWGRVVH
---AAAAATGAAT-
| Original motif | Reverse complement motif |
| Consensus sequence: DDDAACAAWGRVVH | Consensus sequence: HBVMCWTTGTTDDD |
 |
 |
| Dataset #: 2 | Motif ID: 18 | Motif name: Motif 18 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTAGWTWATAA | Consensus sequence: TTATWAWCTAA |
 |
 |
Best Matches for Motif ID 18 (Highest to Lowest)
| Motif ID: | UP00626 |
| Motif name: | VSX2_R200Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0 |
Alignment:
THVMDHVTTAATTAABDRTHDD
-------TTAGWTWATAA----
| Original motif | Reverse complement motif |
| Consensus sequence: THVMDHVTTAATTAABDRTHDD | Consensus sequence: HDHAMDVTTAATTAABDDRBHA |
 |
 |
| Motif ID: | UP00604 |
| Motif name: | NKX2-5 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.00381186 |
Alignment:
TTTAGHTAATDTADATHGWTTA
-TTAGWTWATAA----------
| Original motif | Reverse complement motif |
| Consensus sequence: TAAWCHATDTADATTAHCTAAA | Consensus sequence: TTTAGHTAATDTADATHGWTTA |
 |
 |
| Motif ID: | UP00596 |
| Motif name: | HOXC4_R158L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.00443857 |
Alignment:
HTWDTDHGHTAATTARCDDHHHW
----TTAGWTWATAA--------
| Original motif | Reverse complement motif |
| Consensus sequence: WHHHDDGKTAATTAHCHHADWAH | Consensus sequence: HTWDTDHGHTAATTARCDDHHHW |
 |
 |
| Motif ID: | UP00620 |
| Motif name: | SIX6_H141N_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.00509375 |
Alignment:
YABTAHHTGATAMCMDTHHDAD
--TTAGWTWATAA---------
| Original motif | Reverse complement motif |
| Consensus sequence: DTDHHADRGRTATCAHHTAVTM | Consensus sequence: YABTAHHTGATAMCMDTHHDAD |
 |
 |
| Motif ID: | UP00619 |
| Motif name: | PROP1_R99Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.00541381 |
Alignment:
VHDHADTAATTAMATTAWDCWT
------TTATWAWCTAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: AWGDWTAATYTAATTADTHDHV | Consensus sequence: VHDHADTAATTAMATTAWDCWT |
 |
 |
| Dataset #: 2 | Motif ID: 19 | Motif name: Motif 19 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTCWTAGATTAWA | Consensus sequence: TWTAATCTAWGAA |
 |
 |
Best Matches for Motif ID 19 (Highest to Lowest)
| Motif ID: | UP00609 |
| Motif name: | PAX3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 13 |
| Similarity score: | 0.00606198 |
Alignment:
HBBHDVTAATCGATTADBHDDB
----TWTAATCTAWGAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDDHBDTAATCGATTAVHDBVD | Consensus sequence: HBBHDVTAATCGATTADBHDDB |
 |
 |
| Motif ID: | UP00619 |
| Motif name: | PROP1_R99Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 13 |
| Similarity score: | 0.00857347 |
Alignment:
AWGDWTAATYTAATTADTHDHV
---TWTAATCTAWGAA------
| Original motif | Reverse complement motif |
| Consensus sequence: AWGDWTAATYTAATTADTHDHV | Consensus sequence: VHDHADTAATTAMATTAWDCWT |
 |
 |
| Motif ID: | UP00599 |
| Motif name: | KLF1_R328H_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 13 |
| Similarity score: | 0.0131396 |
Alignment:
THMTWDTAATADATTATBHTTM
----TWTAATCTAWGAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: THMTWDTAATADATTATBHTTM | Consensus sequence: RAAHBATAATDTATTADWAYHA |
 |
 |
| Motif ID: | UP00607 |
| Motif name: | NR2E3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 13 |
| Similarity score: | 0.0148096 |
Alignment:
DVDBHBCAAAGATCAHDHDTDD
----TTCWTAGATTAWA-----
| Original motif | Reverse complement motif |
| Consensus sequence: DVDBHBCAAAGATCAHDHDTDD | Consensus sequence: DDADHHHTGATCTTTGBHVDVD |
 |
 |
| Motif ID: | UP00615 |
| Motif name: | PITX2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 13 |
| Similarity score: | 0.0156306 |
Alignment:
DDTBVRRKGGATTARADWDHAVH
---TTCWTAGATTAWA-------
| Original motif | Reverse complement motif |
| Consensus sequence: DDTBVRRKGGATTARADWDHAVH | Consensus sequence: HVTHHWDTKTAATCCYKMBVADD |
 |
 |
| Dataset #: 2 | Motif ID: 20 | Motif name: Motif 20 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTMAAAGATTT | Consensus sequence: AAATCTTTYAA |
 |
 |
Best Matches for Motif ID 20 (Highest to Lowest)
| Motif ID: | UP00592 |
| Motif name: | GFI1B_A204T_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.00709542 |
Alignment:
TDBHVVGVMCAGATTTVMAHHAD
-----TTMAAAGATTT-------
| Original motif | Reverse complement motif |
| Consensus sequence: DTDHTRBAAATCTGRVCBVHBDA | Consensus sequence: TDBHVVGVMCAGATTTVMAHHAD |
 |
 |
| Motif ID: | UP00607 |
| Motif name: | NR2E3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.00890281 |
Alignment:
DVDBHBCAAAGATCAHDHDTDD
----TTMAAAGATTT-------
| Original motif | Reverse complement motif |
| Consensus sequence: DVDBHBCAAAGATCAHDHDTDD | Consensus sequence: DDADHHHTGATCTTTGBHVDVD |
 |
 |
| Motif ID: | UP00609 |
| Motif name: | PAX3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.0138471 |
Alignment:
HBBHDVTAATCGATTADBHDDB
------AAATCTTTYAA-----
| Original motif | Reverse complement motif |
| Consensus sequence: BDDHBDTAATCGATTAVHDBVD | Consensus sequence: HBBHDVTAATCGATTADBHDDB |
 |
 |
| Motif ID: | UP00592 |
| Motif name: | GFI1B_A204T_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.0230866 |
Alignment:
HAHDDBGCCGTGATTTHDVHDDY
-----TTMAAAGATTT-------
| Original motif | Reverse complement motif |
| Consensus sequence: KDDHVDDAAATCACGGCVDDHTH | Consensus sequence: HAHDDBGCCGTGATTTHDVHDDY |
 |
 |
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.0281874 |
Alignment:
DHVDHDTTTTACGAGVWHHDDG
--------TTMAAAGATTT---
| Original motif | Reverse complement motif |
| Consensus sequence: CDDHHWBCTCGTAAAADHDVHD | Consensus sequence: DHVDHDTTTTACGAGVWHHDDG |
 |
 |
| Dataset #: 2 | Motif ID: 21 | Motif name: Motif 21 |
| Original motif | Reverse complement motif |
| Consensus sequence: ATAAAA | Consensus sequence: TTTTAT |
 |
 |
Best Matches for Motif ID 21 (Highest to Lowest)
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
KBDHVDATTTTATYGGDVVTHH
-------TTTTAT---------
| Original motif | Reverse complement motif |
| Consensus sequence: HHAVVDCCMATAAAATDBDDVY | Consensus sequence: KBDHVDATTTTATYGGDVVTHH |
 |
 |
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0.0141667 |
Alignment:
CDDHHWBCTCGTAAAADHDVHD
----------ATAAAA------
| Original motif | Reverse complement motif |
| Consensus sequence: CDDHHWBCTCGTAAAADHDVHD | Consensus sequence: DHVDHDTTTTACGAGVWHHDDG |
 |
 |
| Motif ID: | UP00589 |
| Motif name: | FOXC1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0387267 |
Alignment:
BHVVDDWATRTAAACAAWSDBDD
---------ATAAAA--------
| Original motif | Reverse complement motif |
| Consensus sequence: BHVVDDWATRTAAACAAWSDBDD | Consensus sequence: DHBHSWTTGTTTAMATWDDVBDB |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 14 |
| Number of overlap: | 6 |
| Similarity score: | 0.0420082 |
Alignment:
CYHAHWTTATDTDTATWBHAADA
----TTTTAT-------------
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 14 |
| Number of overlap: | 6 |
| Similarity score: | 0.0438866 |
Alignment:
CRHAHWTTATWTHTATWBHAAWA
----TTTTAT-------------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Dataset #: 3 | Motif ID: 22 | Motif name: Zfx |
| Original motif | Reverse complement motif |
| Consensus sequence: BBVGCCBVGGCCTV | Consensus sequence: VAGGCCBBGGCVBB |
 |
 |
Best Matches for Motif ID 22 (Highest to Lowest)
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0479447 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
------VAGGCCBBGGCVBB--
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0513 |
Alignment:
THWYHHDCCCCGCCCHHHYADA
---BBVGCCBVGGCCTV-----
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00521 |
| Motif name: | FOXN2_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0532525 |
Alignment:
DVKHRDHDWRGCGTCBHHBHSH
-BBVGCCBVGGCCTV-------
| Original motif | Reverse complement motif |
| Consensus sequence: DVKHRDHDWRGCGTCBHHBHSH | Consensus sequence: HSDBHHBGACGCKWDHHKHRVD |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0534632 |
Alignment:
HDGAVBVGACGCGTCYVBDDBC
-BBVGCCBVGGCCTV-------
| Original motif | Reverse complement motif |
| Consensus sequence: GVHDBBKGACGCGTCVBBTCHH | Consensus sequence: HDGAVBVGACGCGTCYVBDDBC |
 |
 |
| Motif ID: | UP00521 |
| Motif name: | FOXN2_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0583839 |
Alignment:
MWBDVHCGGCGTCVTBAGWSRRD
--BBVGCCBVGGCCTV-------
| Original motif | Reverse complement motif |
| Consensus sequence: MWBDVHCGGCGTCVTBAGWSRRD | Consensus sequence: DMMSWCTBABGACGCCGDBDBWY |
 |
 |
| Dataset #: 3 | Motif ID: 23 | Motif name: Egr1 |
| Original motif | Reverse complement motif |
| Consensus sequence: HGCGTGGGCGK | Consensus sequence: YCGCCCACGCH |
 |
 |
Best Matches for Motif ID 23 (Highest to Lowest)
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0 |
Alignment:
AAHAHWHCTCCCCCRCDHHYVHM
------YCGCCCACGCH------
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.00681456 |
Alignment:
VABVRATGCGKGGGATRDDHHH
------HGCGTGGGCGK-----
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0070197 |
Alignment:
KHHHTHHCCCCCGCAHDHDHHH
----YCGCCCACGCH-------
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 11 |
| Similarity score: | 0.0126311 |
Alignment:
TDTKHHHGGGCGGGGDHHMWHA
--HGCGTGGGCGK---------
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.0135196 |
Alignment:
DBBBSBDGGGGGCGYRDDHHHV
----HGCGTGGGCGK-------
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Dataset #: 3 | Motif ID: 24 | Motif name: SP1 |
| Original motif | Reverse complement motif |
| Consensus sequence: CCCCKCCCCC | Consensus sequence: GGGGGYGGGG |
 |
 |
Best Matches for Motif ID 24 (Highest to Lowest)
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0 |
Alignment:
THWYHHDCCCCGCCCHHHYADA
-------CCCCKCCCCC-----
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 10 |
| Similarity score: | 0.0153751 |
Alignment:
AAHAHWHCTCCCCCRCDHHYVHM
----------CCCCKCCCCC---
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_H322Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 10 |
| Similarity score: | 0.0162022 |
Alignment:
YVVDHDHHHCCCACCCHHDDDDD
--------CCCCKCCCCC-----
| Original motif | Reverse complement motif |
| Consensus sequence: DDDDDHDGGGTGGGHHHDDDVVK | Consensus sequence: YVVDHDHHHCCCACCCHHDDDDD |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 10 |
| Similarity score: | 0.0178026 |
Alignment:
DBBBSBDGGGGGCGYRDDHHHV
GGGGGYGGGG------------
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 10 |
| Similarity score: | 0.0274038 |
Alignment:
HHHDHHDTGCGGGGGHHAHHHY
----GGGGGYGGGG--------
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Dataset #: 3 | Motif ID: 25 | Motif name: TFAP2A |
| Original motif | Reverse complement motif |
| Consensus sequence: GCCBBVRGS | Consensus sequence: SCMVBBGGC |
 |
 |
Best Matches for Motif ID 25 (Highest to Lowest)
| Motif ID: | UP00521 |
| Motif name: | FOXN2_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 14 |
| Number of overlap: | 9 |
| Similarity score: | 0.0115283 |
Alignment:
MWBDVHCGGCGTCVTBAGWSRRD
-SCMVBBGGC-------------
| Original motif | Reverse complement motif |
| Consensus sequence: MWBDVHCGGCGTCVTBAGWSRRD | Consensus sequence: DMMSWCTBABGACGCCGDBDBWY |
 |
 |
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 9 |
| Similarity score: | 0.0158432 |
Alignment:
TDTKHHHGGGCGGGGDHHMWHA
--SCMVBBGGC-----------
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00523 |
| Motif name: | FOXR1_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 14 |
| Number of overlap: | 9 |
| Similarity score: | 0.0171317 |
Alignment:
VVCVHBDVDCAAACAAHYVDBVD
-SCMVBBGGC-------------
| Original motif | Reverse complement motif |
| Consensus sequence: VVCVHBDVDCAAACAAHYVDBVD | Consensus sequence: DVVDBKHTTGTTTGDBDBHVGVV |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 13 |
| Number of overlap: | 9 |
| Similarity score: | 0.0181013 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
------------GCCBBVRGS-
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 13 |
| Number of overlap: | 9 |
| Similarity score: | 0.0214094 |
Alignment:
HHHDHHDTGCGGGGGHHAHHHY
-SCMVBBGGC------------
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Dataset #: 3 | Motif ID: 26 | Motif name: MIZF |
| Original motif | Reverse complement motif |
| Consensus sequence: BAACGTCCGC | Consensus sequence: GCGGACGTTV |
 |
 |
Best Matches for Motif ID 26 (Highest to Lowest)
| Motif ID: | UP00522 |
| Motif name: | FOXN4_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.007999 |
Alignment:
SDYHHHMTAGCGTCBBMTHGADH
-------BAACGTCCGC------
| Original motif | Reverse complement motif |
| Consensus sequence: HDTCDARVBGACGCTAYHHHKHS | Consensus sequence: SDYHHHMTAGCGTCBBMTHGADH |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.0100757 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
-----BAACGTCCGC-------
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0152921 |
Alignment:
KHHHTHHCCCCCGCAHDHDHHH
----BAACGTCCGC--------
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00523 |
| Motif name: | FOXR1_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.0161684 |
Alignment:
DHHBHVRTDAAGTTTGCKBCHVW
-------BAACGTCCGC------
| Original motif | Reverse complement motif |
| Consensus sequence: WBHGBRGCAAACTTHAKVHBHHD | Consensus sequence: DHHBHVRTDAAGTTTGCKBCHVW |
 |
 |
| Motif ID: | UP00521 |
| Motif name: | FOXN2_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.0163494 |
Alignment:
DVKHRDHDWRGCGTCBHHBHSH
--------BAACGTCCGC----
| Original motif | Reverse complement motif |
| Consensus sequence: DVKHRDHDWRGCGTCBHHBHSH | Consensus sequence: HSDBHHBGACGCKWDHHKHRVD |
 |
 |
| Dataset #: 3 | Motif ID: 27 | Motif name: Klf4 |
| Original motif | Reverse complement motif |
| Consensus sequence: DGGGYGKGGC | Consensus sequence: GCCYCMCCCD |
 |
 |
Best Matches for Motif ID 27 (Highest to Lowest)
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0 |
Alignment:
TDTKHHHGGGCGGGGDHHMWHA
------DGGGYGKGGC------
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 10 |
| Similarity score: | 0.0140563 |
Alignment:
AAHAHWHCTCCCCCRCDHHYVHM
-----GCCYCMCCCD--------
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 10 |
| Similarity score: | 0.0236259 |
Alignment:
DBBBSBDGGGGGCGYRDDHHHV
--------DGGGYGKGGC----
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_H322Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.0319872 |
Alignment:
DDDDDHDGGGTGGGHHHDDDVVK
------DGGGYGKGGC-------
| Original motif | Reverse complement motif |
| Consensus sequence: DDDDDHDGGGTGGGHHHDDDVVK | Consensus sequence: YVVDHDHHHCCCACCCHHDDDDD |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 10 |
| Similarity score: | 0.0363573 |
Alignment:
VABVRATGCGKGGGATRDDHHH
----DGGGYGKGGC--------
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Dataset #: 3 | Motif ID: 28 | Motif name: E2F1 |
| Original motif | Reverse complement motif |
| Consensus sequence: TTTSGCGC | Consensus sequence: GCGCSAAA |
 |
 |
Best Matches for Motif ID 28 (Highest to Lowest)
| Motif ID: | UP00523 |
| Motif name: | FOXR1_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
WBHGBRGCAAACTTHAKVHBHHD
---GCGCSAAA------------
| Original motif | Reverse complement motif |
| Consensus sequence: WBHGBRGCAAACTTHAKVHBHHD | Consensus sequence: DHHBHVRTDAAGTTTGCKBCHVW |
 |
 |
| Motif ID: | UP00523 |
| Motif name: | FOXR1_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.00633054 |
Alignment:
DVVDBKHTTGTTTGDBDBHVGVV
----------TTTSGCGC-----
| Original motif | Reverse complement motif |
| Consensus sequence: VVCVHBDVDCAAACAAHYVDBVD | Consensus sequence: DVVDBKHTTGTTTGDBDBHVGVV |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 8 |
| Similarity score: | 0.0144303 |
Alignment:
SDYHHHMTAGCGTCBBMTHGADH
-----TTTSGCGC----------
| Original motif | Reverse complement motif |
| Consensus sequence: HDTCDARVBGACGCTAYHHHKHS | Consensus sequence: SDYHHHMTAGCGTCBBMTHGADH |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 8 |
| Similarity score: | 0.0178986 |
Alignment:
GVHDBBKGACGCGTCVBBTCHH
----TTTSGCGC----------
| Original motif | Reverse complement motif |
| Consensus sequence: GVHDBBKGACGCGTCVBBTCHH | Consensus sequence: HDGAVBVGACGCGTCYVBDDBC |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 8 |
| Similarity score: | 0.0184566 |
Alignment:
DBBBSBDGGGGGCGYRDDHHHV
-----------GCGCSAAA---
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Dataset #: 3 | Motif ID: 29 | Motif name: HIF1AARNT |
| Original motif | Reverse complement motif |
| Consensus sequence: VBACGTGV | Consensus sequence: VCACGTBV |
 |
 |
Best Matches for Motif ID 29 (Highest to Lowest)
| Motif ID: | UP00621 |
| Motif name: | SNAI2_T234I_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.00630757 |
Alignment:
HHDHDADRCACCTGYTADHHAD
-------VBACGTGV-------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDHDADRCACCTGYTADHHAD | Consensus sequence: DTDHDTAKCAGGTGMDTHHDHH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.00804833 |
Alignment:
HHDVDADRCACCTGYTADHHHD
-------VBACGTGV-------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVDADRCACCTGYTADHHHD | Consensus sequence: DHDHDTAKCAGGTGMDTHBDHH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_D119E_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 8 |
| Similarity score: | 0.0083564 |
Alignment:
HHHHDTAKCAGGTGMDTHBDDH
-------VCACGTBV-------
| Original motif | Reverse complement motif |
| Consensus sequence: HDDVDADRCACCTGYTADHHHH | Consensus sequence: HHHHDTAKCAGGTGMDTHBDDH |
 |
 |
| Motif ID: | UP00612 |
| Motif name: | PAX7 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0.0210813 |
Alignment:
VTHDBKSGTCACGGTTHDHDBV
--------VCACGTBV------
| Original motif | Reverse complement motif |
| Consensus sequence: VVDHDHAACCGTGACSRVDHAV | Consensus sequence: VTHDBKSGTCACGGTTHDHDBV |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0215974 |
Alignment:
HDGAVBVGACGCGTCYVBDDBC
--------VCACGTBV------
| Original motif | Reverse complement motif |
| Consensus sequence: GVHDBBKGACGCGTCVBBTCHH | Consensus sequence: HDGAVBVGACGCGTCYVBDDBC |
 |
 |
| Dataset #: 3 | Motif ID: 30 | Motif name: PLAG1 |
| Original motif | Reverse complement motif |
| Consensus sequence: GGGGCCCAAGGGGG | Consensus sequence: CCCCCTTGGGCCCC |
 |
 |
Best Matches for Motif ID 30 (Highest to Lowest)
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0264444 |
Alignment:
RHVMHHDGMGGGGGAGHWHTHTT
GGGGCCCAAGGGGG---------
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_S265Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 14 |
| Similarity score: | 0.0354436 |
Alignment:
VBHDDHARDGGAGACMHDVDVVA
---------GGGGCCCAAGGGGG
| Original motif | Reverse complement motif |
| Consensus sequence: VBHDDHARDGGAGACMHDVDVVA | Consensus sequence: TBVDVDDRGTCTCCHKTHDHHBV |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 14 |
| Similarity score: | 0.0356408 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
-CCCCCTTGGGCCCC-------
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 14 |
| Similarity score: | 0.0363329 |
Alignment:
VBHDDDAADGGAGACMHHVDVDW
---------GGGGCCCAAGGGGG
| Original motif | Reverse complement motif |
| Consensus sequence: VBHDDDAADGGAGACMHHVDVDW | Consensus sequence: WDVDVHDRGTCTCCHTTDDHHBV |
 |
 |
| Motif ID: | UP00521 |
| Motif name: | FOXN2_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0372132 |
Alignment:
MWBDVHCGGCGTCVTBAGWSRRD
-------GGGGCCCAAGGGGG--
| Original motif | Reverse complement motif |
| Consensus sequence: MWBDVHCGGCGTCVTBAGWSRRD | Consensus sequence: DMMSWCTBABGACGCCGDBDBWY |
 |
 |
| Dataset #: 3 | Motif ID: 31 | Motif name: Pax5 |
| Original motif | Reverse complement motif |
| Consensus sequence: DGVBCABTGDWGCGKRRCSR | Consensus sequence: MSGKKRCGCWDCABTGBBCD |
 |
 |
Best Matches for Motif ID 31 (Highest to Lowest)
| Motif ID: | UP00629 |
| Motif name: | ZNF200_H322Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 20 |
| Similarity score: | 0.0431517 |
Alignment:
DDDDDHDGGGTGGGHHHDDDVVK
DGVBCABTGDWGCGKRRCSR---
| Original motif | Reverse complement motif |
| Consensus sequence: DDDDDHDGGGTGGGHHHDDDVVK | Consensus sequence: YVVDHDHHHCCCACCCHHDDDDD |
 |
 |
| Motif ID: | UP00594 |
| Motif name: | HESX1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0480075 |
Alignment:
DTHGBBBWHDVHBHDBVDVBHB
--DGVBCABTGDWGCGKRRCSR
| Original motif | Reverse complement motif |
| Consensus sequence: BHBVHVBDHVHVDHWVBVCHAD | Consensus sequence: DTHGBBBWHDVHBHDBVDVBHB |
 |
 |
| Motif ID: | UP00611 |
| Motif name: | PAX6 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0504226 |
Alignment:
HVBBGBGCVVTCCACAVGCHBHD
-MSGKKRCGCWDCABTGBBCD--
| Original motif | Reverse complement motif |
| Consensus sequence: HVBBGBGCVVTCCACAVGCHBHD | Consensus sequence: DHBHGCVTGTGGAVVGCVCBBVH |
 |
 |
| Motif ID: | UP00599 |
| Motif name: | KLF1_R328H_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 20 |
| Similarity score: | 0.050775 |
Alignment:
RDHTDDBTGTDATGTGHVDWHBW
DGVBCABTGDWGCGKRRCSR---
| Original motif | Reverse complement motif |
| Consensus sequence: WBHWDVHCACATDACABDDADHK | Consensus sequence: RDHTDDBTGTDATGTGHVDWHBW |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0513444 |
Alignment:
DBBBSBDGGGGGCGYRDDHHHV
DGVBCABTGDWGCGKRRCSR--
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Dataset #: 3 | Motif ID: 32 | Motif name: ArntAhr |
| Original motif | Reverse complement motif |
| Consensus sequence: YGCGTG | Consensus sequence: CACGCM |
 |
 |
Best Matches for Motif ID 32 (Highest to Lowest)
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
VABVRATGCGKGGGATRDDHHH
------YGCGTG----------
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0.00272949 |
Alignment:
KHHHTHHCCCCCGCAHDHDHHH
---------CACGCM-------
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00521 |
| Motif name: | FOXN2_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 11 |
| Number of overlap: | 6 |
| Similarity score: | 0.00939422 |
Alignment:
DMMSWCTBABGACGCCGDBDBWY
----------CACGCM-------
| Original motif | Reverse complement motif |
| Consensus sequence: MWBDVHCGGCGTCVTBAGWSRRD | Consensus sequence: DMMSWCTBABGACGCCGDBDBWY |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.0119829 |
Alignment:
SDYHHHMTAGCGTCBBMTHGADH
--------YGCGTG---------
| Original motif | Reverse complement motif |
| Consensus sequence: HDTCDARVBGACGCTAYHHHKHS | Consensus sequence: SDYHHHMTAGCGTCBBMTHGADH |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 6 |
| Similarity score: | 0.0121968 |
Alignment:
AAHAHWHCTCCCCCRCDHHYVHM
-----------CACGCM------
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Dataset #: 3 | Motif ID: 33 | Motif name: Mycn |
| Original motif | Reverse complement motif |
| Consensus sequence: HSCACGTGGC | Consensus sequence: GCCACGTGSD |
 |
 |
Best Matches for Motif ID 33 (Highest to Lowest)
| Motif ID: | UP00605 |
| Motif name: | NKX2-8 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0 |
Alignment:
SBVMAWVTCMAGTGGDBBBDHH
------GCCACGTGSD------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVVVDCCACTYGAVWTYVBS | Consensus sequence: SBVMAWVTCMAGTGGDBBBDHH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_D119E_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.00142642 |
Alignment:
HHHHDTAKCAGGTGMDTHBDDH
------GCCACGTGSD------
| Original motif | Reverse complement motif |
| Consensus sequence: HDDVDADRCACCTGYTADHHHH | Consensus sequence: HHHHDTAKCAGGTGMDTHBDDH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_T234I_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.00196758 |
Alignment:
DTDHDTAKCAGGTGMDTHHDHH
------GCCACGTGSD------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDHDADRCACCTGYTADHHAD | Consensus sequence: DTDHDTAKCAGGTGMDTHHDHH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.00337084 |
Alignment:
DHDHDTAKCAGGTGMDTHBDHH
------GCCACGTGSD------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVDADRCACCTGYTADHHHD | Consensus sequence: DHDHDTAKCAGGTGMDTHBDHH |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.00795147 |
Alignment:
HDGAVBVGACGCGTCYVBDDBC
-----HSCACGTGGC-------
| Original motif | Reverse complement motif |
| Consensus sequence: GVHDBBKGACGCGTCVBBTCHH | Consensus sequence: HDGAVBVGACGCGTCYVBDDBC |
 |
 |
| Dataset #: 3 | Motif ID: 34 | Motif name: Myc |
| Original motif | Reverse complement motif |
| Consensus sequence: VGCACGTGGH | Consensus sequence: DCCACGTGCV |
 |
 |
Best Matches for Motif ID 34 (Highest to Lowest)
| Motif ID: | UP00621 |
| Motif name: | SNAI2_D119E_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0 |
Alignment:
HHHHDTAKCAGGTGMDTHBDDH
------DCCACGTGCV------
| Original motif | Reverse complement motif |
| Consensus sequence: HDDVDADRCACCTGYTADHHHH | Consensus sequence: HHHHDTAKCAGGTGMDTHBDDH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.00177738 |
Alignment:
DHDHDTAKCAGGTGMDTHBDHH
------DCCACGTGCV------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVDADRCACCTGYTADHHHD | Consensus sequence: DHDHDTAKCAGGTGMDTHBDHH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_T234I_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.00250317 |
Alignment:
DTDHDTAKCAGGTGMDTHHDHH
------DCCACGTGCV------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDHDADRCACCTGYTADHHAD | Consensus sequence: DTDHDTAKCAGGTGMDTHHDHH |
 |
 |
| Motif ID: | UP00605 |
| Motif name: | NKX2-8 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 10 |
| Similarity score: | 0.00749721 |
Alignment:
SBVMAWVTCMAGTGGDBBBDHH
------DCCACGTGCV------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVVVDCCACTYGAVWTYVBS | Consensus sequence: SBVMAWVTCMAGTGGDBBBDHH |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.0117827 |
Alignment:
HDGAVBVGACGCGTCYVBDDBC
-----VGCACGTGGH-------
| Original motif | Reverse complement motif |
| Consensus sequence: GVHDBBKGACGCGTCVBBTCHH | Consensus sequence: HDGAVBVGACGCGTCYVBDDBC |
 |
 |
| Dataset #: 3 | Motif ID: 35 | Motif name: GABPA |
| Original motif | Reverse complement motif |
| Consensus sequence: CCGGAAGTGVV | Consensus sequence: VVCACTTCCGG |
 |
 |
Best Matches for Motif ID 35 (Highest to Lowest)
| Motif ID: | UP00586 |
| Motif name: | CRX |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0482888 |
Alignment:
TVDHDVCTAATCCGVTTHDVADD
----VVCACTTCCGG--------
| Original motif | Reverse complement motif |
| Consensus sequence: DDTVDHAABCGGATTAGBDHDVA | Consensus sequence: TVDHDVCTAATCCGVTTHDVADD |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 11 |
| Similarity score: | 0.0483091 |
Alignment:
HHHDHHDTGCGGGGGHHAHHHY
-VVCACTTCCGG----------
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00608 |
| Motif name: | OVOL2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.0542546 |
Alignment:
DWHDTVYADMTAACGGWYDHAYH
-----VVCACTTCCGG-------
| Original motif | Reverse complement motif |
| Consensus sequence: DWHDTVYADMTAACGGWYDHAYH | Consensus sequence: DMTHDMWCCGTTAYDTKVADHWD |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 12 |
| Number of overlap: | 11 |
| Similarity score: | 0.0567917 |
Alignment:
HHHHDMATCCCYCGCATKVBTV
-----------CCGGAAGTGVV
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Motif ID: | UP00612 |
| Motif name: | PAX7 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.0590324 |
Alignment:
VVDHDHAACCGTGACSRVDHAV
--------CCGGAAGTGVV---
| Original motif | Reverse complement motif |
| Consensus sequence: VVDHDHAACCGTGACSRVDHAV | Consensus sequence: VTHDBKSGTCACGGTTHDHDBV |
 |
 |
| Dataset #: 4 | Motif ID: 36 | Motif name: csGCCCCGCCCCsc |
| Original motif | Reverse complement motif |
| Consensus sequence: HVGCCCCGCCCCBB | Consensus sequence: BBGGGGCGGGGCVD |
 |
 |
Best Matches for Motif ID 36 (Highest to Lowest)
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0 |
Alignment:
THWYHHDCCCCGCCCHHHYADA
----HVGCCCCGCCCCBB----
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0197359 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
--HVGCCCCGCCCCBB------
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 14 |
| Similarity score: | 0.0359414 |
Alignment:
KHHHTHHCCCCCGCAHDHDHHH
-----HVGCCCCGCCCCBB---
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.045601 |
Alignment:
VABVRATGCGKGGGATRDDHHH
--BBGGGGCGGGGCVD------
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.0459004 |
Alignment:
AAHAHWHCTCCCCCRCDHHYVHM
-------HVGCCCCGCCCCBB--
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Dataset #: 4 | Motif ID: 37 | Motif name: tkAAATAATAtw |
| Original motif | Reverse complement motif |
| Consensus sequence: HDAAATAATAHW | Consensus sequence: WHTATTATTTDH |
 |
 |
Best Matches for Motif ID 37 (Highest to Lowest)
| Motif ID: | UP00617 |
| Motif name: | POU4F3 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 12 |
| Similarity score: | 0.0220779 |
Alignment:
BHDWVHWTATTAATAAVBHVHA
-----WHTATTATTTDH-----
| Original motif | Reverse complement motif |
| Consensus sequence: THVHVBTTATTAATAWHBWDHB | Consensus sequence: BHDWVHWTATTAATAAVBHVHA |
 |
 |
| Motif ID: | UP00619 |
| Motif name: | PROP1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 12 |
| Similarity score: | 0.0257795 |
Alignment:
HAHTBTAATYWAATTADBTDVG
---------WHTATTATTTDH-
| Original motif | Reverse complement motif |
| Consensus sequence: HAHTBTAATYWAATTADBTDVG | Consensus sequence: CBHAVDTAATTWMATTABADTH |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 12 |
| Similarity score: | 0.0258362 |
Alignment:
CYHAHWTTATDTDTATWBHAADA
--WHTATTATTTDH---------
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00606 |
| Motif name: | NR1H4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 12 |
| Similarity score: | 0.0279126 |
Alignment:
ATTYWABTATWAATDDWTDBHWW
-----WHTATTATTTDH------
| Original motif | Reverse complement motif |
| Consensus sequence: WWHVDAWDDATTWATAVTWMAAT | Consensus sequence: ATTYWABTATWAATDDWTDBHWW |
 |
 |
| Motif ID: | UP00599 |
| Motif name: | KLF1_R328H_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 12 |
| Similarity score: | 0.0289279 |
Alignment:
THMTWDTAATADATTATBHTTM
---------WHTATTATTTDH-
| Original motif | Reverse complement motif |
| Consensus sequence: THMTWDTAATADATTATBHTTM | Consensus sequence: RAAHBATAATDTATTADWAYHA |
 |
 |
| Dataset #: 4 | Motif ID: 38 | Motif name: cccGCCCCGCCCCsb |
| Original motif | Reverse complement motif |
| Consensus sequence: BCCGCCCCGCCCCBB | Consensus sequence: BBGGGGCGGGGCGGB |
 |
 |
Best Matches for Motif ID 38 (Highest to Lowest)
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 15 |
| Similarity score: | 0 |
Alignment:
THWYHHDCCCCGCCCHHHYADA
---BCCGCCCCGCCCCBB----
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 15 |
| Similarity score: | 0.0183446 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
-BCCGCCCCGCCCCBB------
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 15 |
| Similarity score: | 0.0349416 |
Alignment:
KHHHTHHCCCCCGCAHDHDHHH
----BCCGCCCCGCCCCBB---
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 15 |
| Similarity score: | 0.0385263 |
Alignment:
AAHAHWHCTCCCCCRCDHHYVHM
------BCCGCCCCGCCCCBB--
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 15 |
| Similarity score: | 0.0405718 |
Alignment:
VABVRATGCGKGGGATRDDHHH
--BBGGGGCGGGGCGGB-----
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Dataset #: 4 | Motif ID: 39 | Motif name: kCAGCCAATmr |
| Original motif | Reverse complement motif |
| Consensus sequence: DCAGCCAATVR | Consensus sequence: MBATTGGCTGH |
 |
 |
Best Matches for Motif ID 39 (Highest to Lowest)
| Motif ID: | UP00584 |
| Motif name: | ARX_T333N_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.00749807 |
Alignment:
VABDDBVCYAATTRGCBVDTDH
--------MBATTGGCTGH---
| Original motif | Reverse complement motif |
| Consensus sequence: VABDDBVCYAATTRGCBVDTDH | Consensus sequence: HHADVVGCKAATTMGVBDHVTB |
 |
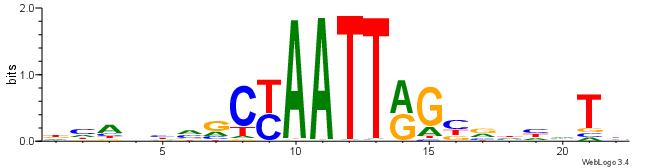 |
| Motif ID: | UP00605 |
| Motif name: | NKX2-8 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0182048 |
Alignment:
HHDVVVDCCACTYGAVWTYVBS
---DCAGCCAATVR--------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVVVDCCACTYGAVWTYVBS | Consensus sequence: SBVMAWVTCMAGTGGDBBBDHH |
 |
 |
| Motif ID: | UP00523 |
| Motif name: | FOXR1_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.0193622 |
Alignment:
VVCVHBDVDCAAACAAHYVDBVD
--------DCAGCCAATVR----
| Original motif | Reverse complement motif |
| Consensus sequence: VVCVHBDVDCAAACAAHYVDBVD | Consensus sequence: DVVDBKHTTGTTTGDBDBHVGVV |
 |
 |
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 11 |
| Similarity score: | 0.0203222 |
Alignment:
HHAVVDCCMATAAAATDBDDVY
--DCAGCCAATVR---------
| Original motif | Reverse complement motif |
| Consensus sequence: HHAVVDCCMATAAAATDBDDVY | Consensus sequence: KBDHVDATTTTATYGGDVVTHH |
 |
 |
| Motif ID: | UP00625 |
| Motif name: | VSX1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.0208017 |
Alignment:
DAGRBBGCTAATTAGBVVBYBH
---DCAGCCAATVR--------
| Original motif | Reverse complement motif |
| Consensus sequence: DAGRBBGCTAATTAGBVVBYBH | Consensus sequence: HBKBBVBCTAATTAGCVVKCTD |
 |
 |
| Dataset #: 4 | Motif ID: 40 | Motif name: kcACCTGCAgc |
| Original motif | Reverse complement motif |
| Consensus sequence: BCACCTGCABC | Consensus sequence: GBTGCAGGTGB |
 |
 |
Best Matches for Motif ID 40 (Highest to Lowest)
| Motif ID: | UP00621 |
| Motif name: | SNAI2_T234I_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0 |
Alignment:
DTDHDTAKCAGGTGMDTHHDHH
----GBTGCAGGTGB-------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDHDADRCACCTGYTADHHAD | Consensus sequence: DTDHDTAKCAGGTGMDTHHDHH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_D119E_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.000881825 |
Alignment:
HHHHDTAKCAGGTGMDTHBDDH
----GBTGCAGGTGB-------
| Original motif | Reverse complement motif |
| Consensus sequence: HDDVDADRCACCTGYTADHHHH | Consensus sequence: HHHHDTAKCAGGTGMDTHBDDH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.0013625 |
Alignment:
DHDHDTAKCAGGTGMDTHBDHH
----GBTGCAGGTGB-------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVDADRCACCTGYTADHHHD | Consensus sequence: DHDHDTAKCAGGTGMDTHBDHH |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0178142 |
Alignment:
KHHHTHHCCCCCGCAHDHDHHH
------BCACCTGCABC-----
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0381181 |
Alignment:
VABVRATGCGKGGGATRDDHHH
----GBTGCAGGTGB-------
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Dataset #: 4 | Motif ID: 41 | Motif name: wwAAATAATAtw |
| Original motif | Reverse complement motif |
| Consensus sequence: HDAAATAATADD | Consensus sequence: DDTATTATTTDH |
 |
 |
Best Matches for Motif ID 41 (Highest to Lowest)
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 12 |
| Similarity score: | 0.0155891 |
Alignment:
CYHAHWTTATDTDTATWBHAADA
--DDTATTATTTDH---------
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 12 |
| Similarity score: | 0.0172798 |
Alignment:
TWTTHVWATAHAWATAAWHTHKG
---------HDAAATAATADD--
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 12 |
| Similarity score: | 0.0186508 |
Alignment:
CYHAHWTTATDTVTATWBHAAWA
--DDTATTATTTDH---------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394W_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 12 |
| Similarity score: | 0.0190508 |
Alignment:
CYHAHWTTATWTDTATWBHAAWA
--DDTATTATTTDH---------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CYHAHWTTATWTDTATWBHAAWA |
 |
 |
| Motif ID: | UP00599 |
| Motif name: | KLF1_R328H_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 12 |
| Similarity score: | 0.0220346 |
Alignment:
THMTWDTAATADATTATBHTTM
---------DDTATTATTTDH-
| Original motif | Reverse complement motif |
| Consensus sequence: THMTWDTAATADATTATBHTTM | Consensus sequence: RAAHBATAATDTATTADWAYHA |
 |
 |
| Dataset #: 4 | Motif ID: 42 | Motif name: sSGTCACGTGACSs |
| Original motif | Reverse complement motif |
| Consensus sequence: SGGTCACGTGACCS | Consensus sequence: SGGTCACGTGACCS |
 |
 |
Best Matches for Motif ID 42 (Highest to Lowest)
| Motif ID: | UP00612 |
| Motif name: | PAX7 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.00859567 |
Alignment:
VTHDBKSGTCACGGTTHDHDBV
-----SGGTCACGTGACCS---
| Original motif | Reverse complement motif |
| Consensus sequence: VVDHDHAACCGTGACSRVDHAV | Consensus sequence: VTHDBKSGTCACGGTTHDHDBV |
 |
 |
| Motif ID: | UP00605 |
| Motif name: | NKX2-8 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0145827 |
Alignment:
HHDVVVDCCACTYGAVWTYVBS
----SGGTCACGTGACCS----
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVVVDCCACTYGAVWTYVBS | Consensus sequence: SBVMAWVTCMAGTGGDBBBDHH |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0151743 |
Alignment:
DHDHDTAKCAGGTGMDTHBDHH
----SGGTCACGTGACCS----
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVDADRCACCTGYTADHHHD | Consensus sequence: DHDHDTAKCAGGTGMDTHBDHH |
 |
 |
| Motif ID: | UP00599 |
| Motif name: | KLF1_R328H_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 14 |
| Similarity score: | 0.0168994 |
Alignment:
WBHWDVHCACATDACABDDADHK
---SGGTCACGTGACCS------
| Original motif | Reverse complement motif |
| Consensus sequence: WBHWDVHCACATDACABDDADHK | Consensus sequence: RDHTDDBTGTDATGTGHVDWHBW |
 |
 |
| Motif ID: | UP00621 |
| Motif name: | SNAI2_D119E_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0.0169037 |
Alignment:
HHHHDTAKCAGGTGMDTHBDDH
----SGGTCACGTGACCS----
| Original motif | Reverse complement motif |
| Consensus sequence: HDDVDADRCACCTGYTADHHHH | Consensus sequence: HHHHDTAKCAGGTGMDTHBDDH |
 |
 |
| Dataset #: 4 | Motif ID: 43 | Motif name: wsTACwGTAsw |
| Original motif | Reverse complement motif |
| Consensus sequence: HVTACWGTABD | Consensus sequence: DBTACWGTAVH |
 |
 |
Best Matches for Motif ID 43 (Highest to Lowest)
| Motif ID: | UP00605 |
| Motif name: | NKX2-8 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0140465 |
Alignment:
SBVMAWVTCMAGTGGDBBBDHH
-----HVTACWGTABD------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDVVVDCCACTYGAVWTYVBS | Consensus sequence: SBVMAWVTCMAGTGGDBBBDHH |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.0186228 |
Alignment:
SDYHHHMTAGCGTCBBMTHGADH
-----HVTACWGTABD-------
| Original motif | Reverse complement motif |
| Consensus sequence: HDTCDARVBGACGCTAYHHHKHS | Consensus sequence: SDYHHHMTAGCGTCBBMTHGADH |
 |
 |
| Motif ID: | UP00608 |
| Motif name: | OVOL2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.0245566 |
Alignment:
DMTHDMWCCGTTAYDTKVADHWD
----HVTACWGTABD--------
| Original motif | Reverse complement motif |
| Consensus sequence: DWHDTVYADMTAACGGWYDHAYH | Consensus sequence: DMTHDMWCCGTTAYDTKVADHWD |
 |
 |
| Motif ID: | UP00624 |
| Motif name: | VENTX_R143C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.0247914 |
Alignment:
CWWVCMVMTABYATATVHDDTHS
------HVTACWGTABD------
| Original motif | Reverse complement motif |
| Consensus sequence: CWWVCMVMTABYATATVHDDTHS | Consensus sequence: SHAHHHVATATMBTAYVRGBWWG |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0.0266567 |
Alignment:
TWTTHVWATAVADATAAWHTHKG
-HVTACWGTABD-----------
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Dataset #: 4 | Motif ID: 44 | Motif name: dhACATTCTkh |
| Original motif | Reverse complement motif |
| Consensus sequence: DHACATTCTGH | Consensus sequence: HCAGAATGTHD |
 |
 |
Best Matches for Motif ID 44 (Highest to Lowest)
| Motif ID: | UP00401 |
| Motif name: | Sox4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 11 |
| Similarity score: | 0 |
Alignment:
HBVMCWTTGTTDDD
-DHACATTCTGH--
| Original motif | Reverse complement motif |
| Consensus sequence: DDDAACAAWGRVVH | Consensus sequence: HBVMCWTTGTTDDD |
 |
 |
| Motif ID: | UP00592 |
| Motif name: | GFI1B_A204T_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 11 |
| Similarity score: | 0.0061565 |
Alignment:
TDBHVVGVMCAGATTTVMAHHAD
--------HCAGAATGTHD----
| Original motif | Reverse complement motif |
| Consensus sequence: DTDHTRBAAATCTGRVCBVHBDA | Consensus sequence: TDBHVVGVMCAGATTTVMAHHAD |
 |
 |
| Motif ID: | UP00630 |
| Motif name: | ZNF655_E327G_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 11 |
| Similarity score: | 0.00867089 |
Alignment:
VVHBDHHKTTAGCATADHDHDH
--------HCAGAATGTHD---
| Original motif | Reverse complement motif |
| Consensus sequence: HDHDDDTATGCTAAYHHDBHVV | Consensus sequence: VVHBDHHKTTAGCATADHDHDH |
 |
 |
| Motif ID: | UP00630 |
| Motif name: | ZNF655_S393L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.00939906 |
Alignment:
VBDCWHHMTTAGCATADHDHDV
--------HCAGAATGTHD---
| Original motif | Reverse complement motif |
| Consensus sequence: BDHDDDTATGCTAAYHHWGHVV | Consensus sequence: VBDCWHHMTTAGCATADHDHDV |
 |
 |
| Motif ID: | UP00630 |
| Motif name: | ZNF655_S393L_R2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 11 |
| Similarity score: | 0.00973712 |
Alignment:
VVDHWHHKTTAGCATADHDDDV
--------HCAGAATGTHD---
| Original motif | Reverse complement motif |
| Consensus sequence: BDHDDDTATGCTAAYHHWDHVV | Consensus sequence: VVDHWHHKTTAGCATADHDDDV |
 |
 |
| Dataset #: 4 | Motif ID: 45 | Motif name: wbgTAAATAww |
| Original motif | Reverse complement motif |
| Consensus sequence: DBGTAAATAHD | Consensus sequence: DHTATTTACBD |
 |
 |
Best Matches for Motif ID 45 (Highest to Lowest)
| Motif ID: | UP00616 |
| Motif name: | POU3F4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0 |
Alignment:
BHGDBDHKTAATTAMBBBGHBC
------DHTATTTACBD-----
| Original motif | Reverse complement motif |
| Consensus sequence: GBHCBBBYTAATTARHHVDCDB | Consensus sequence: BHGDBDHKTAATTAMBBBGHBC |
 |
 |
| Motif ID: | UP00585 |
| Motif name: | BCL6 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 11 |
| Similarity score: | 0.00546358 |
Alignment:
TDRDVVGDTAATTAHCDBADDDH
------DHTATTTACBD------
| Original motif | Reverse complement motif |
| Consensus sequence: HHDHTBDGHTAATTAHCVBDKDA | Consensus sequence: TDRDVVGDTAATTAHCDBADDDH |
 |
 |
| Motif ID: | UP00596 |
| Motif name: | HOXC4_R158L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.00584685 |
Alignment:
WHHHDDGKTAATTAHCHHADWAH
-----DBGTAAATAHD-------
| Original motif | Reverse complement motif |
| Consensus sequence: WHHHDDGKTAATTAHCHHADWAH | Consensus sequence: HTWDTDHGHTAATTARCDDHHHW |
 |
 |
| Motif ID: | UP00589 |
| Motif name: | FOXC1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 11 |
| Similarity score: | 0.00675467 |
Alignment:
BHVVDDWATRTAAACAAWSDBDD
-------DBGTAAATAHD-----
| Original motif | Reverse complement motif |
| Consensus sequence: BHVVDDWATRTAAACAAWSDBDD | Consensus sequence: DHBHSWTTGTTTAMATWDDVBDB |
 |
 |
| Motif ID: | UP00618 |
| Motif name: | POU6F2_E639K_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 6 |
| Number of overlap: | 11 |
| Similarity score: | 0.00679091 |
Alignment:
HTDHTDMDYTAATTAVHBCBHVW
-------DHTATTTACBD-----
| Original motif | Reverse complement motif |
| Consensus sequence: WBHBGVDVTAATTAKHYHADDAH | Consensus sequence: HTDHTDMDYTAATTAVHBCBHVW |
 |
 |
| Dataset #: 5 | Motif ID: 46 | Motif name: TFW1 |
| Original motif | Reverse complement motif |
| Consensus sequence: GTCGCG | Consensus sequence: CGCGAC |
 |
 |
Best Matches for Motif ID 46 (Highest to Lowest)
| Motif ID: | UP00612 |
| Motif name: | PAX7 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0 |
Alignment:
VTHDBKSGTCACGGTTHDHDBV
-------GTCGCG---------
| Original motif | Reverse complement motif |
| Consensus sequence: VVDHDHAACCGTGACSRVDHAV | Consensus sequence: VTHDBKSGTCACGGTTHDHDBV |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 6 |
| Similarity score: | 0.00216016 |
Alignment:
GVHDBBKGACGCGTCVBBTCHH
---------CGCGAC-------
| Original motif | Reverse complement motif |
| Consensus sequence: GVHDBBKGACGCGTCVBBTCHH | Consensus sequence: HDGAVBVGACGCGTCYVBDDBC |
 |
 |
| Motif ID: | UP00521 |
| Motif name: | FOXN2_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 6 |
| Similarity score: | 0.00560752 |
Alignment:
HSDBHHBGACGCKWDHHKHRVD
-------GTCGCG---------
| Original motif | Reverse complement motif |
| Consensus sequence: DVKHRDHDWRGCGTCBHHBHSH | Consensus sequence: HSDBHHBGACGCKWDHHKHRVD |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0120041 |
Alignment:
SDYHHHMTAGCGTCBBMTHGADH
--------CGCGAC---------
| Original motif | Reverse complement motif |
| Consensus sequence: HDTCDARVBGACGCTAYHHHKHS | Consensus sequence: SDYHHHMTAGCGTCBBMTHGADH |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 6 |
| Similarity score: | 0.0150869 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
--------CGCGAC--------
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Dataset #: 5 | Motif ID: 47 | Motif name: TFW2 |
| Original motif | Reverse complement motif |
| Consensus sequence: SCGCGCGG | Consensus sequence: CCGCGCGS |
 |
 |
Best Matches for Motif ID 47 (Highest to Lowest)
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 10 |
| Number of overlap: | 8 |
| Similarity score: | 0 |
Alignment:
TDTKHHHGGGCGGGGDHHMWHA
-----SCGCGCGG---------
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 9 |
| Number of overlap: | 8 |
| Similarity score: | 0.00927263 |
Alignment:
KHHHTHHCCCCCGCAHDHDHHH
--------CCGCGCGS------
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 11 |
| Number of overlap: | 8 |
| Similarity score: | 0.0129979 |
Alignment:
AAHAHWHCTCCCCCRCDHHYVHM
----------CCGCGCGS-----
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 8 |
| Similarity score: | 0.0173097 |
Alignment:
HHHHDMATCCCYCGCATKVBTV
---------CCGCGCGS-----
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 7 |
| Number of overlap: | 8 |
| Similarity score: | 0.0235074 |
Alignment:
HDGAVBVGACGCGTCYVBDDBC
------CCGCGCGS--------
| Original motif | Reverse complement motif |
| Consensus sequence: GVHDBBKGACGCGTCVBBTCHH | Consensus sequence: HDGAVBVGACGCGTCYVBDDBC |
 |
 |
| Dataset #: 5 | Motif ID: 48 | Motif name: TFW3 |
| Original motif | Reverse complement motif |
| Consensus sequence: CGGCYBCGCG | Consensus sequence: CGCGBMGCCG |
 |
 |
Best Matches for Motif ID 48 (Highest to Lowest)
| Motif ID: | UP00522 |
| Motif name: | FOXN4_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 11 |
| Number of overlap: | 10 |
| Similarity score: | 0.0082598 |
Alignment:
SDYHHHMTAGCGTCBBMTHGADH
----------CGGCYBCGCG---
| Original motif | Reverse complement motif |
| Consensus sequence: HDTCDARVBGACGCTAYHHHKHS | Consensus sequence: SDYHHHMTAGCGTCBBMTHGADH |
 |
 |
| Motif ID: | UP00521 |
| Motif name: | FOXN2_primary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.00979086 |
Alignment:
DVKHRDHDWRGCGTCBHHBHSH
-----------CGGCYBCGCG-
| Original motif | Reverse complement motif |
| Consensus sequence: DVKHRDHDWRGCGTCBHHBHSH | Consensus sequence: HSDBHHBGACGCKWDHHKHRVD |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 10 |
| Similarity score: | 0.014104 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
--------CGGCYBCGCG----
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00521 |
| Motif name: | FOXN2_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 8 |
| Number of overlap: | 10 |
| Similarity score: | 0.0164065 |
Alignment:
MWBDVHCGGCGTCVTBAGWSRRD
------CGGCYBCGCG-------
| Original motif | Reverse complement motif |
| Consensus sequence: MWBDVHCGGCGTCVTBAGWSRRD | Consensus sequence: DMMSWCTBABGACGCCGDBDBWY |
 |
 |
| Motif ID: | UP00612 |
| Motif name: | PAX7 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 10 |
| Similarity score: | 0.0172722 |
Alignment:
VVDHDHAACCGTGACSRVDHAV
-CGCGBMGCCG-----------
| Original motif | Reverse complement motif |
| Consensus sequence: VVDHDHAACCGTGACSRVDHAV | Consensus sequence: VTHDBKSGTCACGGTTHDHDBV |
 |
 |
| Dataset #: 5 | Motif ID: 49 | Motif name: TFF1 |
| Original motif | Reverse complement motif |
| Consensus sequence: CGGVGCCGCVGC | Consensus sequence: GCVGCGGCBCCG |
 |
 |
Best Matches for Motif ID 49 (Highest to Lowest)
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 12 |
| Similarity score: | 0.0150923 |
Alignment:
DBBBSBDGGGGGCGYRDDHHHV
------CGGVGCCGCVGC----
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00521 |
| Motif name: | FOXN2_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 10 |
| Number of overlap: | 12 |
| Similarity score: | 0.0150987 |
Alignment:
DMMSWCTBABGACGCCGDBDBWY
---------CGGVGCCGCVGC--
| Original motif | Reverse complement motif |
| Consensus sequence: MWBDVHCGGCGTCVTBAGWSRRD | Consensus sequence: DMMSWCTBABGACGCCGDBDBWY |
 |
 |
| Motif ID: | UP00522 |
| Motif name: | FOXN4_secondary |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 11 |
| Number of overlap: | 12 |
| Similarity score: | 0.021229 |
Alignment:
HDGAVBVGACGCGTCYVBDDBC
CGGVGCCGCVGC----------
| Original motif | Reverse complement motif |
| Consensus sequence: GVHDBBKGACGCGTCVBBTCHH | Consensus sequence: HDGAVBVGACGCGTCYVBDDBC |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 12 |
| Similarity score: | 0.0250573 |
Alignment:
WDVDVHDRGTCTCCHTTDDHHBV
---GCVGCGGCBCCG--------
| Original motif | Reverse complement motif |
| Consensus sequence: VBHDDDAADGGAGACMHHVDVDW | Consensus sequence: WDVDVHDRGTCTCCHTTDDHHBV |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_S265Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 12 |
| Similarity score: | 0.0255393 |
Alignment:
VBHDDHARDGGAGACMHDVDVVA
--------CGGVGCCGCVGC---
| Original motif | Reverse complement motif |
| Consensus sequence: VBHDDHARDGGAGACMHDVDVVA | Consensus sequence: TBVDVDDRGTCTCCHKTHDHHBV |
 |
 |
| Dataset #: 5 | Motif ID: 50 | Motif name: TFF11 |
| Original motif | Reverse complement motif |
| Consensus sequence: GGMGGRGGCGGVGC | Consensus sequence: GCVCCGCCMCCYCC |
 |
 |
Best Matches for Motif ID 50 (Highest to Lowest)
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 5 |
| Number of overlap: | 14 |
| Similarity score: | 0 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
----GCVCCGCCMCCYCC----
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 14 |
| Similarity score: | 0.00204802 |
Alignment:
THWYHHDCCCCGCCCHHHYADA
------GCVCCGCCMCCYCC--
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 14 |
| Similarity score: | 0.0138317 |
Alignment:
AAHAHWHCTCCCCCRCDHHYVHM
------GCVCCGCCMCCYCC---
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 14 |
| Similarity score: | 0.0215847 |
Alignment:
KHHHTHHCCCCCGCAHDHDHHH
-------GCVCCGCCMCCYCC-
| Original motif | Reverse complement motif |
| Consensus sequence: KHHHTHHCCCCCGCAHDHDHHH | Consensus sequence: HHHDHHDTGCGGGGGHHAHHHY |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 14 |
| Similarity score: | 0.0260649 |
Alignment:
VABVRATGCGKGGGATRDDHHH
GGMGGRGGCGGVGC--------
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Dataset #: 5 | Motif ID: 51 | Motif name: TFM2 |
| Original motif | Reverse complement motif |
| Consensus sequence: RGRGGAGRRGGHGGDG | Consensus sequence: CHCCBCCKMCTCCKCM |
 |
 |
Best Matches for Motif ID 51 (Highest to Lowest)
| Motif ID: | UP00627 |
| Motif name: | WT1_REF_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 8 |
| Number of overlap: | 16 |
| Similarity score: | 0.0323246 |
Alignment:
AAHAHWHCTCCCCCRCDHHYVHM
-------CHCCBCCKMCTCCKCM
| Original motif | Reverse complement motif |
| Consensus sequence: AAHAHWHCTCCCCCRCDHHYVHM | Consensus sequence: RHVMHHDGMGGGGGAGHWHTHTT |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_H322Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 16 |
| Similarity score: | 0.033363 |
Alignment:
DDDDDHDGGGTGGGHHHDDDVVK
----RGRGGAGRRGGHGGDG---
| Original motif | Reverse complement motif |
| Consensus sequence: DDDDDHDGGGTGGGHHHDDDVVK | Consensus sequence: YVVDHDHHHCCCACCCHHDDDDD |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R366L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 16 |
| Similarity score: | 0.0368876 |
Alignment:
VABVRATGCGKGGGATRDDHHH
---RGRGGAGRRGGHGGDG---
| Original motif | Reverse complement motif |
| Consensus sequence: VABVRATGCGKGGGATRDDHHH | Consensus sequence: HHHHDMATCCCYCGCATKVBTV |
 |
 |
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 6 |
| Number of overlap: | 16 |
| Similarity score: | 0.0386109 |
Alignment:
BHHDDHMMCGCCCCCHBSBBVH
-----CHCCBCCKMCTCCKCM-
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00587 |
| Motif name: | EGR2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 16 |
| Similarity score: | 0.0406128 |
Alignment:
THWYHHDCCCCGCCCHHHYADA
----CHCCBCCKMCTCCKCM--
| Original motif | Reverse complement motif |
| Consensus sequence: THWYHHDCCCCGCCCHHHYADA | Consensus sequence: TDTKHHHGGGCGGGGDHHMWHA |
 |
 |
| Dataset #: 5 | Motif ID: 52 | Motif name: TFM1 |
| Original motif | Reverse complement motif |
| Consensus sequence: WTKTTTTTHWTTTTTTBT | Consensus sequence: ABAAAAAAWHAAAAARAW |
 |
 |
Best Matches for Motif ID 52 (Highest to Lowest)
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0388799 |
Alignment:
TWTTHVWATAHAWATAAWHTHKG
---ABAAAAAAWHAAAAARAW--
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0401933 |
Alignment:
TDTTHVWATAHADATAAWHTHKG
---ABAAAAAAWHAAAAARAW--
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 18 |
| Similarity score: | 0.0418262 |
Alignment:
TWTTHVWATAVADATAAWHTHKG
---ABAAAAAAWHAAAAARAW--
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394W_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0427632 |
Alignment:
TWTTHVWATAHAWATAAWHTHKG
---ABAAAAAAWHAAAAARAW--
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CYHAHWTTATWTDTATWBHAAWA |
 |
 |
| Motif ID: | UP00606 |
| Motif name: | NR1H4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0435661 |
Alignment:
ATTYWABTATWAATDDWTDBHWW
---WTKTTTTTHWTTTTTTBT--
| Original motif | Reverse complement motif |
| Consensus sequence: WWHVDAWDDATTWATAVTWMAAT | Consensus sequence: ATTYWABTATWAATDDWTDBHWW |
 |
 |
| Dataset #: 5 | Motif ID: 53 | Motif name: TFM3 |
| Original motif | Reverse complement motif |
| Consensus sequence: WWHTTTTTCABAAWTTWA | Consensus sequence: TWAAWTTVTGAAAAAHWW |
 |
 |
Best Matches for Motif ID 53 (Highest to Lowest)
| Motif ID: | UP00626 |
| Motif name: | VSX2_R200P_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 18 |
| Similarity score: | 0.0332522 |
Alignment:
HGDWWHDTTTWATATABWDDHDH
--WWHTTTTTCABAAWTTWA---
| Original motif | Reverse complement motif |
| Consensus sequence: HGDWWHDTTTWATATABWDDHDH | Consensus sequence: HDHDDWBTATATWAAADDWWDCH |
 |
 |
| Motif ID: | UP00606 |
| Motif name: | NR1H4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 4 |
| Number of overlap: | 18 |
| Similarity score: | 0.0360441 |
Alignment:
WWHVDAWDDATTWATAVTWMAAT
--TWAAWTTVTGAAAAAHWW---
| Original motif | Reverse complement motif |
| Consensus sequence: WWHVDAWDDATTWATAVTWMAAT | Consensus sequence: ATTYWABTATWAATDDWTDBHWW |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_F392L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 18 |
| Similarity score: | 0.0382798 |
Alignment:
CYHAHWTTATDTVTATWBHAAWA
-TWAAWTTVTGAAAAAHWW----
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAVADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTVTATWBHAAWA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 18 |
| Similarity score: | 0.0386758 |
Alignment:
TDTTHVWATAHADATAAWHTHKG
---TWAAWTTVTGAAAAAHWW--
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 18 |
| Similarity score: | 0.039146 |
Alignment:
TWTTHVWATAHAWATAAWHTHKG
---TWAAWTTVTGAAAAAHWW--
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Dataset #: 5 | Motif ID: 54 | Motif name: TFM12 |
| Original motif | Reverse complement motif |
| Consensus sequence: CYYCBBCYYYTCCHCCTYYY | Consensus sequence: KKKAGGDGGAKKMGBBGKMG |
 |
 |
Best Matches for Motif ID 54 (Highest to Lowest)
| Motif ID: | UP00600 |
| Motif name: | KLF11 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0322042 |
Alignment:
DBBBSBDGGGGGCGYRDDHHHV
KKKAGGDGGAKKMGBBGKMG--
| Original motif | Reverse complement motif |
| Consensus sequence: BHHDDHMMCGCCCCCHBSBBVH | Consensus sequence: DBBBSBDGGGGGCGYRDDHHHV |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_H322Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 20 |
| Similarity score: | 0.0344106 |
Alignment:
DDDDDHDGGGTGGGHHHDDDVVK
---KKKAGGDGGAKKMGBBGKMG
| Original motif | Reverse complement motif |
| Consensus sequence: DDDDDHDGGGTGGGHHHDDDVVK | Consensus sequence: YVVDHDHHHCCCACCCHHDDDDD |
 |
 |
| Motif ID: | UP00594 |
| Motif name: | HESX1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0386784 |
Alignment:
DTHGBBBWHDVHBHDBVDVBHB
-KKKAGGDGGAKKMGBBGKMG-
| Original motif | Reverse complement motif |
| Consensus sequence: BHBVHVBDHVHVDHWVBVCHAD | Consensus sequence: DTHGBBBWHDVHBHDBVDVBHB |
 |
 |
| Motif ID: | UP00615 |
| Motif name: | PITX2 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0403798 |
Alignment:
DDTBVRRKGGATTARADWDHAVH
-KKKAGGDGGAKKMGBBGKMG--
| Original motif | Reverse complement motif |
| Consensus sequence: DDTBVRRKGGATTARADWDHAVH | Consensus sequence: HVTHHWDTKTAATCCYKMBVADD |
 |
 |
| Motif ID: | UP00629 |
| Motif name: | ZNF200_S265Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0408448 |
Alignment:
VBHDDHARDGGAGACMHDVDVVA
--KKKAGGDGGAKKMGBBGKMG-
| Original motif | Reverse complement motif |
| Consensus sequence: VBHDDHARDGGAGACMHDVDVVA | Consensus sequence: TBVDVDDRGTCTCCHKTHDHHBV |
 |
 |
| Dataset #: 5 | Motif ID: 55 | Motif name: TFM13 |
| Original motif | Reverse complement motif |
| Consensus sequence: ATKAAWTTTTRMAABAHHTW | Consensus sequence: WAHHTVTTYKAAAAWTTRAT |
 |
 |
Best Matches for Motif ID 55 (Highest to Lowest)
| Motif ID: | UP00626 |
| Motif name: | VSX2_R200P_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Reverse Complement |
| Direction: | Backward |
| Position number: | 1 |
| Number of overlap: | 20 |
| Similarity score: | 0.0395806 |
Alignment:
HDHDDWBTATATWAAADDWWDCH
---WAHHTVTTYKAAAAWTTRAT
| Original motif | Reverse complement motif |
| Consensus sequence: HGDWWHDTTTWATATABWDDHDH | Consensus sequence: HDHDDWBTATATWAAADDWWDCH |
 |
 |
| Motif ID: | UP00606 |
| Motif name: | NR1H4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 4 |
| Number of overlap: | 20 |
| Similarity score: | 0.0396581 |
Alignment:
WWHVDAWDDATTWATAVTWMAAT
---WAHHTVTTYKAAAAWTTRAT
| Original motif | Reverse complement motif |
| Consensus sequence: WWHVDAWDDATTWATAVTWMAAT | Consensus sequence: ATTYWABTATWAATDDWTDBHWW |
 |
 |
| Motif ID: | UP00599 |
| Motif name: | KLF1_R328H_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0428521 |
Alignment:
RAAHBATAATDTATTADWAYHA
--ATKAAWTTTTRMAABAHHTW
| Original motif | Reverse complement motif |
| Consensus sequence: THMTWDTAATADATTATBHTTM | Consensus sequence: RAAHBATAATDTATTADWAYHA |
 |
 |
| Motif ID: | UP00604 |
| Motif name: | NKX2-5 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 20 |
| Similarity score: | 0.0431907 |
Alignment:
TAAWCHATDTADATTAHCTAAA
ATKAAWTTTTRMAABAHHTW--
| Original motif | Reverse complement motif |
| Consensus sequence: TAAWCHATDTADATTAHCTAAA | Consensus sequence: TTTAGHTAATDTADATHGWTTA |
 |
 |
| Motif ID: | UP00597 |
| Motif name: | HOXD13_S316C_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 20 |
| Similarity score: | 0.0441188 |
Alignment:
CDDHHWBCTCGTAAAADHDVHD
-WAHHTVTTYKAAAAWTTRAT-
| Original motif | Reverse complement motif |
| Consensus sequence: CDDHHWBCTCGTAAAADHDVHD | Consensus sequence: DHVDHDTTTTACGAGVWHHDDG |
 |
 |
| Dataset #: 5 | Motif ID: 56 | Motif name: TFM11 |
| Original motif | Reverse complement motif |
| Consensus sequence: HDWVAAAHAAAAAMAAAMWWWHBWA | Consensus sequence: TWVHWWWYTTTYTTTTTHTTTVWBH |
 |
 |
Best Matches for Motif ID 56 (Highest to Lowest)
| Motif ID: | UP00626 |
| Motif name: | VSX2_R200P_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Reverse Complement |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 23 |
| Similarity score: | 0.0548446 |
Alignment:
HDHDDWBTATATWAAADDWWDCH--
-TWVHWWWYTTTYTTTTTHTTTVWBH
| Original motif | Reverse complement motif |
| Consensus sequence: HGDWWHDTTTWATATABWDDHDH | Consensus sequence: HDHDDWBTATATWAAADDWWDCH |
 |
 |
| Motif ID: | UP00606 |
| Motif name: | NR1H4 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 3 |
| Number of overlap: | 23 |
| Similarity score: | 0.0574999 |
Alignment:
--WWHVDAWDDATTWATAVTWMAAT
HDWVAAAHAAAAAMAAAMWWWHBWA--
| Original motif | Reverse complement motif |
| Consensus sequence: WWHVDAWDDATTWATAVTWMAAT | Consensus sequence: ATTYWABTATWAATDDWTDBHWW |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_R394L_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Forward |
| Position number: | 2 |
| Number of overlap: | 23 |
| Similarity score: | 0.0606103 |
Alignment:
TWTTHVWATAHAWATAAWHTHKG--
-HDWVAAAHAAAAAMAAAMWWWHBWA
| Original motif | Reverse complement motif |
| Consensus sequence: TWTTHVWATAHAWATAAWHTHKG | Consensus sequence: CRHAHWTTATWTHTATWBHAAWA |
 |
 |
| Motif ID: | UP00619 |
| Motif name: | PROP1_R112Q_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Reverse Complement |
| Direction: | Forward |
| Position number: | 1 |
| Number of overlap: | 23 |
| Similarity score: | 0.0607001 |
Alignment:
VWVTDTWAHYTAATTARBHHWAH--
HDWVAAAHAAAAAMAAAMWWWHBWA
| Original motif | Reverse complement motif |
| Consensus sequence: VWVTDTWAHYTAATTARBHHWAH | Consensus sequence: HTWHHVMTAATTAMHTWAHAVWV |
 |
 |
| Motif ID: | UP00627 |
| Motif name: | WT1_H373Y_R1 |
| Species: | Homo sapiens |
| Matching format of first motif: | Original Motif |
| Matching format of second motif: | Original Motif |
| Direction: | Backward |
| Position number: | 2 |
| Number of overlap: | 23 |
| Similarity score: | 0.0619766 |
Alignment:
--TDTTHVWATAHADATAAWHTHKG
HDWVAAAHAAAAAMAAAMWWWHBWA-
| Original motif | Reverse complement motif |
| Consensus sequence: TDTTHVWATAHADATAAWHTHKG | Consensus sequence: CYHAHWTTATDTDTATWBHAADA |
 |
 |